

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
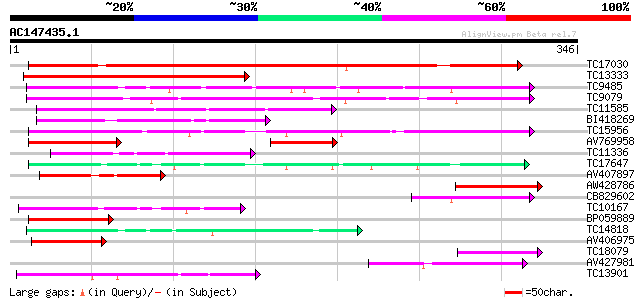
Query= AC147435.1 - phase: 0 /pseudo
(346 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC17030 similar to UP|Q8S995 (Q8S995) Glucosyltransferase-14, pa... 314 1e-86
TC13333 weakly similar to UP|AAR06917 (AAR06917) UDP-glycosyltra... 129 9e-31
TC9485 similar to UP|Q8S999 (Q8S999) Glucosyltransferase-10, par... 103 4e-23
TC9079 weakly similar to UP|Q8W4G1 (Q8W4G1) UDP-glucose glucosyl... 100 5e-22
TC11585 similar to UP|Q9ZWQ5 (Q9ZWQ5) UDP-glycose:flavonoid glyc... 97 3e-21
BI418269 87 4e-18
TC15956 weakly similar to UP|LGT_CITUN (Q9MB73) Limonoid UDP-glu... 76 9e-15
AV769958 75 1e-14
TC11336 similar to UP|Q8S9A5 (Q8S9A5) Glucosyltransferase like p... 70 4e-13
TC17647 weakly similar to UP|AAR06918 (AAR06918) UDP-glycosyltra... 68 3e-12
AV407897 63 8e-11
AW428786 62 1e-10
CB829602 60 5e-10
TC10167 weakly similar to UP|Q9LTA3 (Q9LTA3) Anthocyanidin-3-glu... 57 3e-09
BP059889 56 8e-09
TC14818 weakly similar to UP|Q8S999 (Q8S999) Glucosyltransferase... 52 2e-07
AV406975 48 2e-06
TC18079 weakly similar to UP|UFO5_MANES (Q40287) Flavonol 3-O-gl... 45 2e-05
AV427981 44 4e-05
TC13901 similar to UP|Q9FYU7 (Q9FYU7) UDP-glucose:sinapate gluco... 44 4e-05
>TC17030 similar to UP|Q8S995 (Q8S995) Glucosyltransferase-14, partial (51%)
Length = 1098
Score = 314 bits (805), Expect = 1e-86
Identities = 161/307 (52%), Positives = 209/307 (67%), Gaps = 5/307 (1%)
Frame = +1
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQR-GVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIV 70
HFVLFPL++QGH+IPM+DIA +L+Q+ V VT+ TTP NASRFT S QI+I+
Sbjct: 211 HFVLFPLMSQGHMIPMMDIATILSQQQDVTVTVITTPHNASRFT----HTTKSRSQIRIL 378
Query: 71 TLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFF 130
L FP + GLPEGCEN DM+ S+ + F+A + LQKP E+LF+ LTP P+CIISD
Sbjct: 379 ELEFPHNETGLPEGCENLDMLPSLGTALTFFNAASNLQKPVEQLFNELTPPPNCIISDMC 558
Query: 131 IPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVT 190
+P+T +IA IPRISF G SCF L C+ I K+ + V SE+EYF +PG+PD+I++T
Sbjct: 559 LPYTARIAAKFNIPRISFLGQSCFTLFCLYNIGRHKVRQNVTSETEYFVLPGVPDKIEMT 738
Query: 191 KEQIPGILKGELKE----FGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFV 246
K QIPG + + F + AE SYG ++N+FEELE Y K YKK KNGKVW +
Sbjct: 739 KAQIPGPSGARMDQNRIAFYAETAAAEAASYGVVMNSFEELETEYAKGYKKVKNGKVWCI 918
Query: 247 GPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELAL 306
GPVSL N + + + EH+C+KWLDL + KSV+YACLGS+CNL P QL+EL L
Sbjct: 919 GPVSLSNNNKVSIDE-------DEHYCMKWLDLQKSKSVIYACLGSMCNLTPLQLIELGL 1077
Query: 307 ALEATNR 313
LEA+NR
Sbjct: 1078GLEASNR 1098
>TC13333 weakly similar to UP|AAR06917 (AAR06917) UDP-glycosyltransferase
73E1, partial (10%)
Length = 419
Score = 129 bits (323), Expect = 9e-31
Identities = 64/138 (46%), Positives = 84/138 (60%)
Frame = +3
Query: 9 DVPHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIK 68
D HF+L P ++Q H+IP +AKLLA G+ VTI TP NA+RF V+ +A +S L+I
Sbjct: 3 DQQHFLLVPFMSQSHLIPFTQLAKLLAANGITVTIVLTPLNATRFNMVIDQAKASNLKIH 182
Query: 69 IVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISD 128
L FP K+AGLPEGCEN D V S + F A +L++P E+ L P+CIISD
Sbjct: 183 FKVLPFPCKEAGLPEGCENMDSVTSPQHQPLFFAACNMLKQPLEKWLSELETVPTCIISD 362
Query: 129 FFIPWTIQIAENHKIPRI 146
+PWT A IPR+
Sbjct: 363 ICLPWTSSTATKFNIPRV 416
>TC9485 similar to UP|Q8S999 (Q8S999) Glucosyltransferase-10, partial (75%)
Length = 1195
Score = 103 bits (257), Expect = 4e-23
Identities = 97/336 (28%), Positives = 151/336 (44%), Gaps = 26/336 (7%)
Frame = +1
Query: 11 PHFVLFPLIAQGHIIPMIDIAKLLAQRGVI-VTIFTTPKNASRFTSVLSRAVSSGLQIKI 69
PH V P AQGHI PM+ +AKLL +G VT T N R +GL
Sbjct: 88 PHVVCIPYPAQGHINPMLKLAKLLHFKGGFHVTFVNTEYNHKRLLKSRGPDSLNGLP--- 258
Query: 70 VTLNFPTKQAGLPEGCENFDMVDSIDM------RMNLFHAITLLQKPAEELFDALTPKPS 123
+ F T GLPE + D+ I + L H LL K + D P +
Sbjct: 259 -SFRFETIPDGLPE--TDVDVTQDIPSLCISTRKTCLPHFKKLLSKLNDVSSDV--PPVT 423
Query: 124 CIISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILER--------VDSES 175
CI+SD + +T+ A IP + F ++ M +Q +++E+ D +
Sbjct: 424 CIVSDGCMSFTLDAAIELNIPEVLF--WTTSACGFMGYVQYRELIEKGIIPLKDSSDITN 597
Query: 176 EYF--TVPGIPDQIQVTKEQIPGILKGELKEFGEKMHD-------AEMKSYGEIINTFEE 226
Y T+ +P + + +P L+ + +KM D +K+ I+NTF+
Sbjct: 598 GYLETTIEWLPGMKNIRLKDLPSFLR--TTDPNDKMLDFLTGECQRALKASAIILNTFDA 771
Query: 227 LEKAYVKDYKKEKNGKVWFVGPVSLCNKDGLDKAQRGIIASI--SEHHCLKWLDLHQPKS 284
LE ++ + V+ +GP+ L KD DK + +++ + CLKWLD +P S
Sbjct: 772 LEHDVLEAFSSILP-PVYSIGPLHLLIKDVTDKNLNSLGSNLWKEDSECLKWLDTKEPNS 948
Query: 285 VVYACLGSLCNLIPSQLMELALALEATNRPFIWVIR 320
VVY GS+ + Q++E A L +N+ F+WVIR
Sbjct: 949 VVYVNFGSITVMTSEQMVEFAWGLANSNKTFLWVIR 1056
>TC9079 weakly similar to UP|Q8W4G1 (Q8W4G1) UDP-glucose
glucosyltransferase, partial (65%)
Length = 1585
Score = 100 bits (248), Expect = 5e-22
Identities = 96/338 (28%), Positives = 144/338 (42%), Gaps = 28/338 (8%)
Frame = +1
Query: 11 PHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIV 70
PH VL P QGHI P+ +AKLL RG +T T N R GL +
Sbjct: 16 PHAVLTPSPLQGHINPLFQLAKLLHLRGFHITFVHTEYNHKRLIKSRGPNALDGLPDFV- 192
Query: 71 TLNFPTKQAGLPEGC-----ENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCI 125
F T GL +G + + + DSI + +L LL K L P +C+
Sbjct: 193 ---FETIPDGLEDGDGEVSQDLYSLCDSIRNKCHLPFQ-DLLAKLHRSASAGLVPPVTCL 360
Query: 126 ISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPD 185
+SD + +TIQ A+ ++P + S L QT + + E + G D
Sbjct: 361 VSDLLMTFTIQAAQQLQLPILLLCPASASTFMSFLHFQTLLDKGVIPLKDESYLTNGYLD 540
Query: 186 QIQVTK-EQIPGILKGELK----------------EFGEKMHDAEMKSYGEIINTFEELE 228
TK + IPG+ LK EF ++ D ++ + NTF ELE
Sbjct: 541 ----TKVDWIPGMQNFRLKDLPDFIRTTDPNDIMLEFMVEVADRAKRASAIVFNTFNELE 708
Query: 229 KAYVKDYKKEKNGKVWFVGPVSLCNKDGLDKAQRGIIASISEH------HCLKWLDLHQP 282
+ V ++ +GP L++A + +AS+ + CL+WL+ +P
Sbjct: 709 RD-VLSALSTMLPSLYPIGPFP----SFLNQAPQNQLASLGSNLWKEDTKCLQWLESKEP 873
Query: 283 KSVVYACLGSLCNLIPSQLMELALALEATNRPFIWVIR 320
SVVY GS+ + P QL+E A L +PF+W+IR
Sbjct: 874 GSVVYVNFGSITVMSPEQLLEFAWGLANGKKPFLWIIR 987
>TC11585 similar to UP|Q9ZWQ5 (Q9ZWQ5) UDP-glycose:flavonoid
glycosyltransferase, partial (40%)
Length = 609
Score = 97.4 bits (241), Expect = 3e-21
Identities = 60/183 (32%), Positives = 93/183 (50%)
Frame = +1
Query: 17 PLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVTLNFPT 76
P AQGH+IP++ +A+L+A RG VTI TTP NA F + + G I++ + FP+
Sbjct: 46 PFFAQGHLIPLVHLARLVASRGEHVTIITTPANAQLFDKTIEEDRACGYHIRVHLVKFPS 225
Query: 77 KQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFIPWTIQ 136
Q GLP G EN S ++ H L +P E F +P P +I D W+
Sbjct: 226 TQVGLPGGVENL-FAASDNLTAGKIHMAAHLIQPEIEGFMKQSP-PDVLIPDIMFTWSEA 399
Query: 137 IAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVTKEQIPG 196
A++ IPR+ F+ S F + CM++ S S+S + +PG+P + + + PG
Sbjct: 400 SAKSLGIPRLIFNPISIFDV-CMIEAIKSHPEAFTCSDSGPYHIPGLPHSLTLPIKPSPG 576
Query: 197 ILK 199
+
Sbjct: 577 FAR 585
>BI418269
Length = 520
Score = 87.0 bits (214), Expect = 4e-18
Identities = 51/143 (35%), Positives = 74/143 (51%)
Frame = +2
Query: 17 PLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVTLNFPT 76
P +A GH+IP+ DIA L A RG VTI TTP N + S L + + T+ FP+
Sbjct: 95 PYLAAGHMIPLCDIAILFASRGQHVTIITTPSNTTTIDSSLP-------NLHLHTVPFPS 253
Query: 77 KQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFIPWTIQ 136
+Q GLP+G E+F + + I LL++P +LF P C+++D+ PW
Sbjct: 254 RQVGLPDGIESFSSSADLQTTFKVHQGIKLLREPI-QLFMEHHP-ADCVVADYCFPWIDD 427
Query: 137 IAENHKIPRISFHGFSCFCLHCM 159
+ IPR+SF+ F L M
Sbjct: 428 LTTELHIPRLSFNPSPLFALCAM 496
>TC15956 weakly similar to UP|LGT_CITUN (Q9MB73) Limonoid
UDP-glucosyltransferase (Limonoid glucosyltransferase)
(Limonoid GTase) (LGTase) , partial (65%)
Length = 1169
Score = 75.9 bits (185), Expect = 9e-15
Identities = 81/327 (24%), Positives = 138/327 (41%), Gaps = 18/327 (5%)
Frame = +2
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
H +L AQGHI P++ +AK LA +G VT+ TT + + + I +
Sbjct: 38 HILLVSFPAQGHINPLLRLAKCLAAKGSSVTLATTQTAGKSMRTATNITDKAPTPISDGS 217
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQ--KPAEELFDALTPKPSCIISDF 129
L F GL G ++ + ++ + + + Q K +E D+ + SCII++
Sbjct: 218 LKFEFFDDGL--GPDDDSIRGNLSLYLQQLELVGRQQILKMIKEHADS-NRQISCIINNP 388
Query: 130 FIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKI----------LERVDSESEYFT 179
F+PW + +AE IP +L IQ+S + L S+ +
Sbjct: 389 FLPWVVDVAEEQGIP------------SALLWIQSSAVFTAYYSYFHNLVTFPSKEDPLA 532
Query: 180 VPGIPDQIQVTKE-QIPGILKGE-----LKEFGEKMHDAEMKSYGEIINTFEELEKAYVK 233
+P V K +IP L L + K++ +++T+EELE ++
Sbjct: 533 DVQLPSTSIVLKHCEIPDFLHPSCTYPFLGTLILEQFKNLNKTFCILVDTYEELEHDFI- 709
Query: 234 DYKKEKNGKVWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSL 293
DY K + + PV K K+ + C++WL+ SVVY GS+
Sbjct: 710 DYL----SKTFLIRPVGPLFKSSQAKSAAIRGDFMKSDDCIEWLNSKPQSSVVYVSFGSI 877
Query: 294 CNLIPSQLMELALALEATNRPFIWVIR 320
L Q+ E+A L+ + F+WV++
Sbjct: 878 VYLPQEQVDEIAFGLKKSQVSFLWVMK 958
>AV769958
Length = 442
Score = 75.5 bits (184), Expect = 1e-14
Identities = 32/57 (56%), Positives = 46/57 (80%)
Frame = +1
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIK 68
HFVLFPL++ GH++PMID+A +LA + +IVT+ TTP NA+RF+ +RA SGLQ++
Sbjct: 145 HFVLFPLMSPGHMLPMIDLATILAHQNIIVTVITTPHNAARFSETFTRASESGLQLR 315
Score = 47.4 bits (111), Expect = 4e-06
Identities = 18/41 (43%), Positives = 30/41 (72%)
Frame = +3
Query: 160 LKIQTSKILERVDSESEYFTVPGIPDQIQVTKEQIPGILKG 200
+ + TS++LE ++SEYF +PGIPD+I++T+ Q+P G
Sbjct: 315 MNLLTSQVLENTATDSEYFAIPGIPDKIEITRAQMPSSSNG 437
>TC11336 similar to UP|Q8S9A5 (Q8S9A5) Glucosyltransferase like protein
(Fragment), partial (27%)
Length = 511
Score = 70.5 bits (171), Expect = 4e-13
Identities = 44/125 (35%), Positives = 66/125 (52%)
Frame = +3
Query: 26 PMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVTLNFPTKQAGLPEGC 85
P++D+ LA G+ +TI TPKN +LS ++ I+ + L FP+ +P G
Sbjct: 87 PLLDLTHHLALGGLTITIIITPKNLPILNPLLSTHPNT---IQTLILPFPS-HPNIPAGA 254
Query: 86 ENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFIPWTIQIAENHKIPR 145
EN V + +A++ LQ+P + F P +ISDFF+ WT Q+A IPR
Sbjct: 255 ENLREVGNTG-NYPFINALSKLQQPIIQWFTTHPNPPVALISDFFLGWTHQLATQLSIPR 431
Query: 146 ISFHG 150
I+FHG
Sbjct: 432 IAFHG 446
>TC17647 weakly similar to UP|AAR06918 (AAR06918) UDP-glycosyltransferase
91D1, partial (13%)
Length = 1059
Score = 67.8 bits (164), Expect = 3e-12
Identities = 88/330 (26%), Positives = 130/330 (38%), Gaps = 24/330 (7%)
Frame = -2
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
H V+ P +A GHI P ++AK+LAQ+G VT +PKN R S I +V
Sbjct: 908 HIVMLPWLAMGHIYPYFEVAKVLAQKGHSVTFINSPKNIDRMPKT---PKSLEPFINLVR 738
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMN----LFHAITLLQKPAEELFDALTPKPSCIIS 127
L P + LPEG E+ ++D+ N L A LQ E+ T KP ++
Sbjct: 737 LPLPHIE-HLPEGAES-----TMDIPTNKGCFLKLAYEGLQDAVAEILQ--TSKPDWVLY 582
Query: 128 DFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKI-------LERVDSESEYFTV 180
DF W IA++ IP C H + +K +++ + E
Sbjct: 581 DFAAGWLPPIAKSLNIP----------CAHYNITPAWNKCFFDPPEHVKKSNFSIENICG 432
Query: 181 PGIPDQIQVTKEQIP-------GILKGELKEFGEKMHDAEMKSYGE----IINTFEELEK 229
P + T + P LK E G+ K+Y ++ T ELE
Sbjct: 431 PPTWVPFKTTIKLRPYEFMRAFAALKDE--STGKSASFDLKKAYSSCDLFLLRTSRELEG 258
Query: 230 AYVKDYKKEKNGKVWFVG--PVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVY 287
++ V VG P S+ +D ++ + I WLD +P +VVY
Sbjct: 257 EWLDYLADTYKVPVVPVGLLPPSMQIRDDDEEEKNPDWVEIK-----AWLDTQEPSTVVY 93
Query: 288 ACLGSLCNLIPSQLMELALALEATNRPFIW 317
GS L L ELA ++ + PF W
Sbjct: 92 IGFGSELKLSQQDLTELAHGIKLSGLPFFW 3
>AV407897
Length = 295
Score = 62.8 bits (151), Expect = 8e-11
Identities = 33/77 (42%), Positives = 48/77 (61%)
Frame = +2
Query: 19 IAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVTLNFPTKQ 78
+A GH+IP+ DIA L A RG VTI TTP NA +L +++ S +K+ + FP ++
Sbjct: 80 LAAGHMIPLCDIATLFASRGHHVTIITTPSNA----QILQKSIPSH-NLKLHAVKFPAQE 244
Query: 79 AGLPEGCENFDMVDSID 95
GLP+G E+ V+ ID
Sbjct: 245 LGLPDGVESLSAVNDID 295
>AW428786
Length = 322
Score = 62.0 bits (149), Expect = 1e-10
Identities = 27/53 (50%), Positives = 36/53 (66%)
Frame = +2
Query: 273 CLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALALEATNRPFIWVIREGNKS 325
CL+WLD P+SV+Y GSL L SQ ELALAL+ ++PF+WV+R N +
Sbjct: 164 CLEWLDQQPPQSVIYVSFGSLATLEQSQFNELALALDLLDKPFLWVVRPDNNN 322
>CB829602
Length = 542
Score = 60.1 bits (144), Expect = 5e-10
Identities = 30/77 (38%), Positives = 43/77 (54%), Gaps = 2/77 (2%)
Frame = +1
Query: 246 VGPVSLCNKDGLDKAQRGIIASI--SEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLME 303
+GP+ L + I ++ E CLKWLD +P SV+Y GS+ N+ P QL+E
Sbjct: 7 IGPLDLLLNQVTENRIESIKCNLWKEEPECLKWLDSQEPSSVLYVNFGSVINMTPQQLVE 186
Query: 304 LALALEATNRPFIWVIR 320
LA + + + FIWVIR
Sbjct: 187 LAWGIANSKKKFIWVIR 237
>TC10167 weakly similar to UP|Q9LTA3 (Q9LTA3) Anthocyanidin-3-glucoside
rhamnosyltransferase-like (At5g49690), partial (9%)
Length = 862
Score = 57.4 bits (137), Expect = 3e-09
Identities = 47/144 (32%), Positives = 67/144 (45%), Gaps = 5/144 (3%)
Frame = +3
Query: 6 NINDVP-HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSG 64
N D P H V+ P +A GHI P +AK+LAQ+G +T +PKN R S
Sbjct: 252 NGEDKPLHIVMIPWLAMGHITPYFALAKVLAQKGHFITFINSPKNIDRMP---KPPKSLE 422
Query: 65 LQIKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITL----LQKPAEELFDALTP 120
I +V L P + LPEG E+ ++D+ N + + L LQ E+ T
Sbjct: 423 PFINLVRLPLPPIE-HLPEGAES-----TMDIPTNKGYYLKLAYEGLQDAVAEILQ--TS 578
Query: 121 KPSCIISDFFIPWTIQIAENHKIP 144
KP ++ DF W A++ IP
Sbjct: 579 KPDWVLYDFAADWLPPTAKSLNIP 650
>BP059889
Length = 509
Score = 56.2 bits (134), Expect = 8e-09
Identities = 24/52 (46%), Positives = 35/52 (67%)
Frame = +1
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSS 63
H V PL+A GH++PM+D+AKL A+R V V+I T P N+ +F + R + S
Sbjct: 352 HLVFIPLMAPGHLLPMVDMAKLFARRNVTVSIVTKPLNSIQFRDTIDREIQS 507
>TC14818 weakly similar to UP|Q8S999 (Q8S999) Glucosyltransferase-10,
partial (24%)
Length = 805
Score = 51.6 bits (122), Expect = 2e-07
Identities = 56/210 (26%), Positives = 80/210 (37%), Gaps = 5/210 (2%)
Frame = +2
Query: 11 PHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIV 70
PH V P AQGH+ P + + KLL G +T T N R L GL
Sbjct: 77 PHAVCVPYPAQGHVNPFMQLTKLLRCMGFHITFVNTEYNHKRLAKSLGPEFVKGLP---- 244
Query: 71 TLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKP-----SCI 125
F T GLP D SI + +P +EL + L P SC+
Sbjct: 245 DFRFETIPDGLPP--PEKDATQSIPLLSQ--STSKHCYEPFKELVNKLNSSPEGPPVSCV 412
Query: 126 ISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPD 185
I+D + + +A + I I F S L L+ + + V + E F G D
Sbjct: 413 ITDGVMGFASSVANDLGIQEIQFWTASACGLLAYLQFEELLKRDIVPLKDENFMSDGTLD 592
Query: 186 QIQVTKEQIPGILKGELKEFGEKMHDAEMK 215
T + I G+ LK+F + ++K
Sbjct: 593 ---TTIDWISGMKNMRLKDFPSFVRVTDLK 673
>AV406975
Length = 210
Score = 48.1 bits (113), Expect = 2e-06
Identities = 21/46 (45%), Positives = 30/46 (64%)
Frame = +3
Query: 14 VLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSR 59
+ P I+ H+IP++D A+L A GV VTI TTP NA+ F + + R
Sbjct: 60 IFLPFISTSHLIPVVDTARLFAMHGVDVTIITTPANATIFQTSIDR 197
>TC18079 weakly similar to UP|UFO5_MANES (Q40287) Flavonol
3-O-glucosyltransferase 5 (UDP-glucose flavonoid
3-O-glucosyltransferase 5) , partial (33%)
Length = 889
Score = 44.7 bits (104), Expect = 2e-05
Identities = 20/52 (38%), Positives = 30/52 (57%)
Frame = +3
Query: 274 LKWLDLHQPKSVVYACLGSLCNLIPSQLMELALALEATNRPFIWVIREGNKS 325
L+WLD SV+Y GS + Q+ E+AL LE + F+WV+R N++
Sbjct: 63 LRWLDKQPAGSVIYVSFGSGGTMPQGQMTEIALGLELSQHRFVWVVRPPNEA 218
>AV427981
Length = 312
Score = 43.9 bits (102), Expect = 4e-05
Identities = 33/103 (32%), Positives = 51/103 (49%), Gaps = 6/103 (5%)
Frame = +3
Query: 220 IINTFEELEKAYVKDYKKEKNGKVWFVGPVSL-----CNKDGLDKAQRGIIASISEHHCL 274
++N+F + K ++ ++ N V P++L C D+ + + + CL
Sbjct: 15 LVNSFPDETKLELQSHELPNNKGSQGVCPIALPIAPICR----DQITKSLSFWEEDLSCL 182
Query: 275 KWLDLHQPKSVVYACLGSLCNLIPSQLME-LALALEATNRPFI 316
KWL + SVVY GS + I M+ LALALEA+ RPFI
Sbjct: 183 KWLATQKDNSVVYISFGSWVSPIGEPKMKSLALALEASGRPFI 311
>TC13901 similar to UP|Q9FYU7 (Q9FYU7) UDP-glucose:sinapate
glucosyltransferase , partial (16%)
Length = 558
Score = 43.9 bits (102), Expect = 4e-05
Identities = 43/160 (26%), Positives = 72/160 (44%), Gaps = 11/160 (6%)
Frame = +3
Query: 5 ANINDVP-HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKN-------ASRFTSV 56
A +++ P H +L AQGHI P++ + K LA +G+ VT TT + A+ T+
Sbjct: 24 AMVSESPIHVLLVSFPAQGHISPLLRLGKHLAAKGLFVTFSTTRQKGEEMVVAAAATTTA 203
Query: 57 LSRAVSSG---LQIKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEE 113
+ SS I L F GLPE + + ++ + +L + ++
Sbjct: 204 TANGTSSPPPPTPIGDGFLQFDLYDDGLPEDHPTRNDLKDYSLQXAEYGK-KILPQLIKK 380
Query: 114 LFDALTPKPSCIISDFFIPWTIQIAENHKIPRISFHGFSC 153
DA P SCII++ F+ W ++A IP + +C
Sbjct: 381 HADANRPV-SCIINNPFVSWVCEVAAELDIPCAMYWIHAC 497
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.139 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,432,030
Number of Sequences: 28460
Number of extensions: 94474
Number of successful extensions: 557
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 538
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 542
length of query: 346
length of database: 4,897,600
effective HSP length: 91
effective length of query: 255
effective length of database: 2,307,740
effective search space: 588473700
effective search space used: 588473700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC147435.1


