

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147496.7 + phase: 0
(110 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
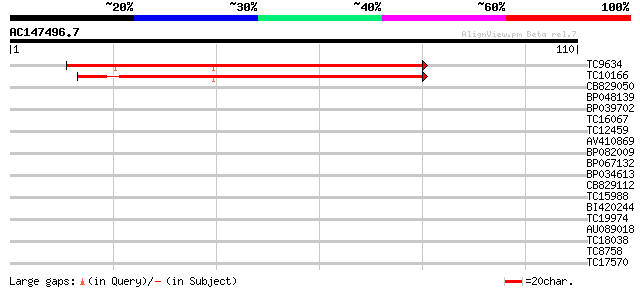
TC9634 similar to UP|Q8W0Z3 (Q8W0Z3) At1g29350/F15D2_27, partial... 67 6e-13
TC10166 weakly similar to UP|Q8W0T8 (Q8W0T8) Gb protein, partial... 65 2e-12
CB829050 26 0.86
BP048139 26 0.86
BP039702 25 1.5
TC16067 UP|Q9ZRY4 (Q9ZRY4) NDX1 homeobox protein, complete 25 1.5
TC12459 25 1.9
AV410869 25 1.9
BP082009 24 3.3
BP067132 24 3.3
BP034613 24 4.3
CB829112 24 4.3
TC15988 similar to PIR|T51353|T51353 NAD ADP-ribosyltransferase ... 24 4.3
BI420244 23 5.6
TC19974 weakly similar to UP|O48721 (O48721) Expressed protein, ... 23 5.6
AU089018 23 7.3
TC18038 similar to GB|CAC04249.1|9886727|PSA297606 PPF-1 protein... 23 7.3
TC8758 similar to GB|AAC77862.2|20197419|AC005623 Argonaute (AGO... 23 7.3
TC17570 similar to UP|O48933 (O48933) Histidinol dehydrogenase (... 23 7.3
>TC9634 similar to UP|Q8W0Z3 (Q8W0Z3) At1g29350/F15D2_27, partial (7%)
Length = 687
Score = 66.6 bits (161), Expect = 6e-13
Identities = 36/72 (50%), Positives = 50/72 (69%), Gaps = 2/72 (2%)
Frame = -3
Query: 12 GGGGGVKA-VKEVVTTAKNVVAAVKEIV-NCTEQEIYDVLRECDMDPNLAVEKLLSQDTF 69
G G KA + + ++ +V ++KEIV N + EIY L++C+MDPN AV +LLSQD F
Sbjct: 514 GAAGHNKASLAGIPPASRKMVQSLKEIVSNIPDNEIYATLKDCNMDPNEAVSRLLSQDPF 335
Query: 70 REVRSKREKRKE 81
EV+SKREK+KE
Sbjct: 334 HEVKSKREKKKE 299
>TC10166 weakly similar to UP|Q8W0T8 (Q8W0T8) Gb protein, partial (5%)
Length = 980
Score = 65.1 bits (157), Expect = 2e-12
Identities = 32/69 (46%), Positives = 47/69 (67%), Gaps = 1/69 (1%)
Frame = +3
Query: 14 GGGVKAVKEVVTTAKNVVAAVKEIV-NCTEQEIYDVLRECDMDPNLAVEKLLSQDTFREV 72
GGG +A + + K + +KEI N ++++IY +L+EC MDPN +KLL QDTF EV
Sbjct: 276 GGGFRA--SIPNSVKKTIQNIKEITGNHSDEDIYAMLKECSMDPNETTQKLLLQDTFHEV 449
Query: 73 RSKREKRKE 81
+ K++KRKE
Sbjct: 450 KRKKDKRKE 476
>CB829050
Length = 559
Score = 26.2 bits (56), Expect = 0.86
Identities = 12/29 (41%), Positives = 18/29 (61%)
Frame = +1
Query: 2 GSESVVNRNSGGGGGVKAVKEVVTTAKNV 30
G++SVV+ +GGG V+EV+ NV
Sbjct: 373 GNQSVVSLEAGGGSRTHQVEEVLEALGNV 459
>BP048139
Length = 567
Score = 26.2 bits (56), Expect = 0.86
Identities = 14/45 (31%), Positives = 27/45 (59%), Gaps = 2/45 (4%)
Frame = +3
Query: 63 LLSQDTFREVRSKREKRKEVIL--LRYVNIDMRLNLSLMMNNVCM 105
L++Q ++++V K ++RK I+ RY+ + +N L N VC+
Sbjct: 408 LMTQYSWKKVLQKSKQRKRYIIKSFRYIIKKL*INKILRKNKVCI 542
>BP039702
Length = 553
Score = 25.4 bits (54), Expect = 1.5
Identities = 11/44 (25%), Positives = 24/44 (54%)
Frame = -2
Query: 62 KLLSQDTFREVRSKREKRKEVILLRYVNIDMRLNLSLMMNNVCM 105
+L + R V S +R+ +L ++++ + +LSL + VC+
Sbjct: 132 RLSCRGILRRVLSP*RRRRPALLCLFISLSL*FSLSLFLYRVCL 1
>TC16067 UP|Q9ZRY4 (Q9ZRY4) NDX1 homeobox protein, complete
Length = 3271
Score = 25.4 bits (54), Expect = 1.5
Identities = 10/25 (40%), Positives = 18/25 (72%)
Frame = +2
Query: 29 NVVAAVKEIVNCTEQEIYDVLRECD 53
++V+AVKE+ QE+Y +LR+ +
Sbjct: 149 DLVSAVKELHGLNSQELYRLLRDAE 223
>TC12459
Length = 797
Score = 25.0 bits (53), Expect = 1.9
Identities = 11/43 (25%), Positives = 19/43 (43%)
Frame = +3
Query: 2 GSESVVNRNSGGGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQE 44
GS GGGGG + EVV + + V+ + + ++
Sbjct: 108 GSSDEEGEGGGGGGGGEGRTEVVVVDETAIGEVRGVAGVSARQ 236
>AV410869
Length = 416
Score = 25.0 bits (53), Expect = 1.9
Identities = 12/40 (30%), Positives = 20/40 (50%)
Frame = -1
Query: 11 SGGGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLR 50
+GGGGG K+ ++ V A C + +YD++R
Sbjct: 386 TGGGGGFKSATLLIAVGAGVAA-------CQVRALYDLVR 288
>BP082009
Length = 110
Score = 24.3 bits (51), Expect = 3.3
Identities = 8/16 (50%), Positives = 15/16 (93%)
Frame = +1
Query: 82 VILLRYVNIDMRLNLS 97
+IL++++NI+ +LNLS
Sbjct: 52 IILMKFINIEEKLNLS 99
>BP067132
Length = 499
Score = 24.3 bits (51), Expect = 3.3
Identities = 8/25 (32%), Positives = 18/25 (72%)
Frame = +1
Query: 86 RYVNIDMRLNLSLMMNNVCMYFDFR 110
+ +++ L+LSL +N +C +F+F+
Sbjct: 28 KLLSLSPSLSLSLSLNKLCXWFNFQ 102
>BP034613
Length = 524
Score = 23.9 bits (50), Expect = 4.3
Identities = 11/30 (36%), Positives = 16/30 (52%)
Frame = -1
Query: 3 SESVVNRNSGGGGGVKAVKEVVTTAKNVVA 32
+E ++ SGGGGG V E + A+ A
Sbjct: 191 TEFRISNTSGGGGGANVVPEGLEAAEGARA 102
>CB829112
Length = 234
Score = 23.9 bits (50), Expect = 4.3
Identities = 15/37 (40%), Positives = 17/37 (45%)
Frame = -3
Query: 2 GSESVVNRNSGGGGGVKAVKEVVTTAKNVVAAVKEIV 38
G S R SGGGGG E T A V +E+V
Sbjct: 118 GGSSGGGRRSGGGGGASGASE--TNAYEVAR*GEELV 14
>TC15988 similar to PIR|T51353|T51353 NAD ADP-ribosyltransferase [imported]
- Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (11%)
Length = 612
Score = 23.9 bits (50), Expect = 4.3
Identities = 11/50 (22%), Positives = 28/50 (56%)
Frame = -1
Query: 61 EKLLSQDTFREVRSKREKRKEVILLRYVNIDMRLNLSLMMNNVCMYFDFR 110
+KLL + ++ + +++ ++ ++ L+LS ++NN+ M +FR
Sbjct: 375 KKLLRTHKAPKQKTSSFQTSPLVMEPHLQ*ELHLDLSSIINNILMVHEFR 226
>BI420244
Length = 571
Score = 23.5 bits (49), Expect = 5.6
Identities = 10/19 (52%), Positives = 11/19 (57%)
Frame = -1
Query: 4 ESVVNRNSGGGGGVKAVKE 22
E V GGGGGV A +E
Sbjct: 115 EEVEEEEEGGGGGVSAEEE 59
>TC19974 weakly similar to UP|O48721 (O48721) Expressed protein, partial
(29%)
Length = 444
Score = 23.5 bits (49), Expect = 5.6
Identities = 19/56 (33%), Positives = 26/56 (45%), Gaps = 5/56 (8%)
Frame = +2
Query: 19 AVKEVVT-----TAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQDTF 69
A K VVT AK + A K +NC E + + R D+D + LS +TF
Sbjct: 170 APKVVVTRERGKNAKLIAALAKHEINCLELTLIEHTRGPDLD---RLPCALSDNTF 328
>AU089018
Length = 512
Score = 23.1 bits (48), Expect = 7.3
Identities = 10/20 (50%), Positives = 13/20 (65%)
Frame = -2
Query: 11 SGGGGGVKAVKEVVTTAKNV 30
SGGGGGV V+ + T K +
Sbjct: 100 SGGGGGVFTVRRLSFTQKRM 41
>TC18038 similar to GB|CAC04249.1|9886727|PSA297606 PPF-1 protein {Pisum
sativum;} , partial (20%)
Length = 586
Score = 23.1 bits (48), Expect = 7.3
Identities = 13/42 (30%), Positives = 20/42 (46%)
Frame = +3
Query: 66 QDTFREVRSKREKRKEVILLRYVNIDMRLNLSLMMNNVCMYF 107
Q RE RSKR KRK + ++ + + L ++ C F
Sbjct: 258 QSFSRERRSKRSKRKRAV*KYHIRLFVAYRGVLYISTDCPIF 383
>TC8758 similar to GB|AAC77862.2|20197419|AC005623 Argonaute (AGO1)-like
protein {Arabidopsis thaliana;} , partial (32%)
Length = 1142
Score = 23.1 bits (48), Expect = 7.3
Identities = 9/18 (50%), Positives = 11/18 (61%)
Frame = -2
Query: 2 GSESVVNRNSGGGGGVKA 19
GS S GGGGG+K+
Sbjct: 157 GSTSAGTMGGGGGGGIKS 104
>TC17570 similar to UP|O48933 (O48933) Histidinol dehydrogenase
(Fragment), partial (51%)
Length = 811
Score = 23.1 bits (48), Expect = 7.3
Identities = 8/18 (44%), Positives = 12/18 (66%)
Frame = -1
Query: 12 GGGGGVKAVKEVVTTAKN 29
GGGGGV+ +KE ++
Sbjct: 70 GGGGGVEGMKEATVPVRS 17
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.135 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,352,023
Number of Sequences: 28460
Number of extensions: 11921
Number of successful extensions: 112
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 111
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 112
length of query: 110
length of database: 4,897,600
effective HSP length: 86
effective length of query: 24
effective length of database: 2,450,040
effective search space: 58800960
effective search space used: 58800960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 47 (22.7 bits)
Medicago: description of AC147496.7


