

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147538.3 - phase: 0
(123 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
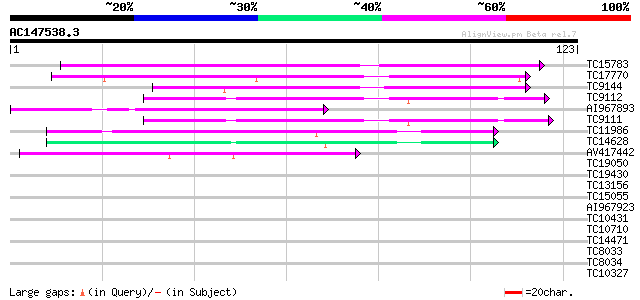
Score E
Sequences producing significant alignments: (bits) Value
TC15783 similar to UP|Q9LJ86 (Q9LJ86) Gb|AAD48513.1 (AT3g22600/F... 58 4e-10
TC17770 weakly similar to UP|Q8VYI9 (Q8VYI9) AT5g64080/MHJ24_6, ... 50 1e-07
TC9144 44 6e-06
TC9112 weakly similar to UP|Q9SI70 (Q9SI70) F23N19.16, partial (... 41 5e-05
AI967893 41 7e-05
TC9111 39 2e-04
TC11986 weakly similar to UP|Q8RYA8 (Q8RYA8) Lipid transfer prec... 39 3e-04
TC14628 similar to UP|NLTP_CICAR (O23758) Nonspecific lipid-tran... 38 4e-04
AV417442 38 6e-04
TC19050 weakly similar to UP|Q9M6T9 (Q9M6T9) Nonspecific lipid-t... 37 0.001
TC19430 weakly similar to UP|NLT3_HORVU (Q43766) Nonspecific lip... 37 0.001
TC13156 weakly similar to UP|NLT3_HORVU (Q43766) Nonspecific lip... 37 0.001
TC15055 weakly similar to UP|O24101 (O24101) MtN5 protein precur... 36 0.002
AI967923 34 0.008
TC10431 similar to UP|Q7XTF6 (Q7XTF6) OJ991214_12.11 protein, pa... 33 0.014
TC10710 similar to UP|Q9SUZ2 (Q9SUZ2) 5B protein like protein, p... 32 0.041
TC14471 similar to UP|O22656 (O22656) Ubiquitin-conjugating enzy... 30 0.12
TC8033 similar to UP|NLTP_CICAR (O23758) Nonspecific lipid-trans... 29 0.20
TC8034 similar to UP|NLTP_CICAR (O23758) Nonspecific lipid-trans... 29 0.20
TC10327 weakly similar to UP|NLT1_PRUDU (Q43017) Nonspecific lip... 28 0.45
>TC15783 similar to UP|Q9LJ86 (Q9LJ86) Gb|AAD48513.1 (AT3g22600/F16J14_17),
partial (48%)
Length = 578
Score = 58.2 bits (139), Expect = 4e-10
Identities = 28/105 (26%), Positives = 49/105 (46%)
Frame = -2
Query: 12 LCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPEC 71
LC++ +++ S C S + + PCL++ TG + +P CC +S+ + P+C
Sbjct: 421 LCLVMVMVATMWTQNAAQSGCTSALTSLSPCLNYITGSSSSPSPSCCSQLSSVVQSSPQC 242
Query: 72 LCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCPSK 116
LC ++ G S + + + L LP C V +S C K
Sbjct: 241 LCSLL----NGGGSSFGITMNQTLALSLPGPCKVQTPPVSQCKGK 119
>TC17770 weakly similar to UP|Q8VYI9 (Q8VYI9) AT5g64080/MHJ24_6, partial
(42%)
Length = 554
Score = 50.1 bits (118), Expect = 1e-07
Identities = 30/113 (26%), Positives = 52/113 (45%), Gaps = 9/113 (7%)
Frame = +3
Query: 10 MCLCVLALII------GGCNGAEDLASKCGSVVQKVIPCLDFATGKAPT--PKKECCDAA 61
+ LCV+A+ N A ++ C +++ K+ CL F T + P+ +CC
Sbjct: 192 LILCVVAICAVDFAHSASSNNAPAPSADCSNLILKMTDCLSFVTNDSTVTKPQGQCCSGL 371
Query: 62 NSIKATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGAN-ISDC 113
S+ T P CLC + S + + K L LP+ C V+ +N +++C
Sbjct: 372 KSVLKTAPSCLCEAFK-----SSSQFGVVLNATKALALPSACKVSASNSVTNC 515
>TC9144
Length = 835
Score = 44.3 bits (103), Expect = 6e-06
Identities = 25/84 (29%), Positives = 40/84 (46%), Gaps = 2/84 (2%)
Frame = +1
Query: 32 CGSVVQKVIPCLDF--ATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
C + + V CL F A P K CC + ++P CLC ++ G P+ +
Sbjct: 205 CFTALTNVSDCLTFVEAGSNLTKPDKGCCPEFAGLIESNPICLCQLL-----GKPDFVGI 369
Query: 90 GIQEDKLLQLPTVCHVNGANISDC 113
I +K ++LP+VC V+ +S C
Sbjct: 370 KINLNKAIKLPSVCGVDTPPVSTC 441
>TC9112 weakly similar to UP|Q9SI70 (Q9SI70) F23N19.16, partial (53%)
Length = 807
Score = 41.2 bits (95), Expect = 5e-05
Identities = 31/89 (34%), Positives = 39/89 (42%), Gaps = 1/89 (1%)
Frame = +1
Query: 30 SKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPE-SKS 88
S S Q +IPC D T + P CCDA T CLC + SP +S
Sbjct: 160 SSIPSCAQNLIPCADHLT--STNPPASCCDAIKKTVETQLTCLCNLFY-----SPGLLQS 318
Query: 89 MGIQEDKLLQLPTVCHVNGANISDCPSKS 117
I D+ LQL C V +++S C S S
Sbjct: 319 FKISTDQALQLSRRCGVT-SDLSSCKSGS 402
>AI967893
Length = 258
Score = 40.8 bits (94), Expect = 7e-05
Identities = 21/69 (30%), Positives = 37/69 (53%)
Frame = +3
Query: 1 MKNQQHQMFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDA 60
M +++ M + L V+A++ C GA S C +V+ + PCL++ TG + TP CC
Sbjct: 63 MAHRKMNMGLILVVMAML---CAGAA-AQSSCTNVLVSLSPCLNYITGNSSTPSSGCCSQ 230
Query: 61 ANSIKATDP 69
++ + P
Sbjct: 231 LATVVRSQP 257
>TC9111
Length = 534
Score = 39.3 bits (90), Expect = 2e-04
Identities = 30/90 (33%), Positives = 39/90 (43%), Gaps = 1/90 (1%)
Frame = +2
Query: 30 SKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPE-SKS 88
S S Q +I C D+ T + P CCDA T CLC + SP +S
Sbjct: 140 SSIPSCAQNLILCADYLT--STNPPASCCDAIKKTVETQLTCLCNLFY-----SPGLLQS 298
Query: 89 MGIQEDKLLQLPTVCHVNGANISDCPSKSK 118
I D+ LQL C V +++S C S K
Sbjct: 299 FKISTDQALQLSRRCGVT-SDLSSCKSGKK 385
>TC11986 weakly similar to UP|Q8RYA8 (Q8RYA8) Lipid transfer precursor
protein (Nonspecific lipid-transfer protein) (LTP),
partial (77%)
Length = 468
Score = 38.5 bits (88), Expect = 3e-04
Identities = 26/103 (25%), Positives = 42/103 (40%), Gaps = 5/103 (4%)
Frame = +2
Query: 9 FMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIK--- 65
F L V+ +++G + + A CG V V PC+ + P+ CC+ ++K
Sbjct: 92 FASLVVMCMVLG--SPFAEAALPCGQVQFTVAPCIGYLRSPGPSVPAPCCNGIRTLKNQA 265
Query: 66 --ATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
D + C ++ T P G+ L LP C VN
Sbjct: 266 KTIADRQGACRCLKSTSLSLP-----GLNLPALASLPGKCGVN 379
>TC14628 similar to UP|NLTP_CICAR (O23758) Nonspecific lipid-transfer
protein precursor (LTP), partial (91%)
Length = 604
Score = 38.1 bits (87), Expect = 4e-04
Identities = 25/103 (24%), Positives = 39/103 (37%), Gaps = 5/103 (4%)
Frame = +1
Query: 9 FMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKAT- 67
F C+ V + + A CGSV+ + PC+ + TG P P +CC ++ A
Sbjct: 94 FSCMVVAICMALVAAPMAEAAISCGSVIGALTPCMPYLTG-GPAPSPQCCAGVKNLNAAA 270
Query: 68 ----DPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
D + C ++ P + LP C VN
Sbjct: 271 STTPDRKTACGCLKNAAGSMP-----NLNAGNAAALPGKCGVN 384
>AV417442
Length = 391
Score = 37.7 bits (86), Expect = 6e-04
Identities = 19/79 (24%), Positives = 35/79 (44%), Gaps = 5/79 (6%)
Frame = +3
Query: 3 NQQHQMFMCLCVLALIIGGCNGAEDLASKCG--SVVQKVIPCLDFAT---GKAPTPKKEC 57
N + + + L + I G + + C ++ PC+ F T G +P EC
Sbjct: 87 NMERFVPLALVLAMAIFMAAPGYAQINTPCNVSAISTSFTPCMSFLTNSSGNGTSPTAEC 266
Query: 58 CDAANSIKATDPECLCYII 76
CD+ S+ + +CLC ++
Sbjct: 267 CDSIKSLTSGGRDCLCLVV 323
>TC19050 weakly similar to UP|Q9M6T9 (Q9M6T9) Nonspecific lipid-transfer
protein (LTP), partial (57%)
Length = 579
Score = 37.0 bits (84), Expect = 0.001
Identities = 28/108 (25%), Positives = 43/108 (38%), Gaps = 5/108 (4%)
Frame = +2
Query: 9 FMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSI---- 64
F L V+ +++G + D A CG + V PC+ + PT CC+ I
Sbjct: 50 FASLAVMVMVLG--SPFADAALPCGQLTFTVAPCIGYLRNPTPTVPAACCNGIRDILRQA 223
Query: 65 -KATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANIS 111
D + +C ++ T P GI L +P C GA +S
Sbjct: 224 KTVPDRQGVCRCLKSTVYNLP-----GINLPALASVPGKC---GAKLS 343
>TC19430 weakly similar to UP|NLT3_HORVU (Q43766) Nonspecific lipid-transfer
protein 3 precursor (LTP 3) (CW20) (CW-20) (CW-19),
partial (53%)
Length = 520
Score = 37.0 bits (84), Expect = 0.001
Identities = 24/103 (23%), Positives = 43/103 (41%), Gaps = 5/103 (4%)
Frame = +3
Query: 9 FMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKAT- 67
F L +L L++ A + A C V++ + PC+ + + P CC A ++ +
Sbjct: 39 FSALVLLMLLVS----ASEAAISCSDVIKDLKPCVSYLVSGSGKPPGACCSGAKALASAV 206
Query: 68 ----DPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
D + C I+ T +KS+ + LP C +N
Sbjct: 207 STSEDKKAACNCIKST------AKSIKMNSQLAKALPGNCGIN 317
>TC13156 weakly similar to UP|NLT3_HORVU (Q43766) Nonspecific lipid-transfer
protein 3 precursor (LTP 3) (CW20) (CW-20) (CW-19),
partial (53%)
Length = 603
Score = 36.6 bits (83), Expect = 0.001
Identities = 24/103 (23%), Positives = 43/103 (41%), Gaps = 5/103 (4%)
Frame = +1
Query: 9 FMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKAT- 67
F L +L L++ A + A C V++ + PC+ + + P CC A ++ +
Sbjct: 28 FSALVLLMLLVS----ASEAAISCSDVIKDLKPCVSYLVSGSGKPPVACCSGAKALASAV 195
Query: 68 ----DPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
D + C I+ T +KS+ + LP C +N
Sbjct: 196 STSEDKKAACNCIKST------AKSIKMNSQLAEALPGNCGIN 306
>TC15055 weakly similar to UP|O24101 (O24101) MtN5 protein precursor,
partial (52%)
Length = 664
Score = 36.2 bits (82), Expect = 0.002
Identities = 28/89 (31%), Positives = 40/89 (44%), Gaps = 1/89 (1%)
Frame = +3
Query: 16 ALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGK-APTPKKECCDAANSIKATDPECLCY 74
AL I GA+ +A C ++ C ATG+ P P K+CCD ++ + CLC
Sbjct: 81 ALFIALLGGAQAVAL-CNIDTSQLKSCRAAATGEHPPPPDKKCCDV---VRQANLPCLC- 245
Query: 75 IIQQTHKGSPESKSMGIQEDKLLQLPTVC 103
K S GI + L+LP+ C
Sbjct: 246 ------KYKSALPSFGINPTQALKLPSEC 314
>AI967923
Length = 302
Score = 33.9 bits (76), Expect = 0.008
Identities = 18/67 (26%), Positives = 29/67 (42%), Gaps = 4/67 (5%)
Frame = +3
Query: 41 PCLDFATGK----APTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSMGIQEDKL 96
PCL + + T CCDA ++ A D C C++++ P + +L
Sbjct: 66 PCLPYVSSPPNNLTGTASTACCDAFSAATAVDSLCFCHLLR-----DPVILGFSLNTTRL 230
Query: 97 LQLPTVC 103
L L +VC
Sbjct: 231 LSLSSVC 251
>TC10431 similar to UP|Q7XTF6 (Q7XTF6) OJ991214_12.11 protein, partial (38%)
Length = 582
Score = 33.1 bits (74), Expect = 0.014
Identities = 21/90 (23%), Positives = 38/90 (41%), Gaps = 6/90 (6%)
Frame = +2
Query: 27 DLASKCG------SVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTH 80
+ A +CG + K+IPC A + + + CC I +P CLC ++
Sbjct: 122 EAAGECGRSTTPDNEAMKLIPCASAAQDENASVSQSCCAQVQKI-GKNPSCLCAVVL--- 289
Query: 81 KGSPESKSMGIQEDKLLQLPTVCHVNGANI 110
S +K G+ + +P C+++ I
Sbjct: 290 --SNMAKMSGVNPKIAITIPKRCNLDNRPI 373
>TC10710 similar to UP|Q9SUZ2 (Q9SUZ2) 5B protein like protein, partial
(58%)
Length = 730
Score = 31.6 bits (70), Expect = 0.041
Identities = 20/70 (28%), Positives = 31/70 (43%)
Frame = +1
Query: 34 SVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSMGIQE 93
S + ++IPC +P P E C N++K+ CLC ++ G P S G+
Sbjct: 4 SALVQLIPCRAAVAPFSPIPPNEAC--CNALKSLGQPCLCVLV----NGPPIS---GVDR 156
Query: 94 DKLLQLPTVC 103
+ LP C
Sbjct: 157 NMASLLPEKC 186
>TC14471 similar to UP|O22656 (O22656) Ubiquitin-conjugating enzyme protein
E2 (Ubiquitin-conjugating enzyme E2) (Ubiquitin-protein
ligase) (Ubiquitin carrier protein) , partial (97%)
Length = 1023
Score = 30.0 bits (66), Expect = 0.12
Identities = 19/65 (29%), Positives = 29/65 (44%)
Frame = -2
Query: 36 VQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSMGIQEDK 95
+Q V PC D G+ K+ CC ++N +T Y+ HK + SK + +
Sbjct: 737 IQLVFPC*DKYHGEK---KEACCRSSNGSLSTRINSNGYVSPSLHKSTMRSKPASYES*Q 567
Query: 96 LLQLP 100
L LP
Sbjct: 566 FL*LP 552
>TC8033 similar to UP|NLTP_CICAR (O23758) Nonspecific lipid-transfer
protein precursor (LTP), partial (85%)
Length = 707
Score = 29.3 bits (64), Expect = 0.20
Identities = 25/104 (24%), Positives = 38/104 (36%), Gaps = 6/104 (5%)
Frame = +3
Query: 9 FMC-LCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKA- 66
F+C L VL++++ A A CG++ +IPC + G A P CC + +
Sbjct: 87 FVCALVVLSMVVAPMAEA---AITCGAIASALIPCTAYIQGGA-GPSPPCCAGVRRVNSA 254
Query: 67 ----TDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
D C ++ P LP C VN
Sbjct: 255 ASSTADRRAACNCLKSAAGSFPR-----FNAGNAASLPGKCGVN 371
>TC8034 similar to UP|NLTP_CICAR (O23758) Nonspecific lipid-transfer
protein precursor (LTP), partial (84%)
Length = 742
Score = 29.3 bits (64), Expect = 0.20
Identities = 25/104 (24%), Positives = 38/104 (36%), Gaps = 6/104 (5%)
Frame = +1
Query: 9 FMC-LCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKA- 66
F+C L VL++++ A A CG++ +IPC + G A P CC + +
Sbjct: 55 FVCALVVLSMVVAPMAEA---AITCGAIASALIPCTAYIQGGA-GPSPPCCAGVRRVNSA 222
Query: 67 ----TDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
D C ++ P LP C VN
Sbjct: 223 ASSTADRRAACNCLKSAAGSFPR-----FNAGNAASLPGKCGVN 339
>TC10327 weakly similar to UP|NLT1_PRUDU (Q43017) Nonspecific lipid-transfer
protein 1 precursor (LTP 1), partial (52%)
Length = 683
Score = 28.1 bits (61), Expect = 0.45
Identities = 20/83 (24%), Positives = 31/83 (37%), Gaps = 5/83 (6%)
Frame = +1
Query: 29 ASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSI-----KATDPECLCYIIQQTHKGS 83
A CG VV + PC+ + +CC+ ++ D +C I+Q S
Sbjct: 157 AVTCGQVVNSLYPCVSYLMNGGNMVPGQCCNGIRNLYGMATNTPDRRAVCTCIKQAVSNS 336
Query: 84 PESKSMGIQEDKLLQLPTVCHVN 106
S + + LP C VN
Sbjct: 337 GFSFT-SYNLNLAAGLPKKCGVN 402
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.134 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,569,490
Number of Sequences: 28460
Number of extensions: 34637
Number of successful extensions: 190
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 189
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 189
length of query: 123
length of database: 4,897,600
effective HSP length: 79
effective length of query: 44
effective length of database: 2,649,260
effective search space: 116567440
effective search space used: 116567440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 49 (23.5 bits)
Medicago: description of AC147538.3


