

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147712.6 + phase: 0
(80 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
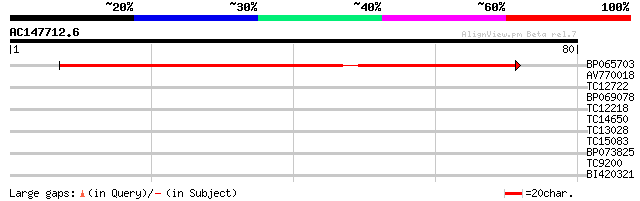
Score E
Sequences producing significant alignments: (bits) Value
BP065703 91 4e-20
AV770018 28 0.23
TC12722 similar to UP|MFP1_TOBAC (Q9M7J4) MAR binding filament-l... 26 1.2
BP069078 26 1.5
TC12218 26 1.5
TC14650 similar to UP|IF3A_TOBAC (Q40554) Eukaryotic translation... 25 2.6
TC13028 similar to UP|Q9C698 (Q9C698) Mysoin-like protein, parti... 25 2.6
TC15083 similar to UP|GAR1_YEAST (P28007) snoRNP protein GAR1, p... 25 3.4
BP073825 24 4.4
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 24 5.8
BI420321 23 7.5
>BP065703
Length = 480
Score = 90.9 bits (224), Expect = 4e-20
Identities = 46/65 (70%), Positives = 54/65 (82%)
Frame = -3
Query: 8 EKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVFSD 67
E +LD +RSS+KKL+EEL +NN IAEL+SETGE +Q+L EEEIR CKNMIKCTV SD
Sbjct: 475 EMELDRERSSRKKLQEELRELNNXIAELSSETGEAAIQKL--EEEIRACKNMIKCTVCSD 302
Query: 68 RPKEV 72
RPKEV
Sbjct: 301 RPKEV 287
>AV770018
Length = 599
Score = 28.5 bits (62), Expect = 0.23
Identities = 20/51 (39%), Positives = 28/51 (54%), Gaps = 3/51 (5%)
Frame = +2
Query: 20 KLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIK---CTVFSD 67
K E L+ V +I E ETG V + ++E+EI VC+ IK CT S+
Sbjct: 260 KFEPNLITVK-RIKESIEETGFKVDEVYDDEQEISVCQVRIKGMACTSCSE 409
>TC12722 similar to UP|MFP1_TOBAC (Q9M7J4) MAR binding filament-like
protein 1-1, partial (7%)
Length = 744
Score = 26.2 bits (56), Expect = 1.2
Identities = 14/42 (33%), Positives = 22/42 (52%)
Frame = +3
Query: 9 KDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEE 50
K+ ++ S KKLEEEL + +I L S+ + V E+
Sbjct: 36 KERETLDSRGKKLEEELASAKGEILRLRSQINSSKVAVSNEK 161
>BP069078
Length = 452
Score = 25.8 bits (55), Expect = 1.5
Identities = 19/54 (35%), Positives = 27/54 (49%), Gaps = 2/54 (3%)
Frame = -2
Query: 7 SEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIR--VCKN 58
SEKDLD R LE E + + +I EL E + +V LE + + C+N
Sbjct: 451 SEKDLDDLRKKLDTLELENIEKDLKIKELEKEK-KNLVTSLEIDHAVLKVTCEN 293
>TC12218
Length = 737
Score = 25.8 bits (55), Expect = 1.5
Identities = 13/46 (28%), Positives = 25/46 (54%)
Frame = +2
Query: 16 SSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIK 61
S ++ +L N ++ E+N++ E + + E+ EIR KN +K
Sbjct: 389 SDAVRVVTQLRNEAEKLKEMNNDLQEKIKELKAEKNEIRDEKNKLK 526
>TC14650 similar to UP|IF3A_TOBAC (Q40554) Eukaryotic translation
initiation factor 3 subunit 10 (eIF-3 theta)
(Eukaryotic translation initiation factor 3 large
subunit) (eIF3a) (PNLA-35), partial (11%)
Length = 978
Score = 25.0 bits (53), Expect = 2.6
Identities = 11/27 (40%), Positives = 17/27 (62%)
Frame = +1
Query: 54 RVCKNMIKCTVFSDRPKEVKTGQLIQL 80
R CKN++K + PKE K +L++L
Sbjct: 34 RSCKNLLKPWIIWKEPKEKKLLRLLKL 114
>TC13028 similar to UP|Q9C698 (Q9C698) Mysoin-like protein, partial (10%)
Length = 605
Score = 25.0 bits (53), Expect = 2.6
Identities = 13/41 (31%), Positives = 24/41 (57%), Gaps = 2/41 (4%)
Frame = +2
Query: 8 EKDLDSKRSSQKKLEEELMNVNNQIAELNS--ETGETVVQQ 46
E + + +KLE+E+ +N +++ NS ET E +V+Q
Sbjct: 374 EDQVKTYEEKVQKLEDEMQEMNEKLSAANSEIETKEGMVKQ 496
>TC15083 similar to UP|GAR1_YEAST (P28007) snoRNP protein GAR1, partial (7%)
Length = 1367
Score = 24.6 bits (52), Expect = 3.4
Identities = 8/15 (53%), Positives = 13/15 (86%)
Frame = +1
Query: 64 VFSDRPKEVKTGQLI 78
+FSDRPK + TG+++
Sbjct: 313 IFSDRPKPIATGKVL 357
>BP073825
Length = 416
Score = 24.3 bits (51), Expect = 4.4
Identities = 10/22 (45%), Positives = 14/22 (63%)
Frame = +3
Query: 1 MSVASASEKDLDSKRSSQKKLE 22
MS + +KDLDS+ S K +E
Sbjct: 198 MSFSGVEDKDLDSRTSGSKTVE 263
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 23.9 bits (50), Expect = 5.8
Identities = 14/35 (40%), Positives = 19/35 (54%)
Frame = +1
Query: 18 QKKLEEELMNVNNQIAELNSETGETVVQQLEEEEE 52
Q KLE E+ + E+ E E V ++ EEEEE
Sbjct: 166 QNKLEAEV----QEYEEVEEEVEEEVEEEEEEEEE 258
>BI420321
Length = 450
Score = 23.5 bits (49), Expect = 7.5
Identities = 14/54 (25%), Positives = 26/54 (47%)
Frame = -1
Query: 15 RSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVFSDR 68
R ++ +++ + + EL +E + + EEE E RV + CT+F R
Sbjct: 213 RRGSSSVQADIVCNDFGVRELRAEKEKEKGKWREEEGEERVWGDFSVCTIFIGR 52
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.304 0.122 0.303
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 806,710
Number of Sequences: 28460
Number of extensions: 5680
Number of successful extensions: 38
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 36
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 37
length of query: 80
length of database: 4,897,600
effective HSP length: 56
effective length of query: 24
effective length of database: 3,303,840
effective search space: 79292160
effective search space used: 79292160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.9 bits)
S2: 48 (23.1 bits)
Medicago: description of AC147712.6


