

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
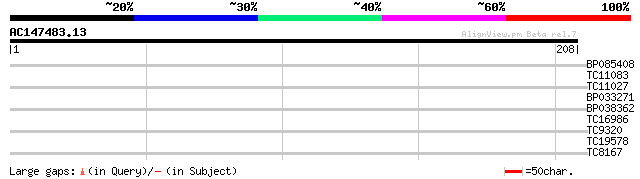
Query= AC147483.13 - phase: 0
(208 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP085408 27 2.0
TC11083 weakly similar to UP|AAQ82687 (AAQ82687) Epa5p, partial ... 27 3.3
TC11027 homologue to UP|Q39828 (Q39828) SDL5A, partial (44%) 26 4.4
BP033271 26 5.7
BP038362 26 5.7
TC16986 similar to UP|AAR86687 (AAR86687) Glycerol kinase , par... 26 5.7
TC9320 similar to UP|IF3B_ORYSA (Q94HF1) Eukaryotic translation ... 25 7.5
TC19578 similar to UP|Q8LPN7 (Q8LPN7) AT3g19950/MPN9_19, partial... 25 9.7
TC8167 similar to PIR|JC7519|JC7519 subtilisin-like serine prote... 25 9.7
>BP085408
Length = 350
Score = 27.3 bits (59), Expect = 2.0
Identities = 15/48 (31%), Positives = 23/48 (47%)
Frame = +3
Query: 63 ILRTNVEEKLVHTLVQFYDPSFRCFTFPDFQVVPTLEAYSHLLDSPTA 110
I++TN ++KL+H + P +V+PT EA H P A
Sbjct: 117 IIKTNGQDKLIHNTRELCAPHLL*LPPHHLRVMPTWEARCHAGVGPVA 260
>TC11083 weakly similar to UP|AAQ82687 (AAQ82687) Epa5p, partial (5%)
Length = 518
Score = 26.6 bits (57), Expect = 3.3
Identities = 11/25 (44%), Positives = 16/25 (64%)
Frame = -3
Query: 7 CIHRRIGQIKNFVNITGMKKTKSYK 31
C+H+ I QIK+ N +K+T YK
Sbjct: 444 CLHKNIHQIKSDENTAKIKRTN*YK 370
>TC11027 homologue to UP|Q39828 (Q39828) SDL5A, partial (44%)
Length = 826
Score = 26.2 bits (56), Expect = 4.4
Identities = 14/40 (35%), Positives = 24/40 (60%)
Frame = -2
Query: 122 LTPLVVAKDLYLKTSDVSNHLTTKSHIRGFTSKYLLEQAT 161
L P +++L +K D+ N +T++SH KYL++ AT
Sbjct: 540 LQPFQSSRELIVK--DIINFITSRSHAIKMIFKYLIK*AT 427
>BP033271
Length = 541
Score = 25.8 bits (55), Expect = 5.7
Identities = 16/37 (43%), Positives = 18/37 (48%)
Frame = +1
Query: 122 LTPLVVAKDLYLKTSDVSNHLTTKSHIRGFTSKYLLE 158
LTP L L+TS L T+ IRGFT L E
Sbjct: 184 LTPTXFFTPLTLRTSHHRTVLQTRRRIRGFTVCILTE 294
>BP038362
Length = 274
Score = 25.8 bits (55), Expect = 5.7
Identities = 12/36 (33%), Positives = 20/36 (55%)
Frame = -3
Query: 92 FQVVPTLEAYSHLLDSPTAEKTPFTGPGTSLTPLVV 127
F PT+EA+S+L + ++ P LTPL++
Sbjct: 209 FPFSPTVEAFSYLGHGTSPWESSHPSPTLDLTPLIL 102
>TC16986 similar to UP|AAR86687 (AAR86687) Glycerol kinase , partial (24%)
Length = 491
Score = 25.8 bits (55), Expect = 5.7
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = +3
Query: 130 DLYLKTSDVSNHLTTKSHIRG 150
DL LKT +N++ TK H+ G
Sbjct: 45 DLNLKTQSNNNNIETKKHVEG 107
>TC9320 similar to UP|IF3B_ORYSA (Q94HF1) Eukaryotic translation initiation
factor 3 subunit 11 (eIF-3 p25) (eIF3k), partial (97%)
Length = 1086
Score = 25.4 bits (54), Expect = 7.5
Identities = 13/31 (41%), Positives = 19/31 (60%)
Frame = +1
Query: 158 EQATCQDALEAILALLIYGLILFPSLDNFVD 188
EQA A+E +LAL Y + P L+N+V+
Sbjct: 94 EQAQVGYAVEQLLALNPYNPDILPDLENYVN 186
>TC19578 similar to UP|Q8LPN7 (Q8LPN7) AT3g19950/MPN9_19, partial (16%)
Length = 507
Score = 25.0 bits (53), Expect = 9.7
Identities = 9/24 (37%), Positives = 14/24 (57%)
Frame = +3
Query: 93 QVVPTLEAYSHLLDSPTAEKTPFT 116
Q++PTL SH + +PT P +
Sbjct: 162 QIIPTLTLTSHCISAPTLPPIPIS 233
>TC8167 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase -
Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (65%)
Length = 2760
Score = 25.0 bits (53), Expect = 9.7
Identities = 23/73 (31%), Positives = 34/73 (46%)
Frame = -3
Query: 75 TLVQFYDPSFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTGPGTSLTPLVVAKDLYLK 134
T+ F P+F +F + V+PT+E S T T P +P +V +YL
Sbjct: 2272 TVYLFLSPTFPNSSFVGYTVIPTMEGPS--------TTTS*TEPA---SPTLVTVLVYLT 2126
Query: 135 TSDVSNHLTTKSH 147
T SN TT+S+
Sbjct: 2125 TFFASN--TTESN 2093
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.139 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,330,065
Number of Sequences: 28460
Number of extensions: 40488
Number of successful extensions: 201
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 200
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 201
length of query: 208
length of database: 4,897,600
effective HSP length: 86
effective length of query: 122
effective length of database: 2,450,040
effective search space: 298904880
effective search space used: 298904880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 53 (25.0 bits)
Medicago: description of AC147483.13


