

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
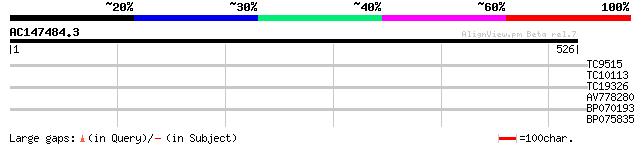
Query= AC147484.3 - phase: 0 /pseudo
(526 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC9515 similar to UP|O49081 (O49081) Clp-like energy-dependent p... 31 0.44
TC10113 UP|RR11_LOTJA (Q9BBQ3) Chloroplast 30S ribosomal protein... 31 0.57
TC19326 weakly similar to UP|Q9ASX8 (Q9ASX8) At1g27760/T22C5_5, ... 30 0.97
AV778280 29 2.2
BP070193 28 2.8
BP075835 28 4.8
>TC9515 similar to UP|O49081 (O49081) Clp-like energy-dependent protease
(ATP-dependent Clp protease proteolytic subunit)
(Endopeptidase Clp) , partial (74%)
Length = 1158
Score = 31.2 bits (69), Expect = 0.44
Identities = 28/94 (29%), Positives = 40/94 (41%), Gaps = 7/94 (7%)
Frame = -3
Query: 116 PSVCREFVARVSDRVSSGALGWSSRQLHSLHCLGVLLNCEKEDDLHVHIKDVYSLISVLV 175
PS+C +R++ R ++ LH LGVL + D H+H SL L
Sbjct: 769 PSLCTVAPSRLAVR*KHYTSCQFYGEIAQLHSLGVL---HPQHDPHIHSGVDVSLCGNLA 599
Query: 176 TGLQLPSEEIRGEVLF-------VLYKLFALHST 202
PS +GEVL +LY+ +AL T
Sbjct: 598 LNSAFPSPHQQGEVLLGLPLLRNILYQPWALSKT 497
>TC10113 UP|RR11_LOTJA (Q9BBQ3) Chloroplast 30S ribosomal protein S11,
complete
Length = 879
Score = 30.8 bits (68), Expect = 0.57
Identities = 22/57 (38%), Positives = 30/57 (52%), Gaps = 2/57 (3%)
Frame = +3
Query: 419 LDLSKS--IEEAVKQAILACLYVSERNINQILQCFYLLKEAYAYSHDENSTHISKLE 473
L+LSK I + + IL Y RNI I Q FY +K YS+ N+ +I K+E
Sbjct: 186 LNLSKKNYILYKI*EYILYYKYYRIRNIIYIFQKFYSIKNCILYSYK*NTINIYKIE 356
>TC19326 weakly similar to UP|Q9ASX8 (Q9ASX8) At1g27760/T22C5_5, partial
(9%)
Length = 677
Score = 30.0 bits (66), Expect = 0.97
Identities = 23/65 (35%), Positives = 32/65 (48%)
Frame = +2
Query: 22 HRSSLNLQTNQGGTICLVCFSNLISNPSSPTIHVSYALSQLSRSLSIPQFLHSIVTFHPH 81
H +LNL NQ FS+ PS + V+Y + LS+PQFL V+ + H
Sbjct: 77 HHLNLNLNLNQTKCCR*SPFSSRFLYPSILLLPVAYFFILVQFRLSLPQFLIR-VSNNAH 253
Query: 82 FLVSP 86
+VSP
Sbjct: 254 NIVSP 268
>AV778280
Length = 555
Score = 28.9 bits (63), Expect = 2.2
Identities = 18/56 (32%), Positives = 28/56 (49%), Gaps = 4/56 (7%)
Frame = -2
Query: 430 KQAILACLYVSERNINQILQCFY----LLKEAYAYSHDENSTHISKLELRSGILDI 481
K++ +CL + + +I IL C Y L + YS D+ IS L L+ +L I
Sbjct: 473 KKSGKSCLCIMQCSIGDILTCSYELGVFLCQKLLYSSDKYIMEISCLNLKKSVLSI 306
>BP070193
Length = 476
Score = 28.5 bits (62), Expect = 2.8
Identities = 15/49 (30%), Positives = 25/49 (50%), Gaps = 9/49 (18%)
Frame = +2
Query: 47 NPSSPTIHVSYALSQLSRSLSIPQFLH---------SIVTFHPHFLVSP 86
NPS +H+S+ L+ + SL+ P +H S++T P F +P
Sbjct: 134 NPSCHLLHLSHGLAATTTSLASPPHIHGRAATTTSFSLLTSFPSFSTAP 280
>BP075835
Length = 379
Score = 27.7 bits (60), Expect = 4.8
Identities = 14/40 (35%), Positives = 23/40 (57%)
Frame = -3
Query: 434 LACLYVSERNINQILQCFYLLKEAYAYSHDENSTHISKLE 473
+A + S++NI ++C Y+L YA+SHD H + E
Sbjct: 308 VATFHGSQKNILVRVKC*YILILTYAHSHDWYPCHSNSHE 189
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.137 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,824,935
Number of Sequences: 28460
Number of extensions: 146693
Number of successful extensions: 842
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 839
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 842
length of query: 526
length of database: 4,897,600
effective HSP length: 94
effective length of query: 432
effective length of database: 2,222,360
effective search space: 960059520
effective search space used: 960059520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC147484.3


