

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147496.2 - phase: 0 /pseudo
(550 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
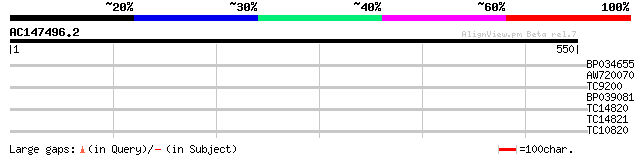
Score E
Sequences producing significant alignments: (bits) Value
BP034655 33 0.091
AW720070 32 0.20
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 28 3.8
BP039081 28 5.0
TC14820 homologue to UP|HS80_LYCES (P36181) Heat shock cognate p... 28 5.0
TC14821 homologue to UP|AAR12195 (AAR12195) Molecular chaperone ... 28 5.0
TC10820 weakly similar to UP|Q9M9Y0 (Q9M9Y0) F4H5.19 protein, pa... 27 6.6
>BP034655
Length = 517
Score = 33.5 bits (75), Expect = 0.091
Identities = 19/67 (28%), Positives = 35/67 (51%), Gaps = 3/67 (4%)
Frame = +2
Query: 459 LSPSEVRDDQKKRKEKYEKEKRKIKRKEREFA*KKK---KRRVSRNSHEQTAPIFTTLQK 515
+SP ++ D K K +K+K+K K+K+++ KKK KR + + + T P+ +
Sbjct: 107 ISPPQILDSPTTMKRKKKKKKKKKKKKKKKKKKKKKKKTKRMMKKTKMKTTFPLHRRIGF 286
Query: 516 VALLTNT 522
+ T T
Sbjct: 287 LPTFTRT 307
>AW720070
Length = 484
Score = 32.3 bits (72), Expect = 0.20
Identities = 17/58 (29%), Positives = 34/58 (58%)
Frame = +2
Query: 467 DQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQTAPIFTTLQKVALLTNTQN 524
++K++K+K +++K+K +KE +KKK++ + + TAP L + LL Q+
Sbjct: 107 EKKRKKKKKKRKKKKQVKKESSQQQEKKKKKKKKAAVSTTAPFLHPLISLYLLLLPQS 280
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 28.1 bits (61), Expect = 3.8
Identities = 12/33 (36%), Positives = 23/33 (69%)
Frame = +2
Query: 465 RDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRR 497
+ +KK ++K+ ++K++ KRK R KKKK++
Sbjct: 200 KKSKKK*RKKWRRKKKRRKRKRRVKKKKKKKKK 298
>BP039081
Length = 543
Score = 27.7 bits (60), Expect = 5.0
Identities = 12/43 (27%), Positives = 21/43 (47%)
Frame = +3
Query: 297 VENHEGYVEEEAEEIPSGDLFMIRRFLGNQAKEEESNQRETLF 339
V NH G+ ++E + F R+ G KEEE + ++ +
Sbjct: 12 VPNHTGFNKDECGTVDEDGRFCSNRWKGYNEKEEEGSPQDNYY 140
>TC14820 homologue to UP|HS80_LYCES (P36181) Heat shock cognate protein 80,
partial (53%)
Length = 1180
Score = 27.7 bits (60), Expect = 5.0
Identities = 13/40 (32%), Positives = 23/40 (57%)
Frame = +1
Query: 466 DDQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQ 505
+D + KEK EK+K+KIK E++ K++ + E+
Sbjct: 778 EDVDEEKEKEEKKKKKIKEVSHEWSLVNKQKPIWMRKPEE 897
>TC14821 homologue to UP|AAR12195 (AAR12195) Molecular chaperone Hsp90-1,
partial (29%)
Length = 612
Score = 27.7 bits (60), Expect = 5.0
Identities = 13/40 (32%), Positives = 23/40 (57%)
Frame = +3
Query: 466 DDQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQ 505
+D + KEK EK+K+KIK E++ K++ + E+
Sbjct: 117 EDVDEEKEKEEKKKKKIKEVSHEWSLVNKQKPIWMRKPEE 236
>TC10820 weakly similar to UP|Q9M9Y0 (Q9M9Y0) F4H5.19 protein, partial (7%)
Length = 619
Score = 27.3 bits (59), Expect = 6.6
Identities = 19/43 (44%), Positives = 23/43 (53%), Gaps = 2/43 (4%)
Frame = +2
Query: 466 DDQKKRKEKYEKEKRKI--KRKEREFA*KKKKRRVSRNSHEQT 506
D+ KKRK K E EKRK K +E KK+RR R +T
Sbjct: 119 DEMKKRKLK-ENEKRKAHEAEKAKEEQLSKKRRREERREKYRT 244
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.358 0.160 0.569
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,732,595
Number of Sequences: 28460
Number of extensions: 112593
Number of successful extensions: 1203
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 897
Number of HSP's successfully gapped in prelim test: 24
Number of HSP's that attempted gapping in prelim test: 282
Number of HSP's gapped (non-prelim): 941
length of query: 550
length of database: 4,897,600
effective HSP length: 95
effective length of query: 455
effective length of database: 2,193,900
effective search space: 998224500
effective search space used: 998224500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC147496.2


