

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
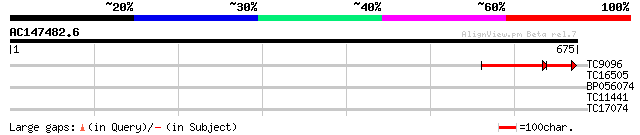
Query= AC147482.6 + phase: 2 /pseudo
(675 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC9096 similar to UP|Q9LD60 (Q9LD60) Spliceosomal-like protein, ... 113 1e-38
TC16505 similar to GB|AAN64529.1|24797034|BT001138 At5g58299/At5... 30 1.7
BP056074 28 6.3
TC11441 similar to PIR|S54157|S54157 extensin-like protein - cow... 27 8.2
TC17074 27 8.2
>TC9096 similar to UP|Q9LD60 (Q9LD60) Spliceosomal-like protein, partial
(10%)
Length = 718
Score = 113 bits (283), Expect(2) = 1e-38
Identities = 60/79 (75%), Positives = 61/79 (76%)
Frame = +3
Query: 562 NFMLVM*SHACKRPLLYQVVESAF*MEQLWEVLEHYMLLLHEMMLISSLIWRCI*GRIIH 621
N MLVM S ACKR LLY VVES MEQLWEVL H MLLLH MMLIS LIWRCI* R IH
Sbjct: 6 NLMLVMWSQACKRHLLYLVVESVLYMEQLWEVLGHCMLLLHVMMLISFLIWRCI*CRTIH 185
Query: 622 LCVEEIIWLIDPPIFLLSL 640
LCVEEI WLID PIF L +
Sbjct: 186 LCVEEITWLIDLPIFPLRM 242
Score = 63.9 bits (154), Expect(2) = 1e-38
Identities = 30/35 (85%), Positives = 32/35 (90%)
Frame = +2
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQFPTLPMDLQRKIADELDRT EIL+KLEE +
Sbjct: 257 LCEQFPTLPMDLQRKIADELDRTPAEILRKLEEVR 361
>TC16505 similar to GB|AAN64529.1|24797034|BT001138 At5g58299/At5g58299
{Arabidopsis thaliana;}, partial (16%)
Length = 649
Score = 29.6 bits (65), Expect = 1.7
Identities = 13/34 (38%), Positives = 20/34 (58%)
Frame = +3
Query: 607 ISSLIWRCI*GRIIHLCVEEIIWLIDPPIFLLSL 640
++S W+C+ GR++ +IWLI P F L L
Sbjct: 18 LASSFWKCLPGRLLCHLQAAMIWLISPAGFSLLL 119
>BP056074
Length = 485
Score = 27.7 bits (60), Expect = 6.3
Identities = 18/62 (29%), Positives = 29/62 (46%), Gaps = 1/62 (1%)
Frame = -1
Query: 591 WEVLEHYMLLLHEMMLISSLIWRCI*GRIIHLCVEEIIWLID-PPIFLLSLCEQFPTLPM 649
WEV E + L++ + SL W ++ V +I+ I PP F+ + + LP
Sbjct: 248 WEVCESHHLIIFHCLFFFSLCWMSY--SLVKT*VYHLIFFIRLPPSFICTYLRERAYLPD 75
Query: 650 DL 651
DL
Sbjct: 74 DL 69
>TC11441 similar to PIR|S54157|S54157 extensin-like protein - cowpea
(fragment) {Vigna unguiculata;} , partial (6%)
Length = 541
Score = 27.3 bits (59), Expect = 8.2
Identities = 16/67 (23%), Positives = 30/67 (43%)
Frame = -3
Query: 92 HMTRQSESYHWILMIVCRL*VYRVFLQLQNRFFSLKFKHLLAVRMGQITPLAFS*RWFAE 151
H + + HW++M++C +Y + +++ L K L+ M T + RW
Sbjct: 206 H*EEKDDHGHWLVMVLC---LYLLVVEIG----LLALKVLVKEVMMMTTVVTVEKRWLFG 48
Query: 152 WCTI*NC 158
WC + C
Sbjct: 47 WCLMSYC 27
>TC17074
Length = 573
Score = 27.3 bits (59), Expect = 8.2
Identities = 12/32 (37%), Positives = 21/32 (65%)
Frame = +2
Query: 70 LLVWTLLQCPKADKDPVFLQLDHMTRQSESYH 101
L++W++ C K P+ LQ+ H++RQS +H
Sbjct: 11 LVLWSI-SCDK*ATTPMLLQVLHLSRQSYYFH 103
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.360 0.160 0.619
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,762,647
Number of Sequences: 28460
Number of extensions: 254000
Number of successful extensions: 2684
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1978
Number of HSP's successfully gapped in prelim test: 65
Number of HSP's that attempted gapping in prelim test: 657
Number of HSP's gapped (non-prelim): 2114
length of query: 675
length of database: 4,897,600
effective HSP length: 96
effective length of query: 579
effective length of database: 2,165,440
effective search space: 1253789760
effective search space used: 1253789760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC147482.6


