

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147714.16 - phase: 0
(322 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
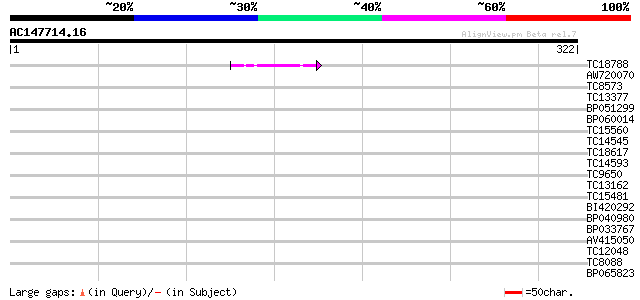
Score E
Sequences producing significant alignments: (bits) Value
TC18788 weakly similar to UP|Q9SSA6 (Q9SSA6) F4P13.1 protein, pa... 44 4e-05
AW720070 36 0.008
TC8573 similar to UP|Q9SIH5 (Q9SIH5) At2g36090 protein, partial ... 36 0.010
TC13377 similar to UP|Q9M5G4 (Q9M5G4) CDK-activating kinase, par... 35 0.017
BP051299 35 0.017
BP060014 35 0.022
TC15560 similar to SP|Q03173|NDPP_MOUSE NPC derived proline rich... 35 0.022
TC14545 similar to UP|Q9FT78 (Q9FT78) P23 co-chaperone, partial ... 33 0.065
TC18617 weakly similar to UP|Q9FPQ5 (Q9FPQ5) Gamete-specific hyd... 33 0.065
TC14593 similar to UP|Q8H1U3 (Q8H1U3) CDC26, partial (31%) 33 0.085
TC9650 similar to UP|O49485 (O49485) Phosphoglycerate dehydrogen... 33 0.085
TC13162 similar to UP|Q8C0D7 (Q8C0D7) CANDIDATE tumor suppressor... 33 0.085
TC15481 similar to UP|Q9LUR1 (Q9LUR1) RING zinc finger protein-l... 32 0.11
BI420292 32 0.11
BP040980 32 0.14
BP033767 32 0.14
AV415050 32 0.19
TC12048 weakly similar to UP|Q9LGA3 (Q9LGA3) ESTs D22655(C0749),... 31 0.25
TC8088 similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like arabinogalac... 31 0.25
BP065823 31 0.32
>TC18788 weakly similar to UP|Q9SSA6 (Q9SSA6) F4P13.1 protein, partial (7%)
Length = 951
Score = 43.9 bits (102), Expect = 4e-05
Identities = 20/52 (38%), Positives = 30/52 (57%)
Frame = +1
Query: 126 DEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQC 177
DE +C VC D + +D ++ CD C+ H C PL + IP+G+W+C C
Sbjct: 103 DEGVCKVC-GLDKD-DDSVLLCDTCDAEYHTYCLNPPLAR-IPEGNWYCPSC 249
>AW720070
Length = 484
Score = 36.2 bits (82), Expect = 0.008
Identities = 28/103 (27%), Positives = 50/103 (48%), Gaps = 9/103 (8%)
Frame = +1
Query: 21 QQQQQQEEQDLSHCSLLPT---KKRKESRNSSLFHTPPSSPTPPPSTYSLPTKKRITALQ 77
++++++EE++ ++ T ++ +E +SS H P SSP+ + S P+
Sbjct: 121 EEEEEEEEEEAGEEGIITTTREEEEEEEESSSKHHCPFSSPSNFSLSSSSPSITDYNNTH 300
Query: 78 PHLHHHN----NIP--NDAVPLIDLNVEYSPSLPSATPIEKQS 114
HLHHHN NIP N + + + +SP +AT S
Sbjct: 301 -HLHHHNHHVMNIPSQNSWLGMTHDDPHHSPEANTATESHNSS 426
>TC8573 similar to UP|Q9SIH5 (Q9SIH5) At2g36090 protein, partial (17%)
Length = 515
Score = 35.8 bits (81), Expect = 0.010
Identities = 21/65 (32%), Positives = 31/65 (47%), Gaps = 3/65 (4%)
Frame = +3
Query: 53 TPPSSPTPPPSTYSLPTKK--RITALQPHLHHHNNIPNDAVPLIDLNVEYSPSLP-SATP 109
TP S P+PPP+ S PT T L P +H + + P +PS P S+ P
Sbjct: 321 TPSSPPSPPPTALSSPTPSLPSTTLLSPTSYHPRRLRRSSSPPWTYTTRANPSFPESSAP 500
Query: 110 IEKQS 114
+K++
Sbjct: 501 RQKRA 515
>TC13377 similar to UP|Q9M5G4 (Q9M5G4) CDK-activating kinase, partial (26%)
Length = 439
Score = 35.0 bits (79), Expect = 0.017
Identities = 18/54 (33%), Positives = 27/54 (49%)
Frame = +3
Query: 8 TDLPPNKRLRFIYQQQQQQEEQDLSHCSLLPTKKRKESRNSSLFHTPPSSPTPP 61
T PP+++ R + Q + S T+ S +SS+ +PPSSPTPP
Sbjct: 195 TTSPPSEKSRLSRCSKASQTSSSCTSTSGKKTRTPSSSSSSSVLTSPPSSPTPP 356
>BP051299
Length = 539
Score = 35.0 bits (79), Expect = 0.017
Identities = 19/72 (26%), Positives = 36/72 (49%), Gaps = 1/72 (1%)
Frame = +3
Query: 36 LLPTKKRKESRNSSLFH-TPPSSPTPPPSTYSLPTKKRITALQPHLHHHNNIPNDAVPLI 94
L P S ++LF PP S +PPPST + P+ + +TA + + + +
Sbjct: 177 LFPISSTTPSAATTLFSPNPPPSASPPPSTVNSPSSESVTASSASPPPKTAVISSSATPL 356
Query: 95 DLNVEYSPSLPS 106
+++ +SP+ P+
Sbjct: 357 FVSISHSPNPPT 392
>BP060014
Length = 366
Score = 34.7 bits (78), Expect = 0.022
Identities = 23/71 (32%), Positives = 30/71 (41%), Gaps = 1/71 (1%)
Frame = +3
Query: 211 VHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLE-KCYVCGCFDGCALVCSEQKCGLGF 269
VH C P+V+FVD E K R + KC CG G AL C + C +
Sbjct: 147 VHRCCVDWAPQVYFVD----ETCKNLKAEVARGAKLKCSTCG-LKGAALGCYVKSCKRTY 311
Query: 270 HITCGIKEDLC 280
H+ C + C
Sbjct: 312 HVPCAMDVSTC 344
>TC15560 similar to SP|Q03173|NDPP_MOUSE NPC derived proline rich protein
1 (NDPP-1). {Mus musculus;}, partial (8%)
Length = 1134
Score = 34.7 bits (78), Expect = 0.022
Identities = 21/57 (36%), Positives = 28/57 (48%), Gaps = 3/57 (5%)
Frame = +1
Query: 35 SLLPTKKRKESRNSSLFHTPPSSPT---PPPSTYSLPTKKRITALQPHLHHHNNIPN 88
SL+ K + E + HTPP SP+ P +T SLP + PHL H N P+
Sbjct: 13 SLINPKPKNEEQLKPQAHTPPLSPSQTNPTTTTLSLPPPLLLLPPPPHLLHLNLHPH 183
>TC14545 similar to UP|Q9FT78 (Q9FT78) P23 co-chaperone, partial (46%)
Length = 1127
Score = 33.1 bits (74), Expect = 0.065
Identities = 16/49 (32%), Positives = 25/49 (50%)
Frame = -1
Query: 33 HCSLLPTKKRKESRNSSLFHTPPSSPTPPPSTYSLPTKKRITALQPHLH 81
HC + K +E +S FH+ SS + PS + P+K T++ H H
Sbjct: 650 HCIITHPTKFREIHSSQSFHSSQSSHSTHPSHSTHPSKSSKTSITAHTH 504
>TC18617 weakly similar to UP|Q9FPQ5 (Q9FPQ5) Gamete-specific
hydroxyproline-rich glycoprotein a2, partial (13%)
Length = 578
Score = 33.1 bits (74), Expect = 0.065
Identities = 23/65 (35%), Positives = 30/65 (45%), Gaps = 8/65 (12%)
Frame = +3
Query: 53 TPPSSPTPP---PSTYSLPT----KKRITALQPHLHHHNNIPNDAVPLIDLNVEYSPSLP 105
+PP PTPP PS++SL T K L P HH + P P + + S P
Sbjct: 198 SPPPGPTPPPPAPSSHSLSTLAGSSKSPPCLTPPRHHRHQTPPPQKPTSSASTAATASRP 377
Query: 106 -SATP 109
SA+P
Sbjct: 378 RSASP 392
>TC14593 similar to UP|Q8H1U3 (Q8H1U3) CDC26, partial (31%)
Length = 1112
Score = 32.7 bits (73), Expect = 0.085
Identities = 24/79 (30%), Positives = 36/79 (45%), Gaps = 10/79 (12%)
Frame = +2
Query: 39 TKKRKESRNSSLFHTPPSSPTPPPSTYSL----------PTKKRITALQPHLHHHNNIPN 88
+K +K+SRN + PP+ P PP L P KRI P LH ++ +
Sbjct: 53 SKTKKKSRNIAAVTPPPTPPQQPPLAPPLSFVSSIAPRIPLPKRIALASP-LHKPFSVRS 229
Query: 89 DAVPLIDLNVEYSPSLPSA 107
++ D+ VE LP+A
Sbjct: 230 LSLSPSDMEVEVKLRLPNA 286
>TC9650 similar to UP|O49485 (O49485) Phosphoglycerate dehydrogenase-like
protein , partial (47%)
Length = 1041
Score = 32.7 bits (73), Expect = 0.085
Identities = 20/72 (27%), Positives = 30/72 (40%)
Frame = +2
Query: 38 PTKKRKESRNSSLFHTPPSSPTPPPSTYSLPTKKRITALQPHLHHHNNIPNDAVPLIDLN 97
P S ++ P SS +PPPST + P+ ++ + + P P I
Sbjct: 107 PPPSPSPSAATATVPAPSSSSSPPPSTPNPPSSSPRSSAKRVSNSSRTSPTSTAPTISPL 286
Query: 98 VEYSPSLPSATP 109
+P PSATP
Sbjct: 287 RSSAPRSPSATP 322
>TC13162 similar to UP|Q8C0D7 (Q8C0D7) CANDIDATE tumor suppressor P33 ING1
homolog, partial (21%)
Length = 508
Score = 32.7 bits (73), Expect = 0.085
Identities = 19/81 (23%), Positives = 33/81 (40%), Gaps = 1/81 (1%)
Frame = +2
Query: 107 ATPIEKQSQKQDIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQ 166
A P D++ VD + C C+ +V CD + + +G +++
Sbjct: 32 AQPTSVNPTGMDLDLPVDPNAPTYCFCNQVSYGE---MVACDNPDCKIEWFHFGCVGLRE 202
Query: 167 IPDGDWFCDQ-CRFKNDIDTD 186
P G W+C Q C +K D +
Sbjct: 203 QPQGKWYCSQLCSYKKLADKE 265
>TC15481 similar to UP|Q9LUR1 (Q9LUR1) RING zinc finger protein-like,
partial (32%)
Length = 706
Score = 32.3 bits (72), Expect = 0.11
Identities = 25/75 (33%), Positives = 32/75 (42%)
Frame = +3
Query: 11 PPNKRLRFIYQQQQQQEEQDLSHCSLLPTKKRKESRNSSLFHTPPSSPTPPPSTYSLPTK 70
P KRLR Q Q+Q+ L H SLL + + PP SP PPP P+
Sbjct: 63 PHAKRLRRQRLQPQRQDHALLHHPSLLHRRLNGLPPHLRPLVPPPRSP-PPPPQPPPPSS 239
Query: 71 KRITALQPHLHHHNN 85
PH HH++
Sbjct: 240 PARVLP*PHQPHHSS 284
>BI420292
Length = 553
Score = 32.3 bits (72), Expect = 0.11
Identities = 19/72 (26%), Positives = 30/72 (41%), Gaps = 8/72 (11%)
Frame = +2
Query: 111 EKQSQKQDIEFEVDDDEI-LCCVCHS----TDANAEDPIVFCDGCNLMVHASC---YGNP 162
E + Q F V+ DE C +C D ++ CD C H C +
Sbjct: 275 ENKDVDQGRSFLVEHDEGGQCALCGGHDFIKDVFGPRTVMICDQCEKEYHVECLKDHNKQ 454
Query: 163 LVKQIPDGDWFC 174
++++P+G WFC
Sbjct: 455 NLEKLPEGKWFC 490
Score = 30.4 bits (67), Expect = 0.42
Identities = 22/74 (29%), Positives = 30/74 (39%), Gaps = 2/74 (2%)
Frame = +2
Query: 129 LCCVCHSTDANAEDPIVFCDG--CNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTD 186
LC +C +N D ++ CD C H C P +IP G W+C C + + D
Sbjct: 131 LCSLC----SNGGD-LISCDRDKCTRAFHLKCARLP---RIPSGLWYCKYCENETQENKD 286
Query: 187 TGPIRCSLCPTKEG 200
R L EG
Sbjct: 287 VDQGRSFLVEHDEG 328
>BP040980
Length = 452
Score = 32.0 bits (71), Expect = 0.14
Identities = 14/22 (63%), Positives = 19/22 (85%)
Frame = -1
Query: 301 SQIWEKNKGSGKYKIVAVEDKE 322
S +WEK + SG+Y+IVAVEDK+
Sbjct: 452 SLLWEKLQLSGEYQIVAVEDKK 387
>BP033767
Length = 541
Score = 32.0 bits (71), Expect = 0.14
Identities = 26/78 (33%), Positives = 39/78 (49%), Gaps = 5/78 (6%)
Frame = +2
Query: 32 SHCSLLPTKKRKESRNSSLFHTPPSS-PTPPPST---YSLPTKKRITA-LQPHLHHHNNI 86
S +L+ T ++E +L PS+ TP P+T SL +K +I L PH HHH
Sbjct: 101 SSATLIRTHHQQEDPPPTLHTRLPSTLSTPKPTTTITISLTSKPKIPIFLHPHNHHHPKC 280
Query: 87 PNDAVPLIDLNVEYSPSL 104
+ +P I ++ S SL
Sbjct: 281 LHTFLPHIPISTTTSFSL 334
Score = 27.7 bits (60), Expect = 2.7
Identities = 13/41 (31%), Positives = 18/41 (43%)
Frame = +1
Query: 52 HTPPSSPTPPPSTYSLPTKKRITALQPHLHHHNNIPNDAVP 92
H PP PPP +++ HLHHH +P + P
Sbjct: 241 HFPPPPQPPPPKM-----PPHLSSPHTHLHHHLLLPPPSPP 348
>AV415050
Length = 357
Score = 31.6 bits (70), Expect = 0.19
Identities = 17/51 (33%), Positives = 22/51 (42%), Gaps = 1/51 (1%)
Frame = +2
Query: 128 ILCCVCHSTDANAEDPIVFCDGCNL-MVHASCYGNPLVKQIPDGDWFCDQC 177
++C C D D VFCD C + +H C P V D W C+ C
Sbjct: 104 MVCLKCG--DEGFSDTFVFCDKCQVYALHRYCLDGP-VNFYDDVTWLCEDC 247
>TC12048 weakly similar to UP|Q9LGA3 (Q9LGA3) ESTs D22655(C0749), partial
(20%)
Length = 697
Score = 31.2 bits (69), Expect = 0.25
Identities = 21/78 (26%), Positives = 37/78 (46%)
Frame = -3
Query: 8 TDLPPNKRLRFIYQQQQQQEEQDLSHCSLLPTKKRKESRNSSLFHTPPSSPTPPPSTYSL 67
++ P + R + IYQ++ +++ H S P + +S+ + PS +PPP + S
Sbjct: 542 SEYPADSRTKPIYQREMSPDQK---HQSSTPPPPQSP*ASSA---SSPSPSSPPPPSRS- 384
Query: 68 PTKKRITALQPHLHHHNN 85
QPH H H+N
Sbjct: 383 ---------QPHPHRHHN 357
>TC8088 similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like
arabinogalactan-protein 2, partial (69%)
Length = 1527
Score = 31.2 bits (69), Expect = 0.25
Identities = 24/80 (30%), Positives = 30/80 (37%), Gaps = 2/80 (2%)
Frame = +2
Query: 32 SHCSLLPTKKRKESRNSSLFH--TPPSSPTPPPSTYSLPTKKRITALQPHLHHHNNIPND 89
S+ LP S + FH PP P PP + +LP T P P+
Sbjct: 41 SNSHYLPLTLLSSSSS*QPFHFLPPPPQPPPPMAATTLPAFSPNTPTSP--PSTTTSPSP 214
Query: 90 AVPLIDLNVEYSPSLPSATP 109
P + SPS PS TP
Sbjct: 215 TSPARSTTAKPSPSSPSTTP 274
>BP065823
Length = 534
Score = 30.8 bits (68), Expect = 0.32
Identities = 17/47 (36%), Positives = 27/47 (57%)
Frame = +2
Query: 40 KKRKESRNSSLFHTPPSSPTPPPSTYSLPTKKRITALQPHLHHHNNI 86
K +K ++ SS FH P P P+T S KR+ + +++HH+NI
Sbjct: 50 KMKKITKYSSSFHFP*--PK*VPTTSSXSPYKRLPCVWVYIYHHHNI 184
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.137 0.445
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,060,367
Number of Sequences: 28460
Number of extensions: 164000
Number of successful extensions: 2536
Number of sequences better than 10.0: 190
Number of HSP's better than 10.0 without gapping: 2136
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2418
length of query: 322
length of database: 4,897,600
effective HSP length: 90
effective length of query: 232
effective length of database: 2,336,200
effective search space: 541998400
effective search space used: 541998400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC147714.16


