

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147712.16 + phase: 0
(236 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
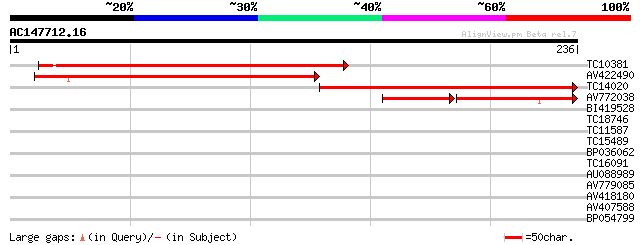
Score E
Sequences producing significant alignments: (bits) Value
TC10381 similar to UP|Q9LTU6 (Q9LTU6) Gb|AAF50915.1, partial (48%) 211 9e-56
AV422490 198 6e-52
TC14020 similar to UP|Q9LTU6 (Q9LTU6) Gb|AAF50915.1, partial (48%) 197 1e-51
AV772038 70 2e-23
BI419528 28 1.1
TC18746 weakly similar to UP|Q7XIG8 (Q7XIG8) AcinusL protein-lik... 28 1.8
TC11587 homologue to UP|Q9FN36 (Q9FN36) Gb|AAF34833.1, partial (... 27 2.4
TC15489 similar to UP|DDL_BACSU (P96612) D-alanine--D-alanine li... 27 3.1
BP036062 27 4.0
TC16091 26 6.9
AU088989 26 6.9
AV779085 26 6.9
AV418180 25 9.0
AV407588 25 9.0
BP054799 25 9.0
>TC10381 similar to UP|Q9LTU6 (Q9LTU6) Gb|AAF50915.1, partial (48%)
Length = 437
Score = 211 bits (537), Expect = 9e-56
Identities = 103/129 (79%), Positives = 112/129 (85%)
Frame = +2
Query: 13 SCSLNSCTMASSSMEVDNVLSEQKVLPPKPKFEPLKPHEMPGAAVQFRKVSVPPHRYTPL 72
SC++ S A S+MEV+ SE KVLPPKP FEPLKPHEM VQFRKVSVPPHRYTPL
Sbjct: 53 SCAMQS-NEAPSTMEVEASPSEPKVLPPKPLFEPLKPHEMSDGQVQFRKVSVPPHRYTPL 229
Query: 73 KKVWMDIYTPVFEQMKIDIRMNLKGRKVELKTRHDTPDISNLQKCADFVHAFMLGFDVID 132
KK WMD+YTP++EQMKID+RMNLK RKVELKTR DTPDISNLQKCADFVHAFMLGFDVID
Sbjct: 230 KKAWMDLYTPIYEQMKIDVRMNLKARKVELKTRRDTPDISNLQKCADFVHAFMLGFDVID 409
Query: 133 AIAILRLDE 141
AIA+LRLDE
Sbjct: 410 AIALLRLDE 436
>AV422490
Length = 471
Score = 198 bits (504), Expect = 6e-52
Identities = 99/123 (80%), Positives = 104/123 (84%), Gaps = 4/123 (3%)
Frame = +2
Query: 11 LFSCSLNSCTMAS----SSMEVDNVLSEQKVLPPKPKFEPLKPHEMPGAAVQFRKVSVPP 66
+F S NSCTM S SSMEV+ V SE K+LPPKPKFEPLK HEM VQFRKVSVPP
Sbjct: 101 VFRYS*NSCTMQSNEAPSSMEVEAVPSEPKLLPPKPKFEPLKSHEMSDGQVQFRKVSVPP 280
Query: 67 HRYTPLKKVWMDIYTPVFEQMKIDIRMNLKGRKVELKTRHDTPDISNLQKCADFVHAFML 126
HRYTPLKK WMD+YTPV+EQMKIDIRMNLK RKVELKTR DTPDISNLQKCADFVHAFML
Sbjct: 281 HRYTPLKKAWMDLYTPVYEQMKIDIRMNLKARKVELKTRRDTPDISNLQKCADFVHAFML 460
Query: 127 GFD 129
GFD
Sbjct: 461 GFD 469
>TC14020 similar to UP|Q9LTU6 (Q9LTU6) Gb|AAF50915.1, partial (48%)
Length = 546
Score = 197 bits (501), Expect = 1e-51
Identities = 101/107 (94%), Positives = 105/107 (97%)
Frame = +2
Query: 130 VIDAIAILRLDELYVESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKFTIENASKTRIVIA 189
VIDAIA+LRLDELY+ESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKF IENASKTRIVIA
Sbjct: 2 VIDAIALLRLDELYIESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKFAIENASKTRIVIA 181
Query: 190 DTKIHILGSFANIKIARDSLCYLIMGSPAGKVYSKLRAVTSRMAERF 236
D+KIHILGSFANIKIARDSLC LIMGSPAGKVYSKLRAVTSR+AERF
Sbjct: 182 DSKIHILGSFANIKIARDSLCSLIMGSPAGKVYSKLRAVTSRLAERF 322
>AV772038
Length = 465
Score = 70.1 bits (170), Expect(2) = 2e-23
Identities = 37/51 (72%), Positives = 43/51 (83%), Gaps = 1/51 (1%)
Frame = -2
Query: 187 VIADTKIHILGSFANIKIARDSLCYLIMGSPAG-KVYSKLRAVTSRMAERF 236
VIAD++IHILGSFA +KIARDSLC LI+GSPAG + RAVTSR+AERF
Sbjct: 368 VIADSQIHILGSFAYLKIARDSLCSLILGSPAGQRCTPNYRAVTSRLAERF 216
Score = 54.7 bits (130), Expect(2) = 2e-23
Identities = 26/30 (86%), Positives = 28/30 (92%)
Frame = -3
Query: 156 RGDHLSRAIGRLSGKGGKTKFTIENASKTR 185
RGDHLSRAIGRLSG GGKTKF IENAS+T+
Sbjct: 463 RGDHLSRAIGRLSG*GGKTKFAIENASQTQ 374
>BI419528
Length = 371
Score = 28.5 bits (62), Expect = 1.1
Identities = 11/28 (39%), Positives = 16/28 (56%)
Frame = +2
Query: 1 MLHLLCFFNQLFSCSLNSCTMASSSMEV 28
M+H++ +N LF CS C M + S V
Sbjct: 68 MIHVMISYNHLFYCSCQFCKMGTRSWHV 151
>TC18746 weakly similar to UP|Q7XIG8 (Q7XIG8) AcinusL protein-like, partial
(6%)
Length = 709
Score = 27.7 bits (60), Expect = 1.8
Identities = 17/63 (26%), Positives = 31/63 (48%), Gaps = 1/63 (1%)
Frame = +3
Query: 13 SCSLNSCTMASSSMEVDNVLSEQKVLPPKPKFEPLKPHEMPGAAVQF-RKVSVPPHRYTP 71
S S+N ++ DNV E+ ++ P+ EP +++PG V V P + TP
Sbjct: 195 SVSINQKNELKDTIIADNVKLEEDIVRPEMVEEPSSNNDIPGHDVSHSMDVGEPHKKKTP 374
Query: 72 LKK 74
+++
Sbjct: 375 VEE 383
>TC11587 homologue to UP|Q9FN36 (Q9FN36) Gb|AAF34833.1, partial (36%)
Length = 697
Score = 27.3 bits (59), Expect = 2.4
Identities = 18/47 (38%), Positives = 24/47 (50%), Gaps = 4/47 (8%)
Frame = +3
Query: 7 FFNQLFSCSLNSCTMASSSMEVDNVLSEQKVLPPKPKF----EPLKP 49
+ +QLFS + NS SSS + + LS + PP P F PL P
Sbjct: 414 YLSQLFSSTRNS----SSSTSIPSKLSTIPIRPPPPLFNWGQRPLNP 542
>TC15489 similar to UP|DDL_BACSU (P96612) D-alanine--D-alanine ligase
(D-alanylalanine synthetase) (D-Ala-D-Ala ligase) ,
partial (5%)
Length = 638
Score = 26.9 bits (58), Expect = 3.1
Identities = 10/25 (40%), Positives = 16/25 (64%)
Frame = +1
Query: 4 LLCFFNQLFSCSLNSCTMASSSMEV 28
++ F LFSCS N C + +S M++
Sbjct: 535 IIVIFFSLFSCSFNLCQLKNSFMKI 609
>BP036062
Length = 512
Score = 26.6 bits (57), Expect = 4.0
Identities = 16/73 (21%), Positives = 30/73 (40%)
Frame = -2
Query: 4 LLCFFNQLFSCSLNSCTMASSSMEVDNVLSEQKVLPPKPKFEPLKPHEMPGAAVQFRKVS 63
LL +L + +SC S++ N+ + +PP K +P+E P A + +
Sbjct: 274 LLSILRKLITFFSSSCKEQ*SNLNFFNMSKNLQHIPPNNKQMENQPNESPAARFKIETLP 95
Query: 64 VPPHRYTPLKKVW 76
P + +W
Sbjct: 94 ANFQFQGPRRTIW 56
>TC16091
Length = 627
Score = 25.8 bits (55), Expect = 6.9
Identities = 9/16 (56%), Positives = 13/16 (81%)
Frame = +2
Query: 6 CFFNQLFSCSLNSCTM 21
C F+QL SC++ SCT+
Sbjct: 356 CLFSQLDSCTILSCTL 403
>AU088989
Length = 186
Score = 25.8 bits (55), Expect = 6.9
Identities = 18/63 (28%), Positives = 33/63 (51%)
Frame = -2
Query: 98 RKVELKTRHDTPDISNLQKCADFVHAFMLGFDVIDAIAILRLDELYVESFEIKDVKTLRG 157
RK+EL ++ ++ + F F+L +++A+ LR L+ + F +K K +R
Sbjct: 179 RKIELXCXGESLEVVLXVRFTGFGVLFLLXLLLVEAVXFLR---LFCDFFAMKAHK-IRE 12
Query: 158 DHL 160
DHL
Sbjct: 11 DHL 3
>AV779085
Length = 508
Score = 25.8 bits (55), Expect = 6.9
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = +3
Query: 91 IRMNLKGRKVELKTRHDTPDISNLQK 116
I++ L GR+ ELK H TP Q+
Sbjct: 273 IKLGLSGRRQELKILHQTPQSDAKQR 350
>AV418180
Length = 363
Score = 25.4 bits (54), Expect = 9.0
Identities = 11/16 (68%), Positives = 11/16 (68%)
Frame = +1
Query: 41 KPKFEPLKPHEMPGAA 56
KPKFE LKP E P A
Sbjct: 28 KPKFELLKPEEAPRVA 75
>AV407588
Length = 277
Score = 25.4 bits (54), Expect = 9.0
Identities = 11/28 (39%), Positives = 17/28 (60%)
Frame = -2
Query: 181 ASKTRIVIADTKIHILGSFANIKIARDS 208
+SKT IV+ K ++ F N++ RDS
Sbjct: 96 SSKTAIVVVCVKAQVIFLFCNLQTERDS 13
>BP054799
Length = 430
Score = 25.4 bits (54), Expect = 9.0
Identities = 13/35 (37%), Positives = 20/35 (57%), Gaps = 3/35 (8%)
Frame = -1
Query: 111 ISNLQKCADFVHAFML---GFDVIDAIAILRLDEL 142
+ +L C DFVH F+L G ++A++RL L
Sbjct: 205 VDSLCWCLDFVHMFLLCDSGLHYATSLALMRLMNL 101
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.138 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,957,355
Number of Sequences: 28460
Number of extensions: 53872
Number of successful extensions: 319
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 318
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 318
length of query: 236
length of database: 4,897,600
effective HSP length: 87
effective length of query: 149
effective length of database: 2,421,580
effective search space: 360815420
effective search space used: 360815420
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 54 (25.4 bits)
Medicago: description of AC147712.16


