

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
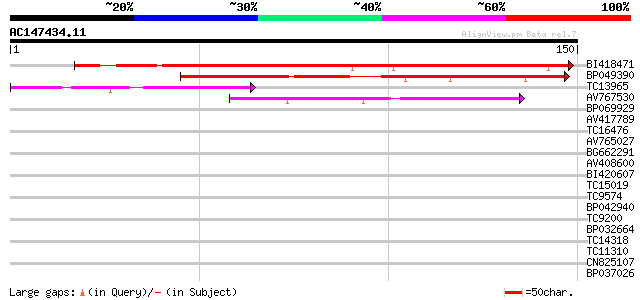
Query= AC147434.11 - phase: 0
(150 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BI418471 112 3e-26
BP049390 71 7e-14
TC13965 weakly similar to UP|HS11_MEDSA (P27879) 18.1 kDa class ... 48 8e-07
AV767530 46 2e-06
BP069929 37 0.002
AV417789 36 0.002
TC16476 similar to UP|Q9LJ93 (Q9LJ93) Emb|CAB10262.1, partial (10%) 33 0.027
AV765027 30 0.23
BG662291 28 0.67
AV408600 27 1.5
BI420607 26 2.5
TC15019 UP|Q9FY16 (Q9FY16) Ferredoxin-nitrite reductase precurso... 26 2.5
TC9574 26 3.3
BP042940 26 3.3
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 25 4.3
BP032664 25 5.7
TC14318 similar to UP|O64739 (O64739) Ran-binding protein (AtRan... 25 5.7
TC11310 similar to UP|BAD11331 (BAD11331) BRI1-KD interacting pr... 25 5.7
CN825107 25 5.7
BP037026 25 5.7
>BI418471
Length = 574
Score = 112 bits (280), Expect = 3e-26
Identities = 61/138 (44%), Positives = 85/138 (61%), Gaps = 6/138 (4%)
Frame = +2
Query: 18 PFSMDIWDPLQGFPSSARETTALANTRVDWKETQEAHVFSVDLPGLKKEEVKVEIEDGNV 77
PF +W G P RE + + +DW E+ AH+ +++PG K+++KV+IEDGN+
Sbjct: 68 PFGRFLW----GHPPIYREWSG-STPLLDWLESPTAHILKINVPGFSKDDIKVQIEDGNI 232
Query: 78 LQISGERNKEQE-EKDDKWHRVER----SSGKFMRRFRLPENVKMDQVKAGMENGVLTVT 132
L + GE KE+ KD WH ER G F R LPENVK+DQ+KA +ENGVLTV
Sbjct: 233 LHVKGEGGKEEALAKDTVWHVAERGIGNGKGDFSRAIELPENVKVDQIKAHVENGVLTVL 412
Query: 133 VPKEEEKKS-EVKSIEIS 149
VPKE KS +V+++ I+
Sbjct: 413 VPKEAAPKSPKVRNVNIT 466
>BP049390
Length = 441
Score = 71.2 bits (173), Expect = 7e-14
Identities = 45/106 (42%), Positives = 69/106 (64%), Gaps = 3/106 (2%)
Frame = -1
Query: 46 DWKETQEAHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSG-K 104
D +ET++A + V++PGL KE+VK+ +E N L I GE KE EE E G +
Sbjct: 378 DARETEDALLLRVNMPGLGKEDVKIAVEQ-NTLTIRGEGPKESEE--------EVGGGLR 226
Query: 105 FMRRFRLPENV-KMDQVKAGMENGVLTVTVPK-EEEKKSEVKSIEI 148
+ R LP+ + ++DQ+KA M+NGVL VTVPK +EE++S+V ++ +
Sbjct: 225 YTSRIDLPDQLYQIDQIKAEMKNGVLKVTVPKMKEEERSDVFNVPV 88
>TC13965 weakly similar to UP|HS11_MEDSA (P27879) 18.1 kDa class I heat
shock protein (Fragment), partial (22%)
Length = 348
Score = 47.8 bits (112), Expect = 8e-07
Identities = 28/79 (35%), Positives = 40/79 (50%), Gaps = 14/79 (17%)
Frame = -2
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWD--------------PLQGFPSSARETTALANTRVD 46
MS++P+ GR+ + +P IWD P FPS E + + N+ ++
Sbjct: 224 MSILPNSLFGRRRS--EPHRSHIWDLFQDHGFGAARTSTPHMAFPS---EPSPMVNSHIE 60
Query: 47 WKETQEAHVFSVDLPGLKK 65
WKET EAHV LPGLK+
Sbjct: 59 WKETPEAHVGKAHLPGLKR 3
>AV767530
Length = 476
Score = 46.2 bits (108), Expect = 2e-06
Identities = 25/90 (27%), Positives = 48/90 (52%), Gaps = 12/90 (13%)
Frame = -2
Query: 59 DLPGLKKEEVKVEI-EDGNVLQISGERNKEQEEKD-----------DKWHRVERSSGKFM 106
+ PG ++ + +E+ +D +VL +SG KE + + + W VE +F
Sbjct: 457 ETPGFPRDRLNIELSDDHSVLTVSGSMRKETKADEAGEAGGAQTQRNPWSSVEER--QFS 284
Query: 107 RRFRLPENVKMDQVKAGMENGVLTVTVPKE 136
+ +RLP + ++ +KA E+GVL ++VP +
Sbjct: 283 QSYRLPRDADVESIKADYEHGVLAISVPSK 194
>BP069929
Length = 421
Score = 36.6 bits (83), Expect = 0.002
Identities = 17/33 (51%), Positives = 28/33 (84%), Gaps = 1/33 (3%)
Frame = -2
Query: 117 MDQVKAGMENGVLTVTVPK-EEEKKSEVKSIEI 148
+DQ++A M+NGVL VTVPK +EE++S+V ++ +
Sbjct: 420 IDQIQAEMKNGVLKVTVPKMKEEERSDVFNVTV 322
>AV417789
Length = 331
Score = 36.2 bits (82), Expect = 0.002
Identities = 23/80 (28%), Positives = 38/80 (46%)
Frame = +1
Query: 56 FSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENV 115
F+V LPG +++++KV++ L++ GER ++W R F F +P
Sbjct: 118 FTVMLPGYRRDQLKVQVTSKPALKLMGER----LIVGNRWRR-------FSLEFPIPSEY 264
Query: 116 KMDQVKAGMENGVLTVTVPK 135
D V A E G L++ K
Sbjct: 265 DTDDVTATFEGGRLSIKFGK 324
>TC16476 similar to UP|Q9LJ93 (Q9LJ93) Emb|CAB10262.1, partial (10%)
Length = 1144
Score = 32.7 bits (73), Expect = 0.027
Identities = 24/83 (28%), Positives = 38/83 (44%)
Frame = +1
Query: 53 AHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLP 112
AH + LPG+ K + + DG + ++G + ++ D W R+RLP
Sbjct: 364 AHAVEI-LPGITK--IVIRRLDGGDVAVTGHQRQQPRFGADLW------------RYRLP 498
Query: 113 ENVKMDQVKAGMENGVLTVTVPK 135
+ ++V A G L VTVPK
Sbjct: 499 PWTQPEKVTAVCTGGKLVVTVPK 567
>AV765027
Length = 491
Score = 29.6 bits (65), Expect = 0.23
Identities = 15/30 (50%), Positives = 19/30 (63%)
Frame = -2
Query: 106 MRRFRLPENVKMDQVKAGMENGVLTVTVPK 135
M RFRLPE+ + + A +G L VTVPK
Sbjct: 370 MWRFRLPESTQPELASAIFVDGELIVTVPK 281
>BG662291
Length = 394
Score = 28.1 bits (61), Expect = 0.67
Identities = 16/58 (27%), Positives = 27/58 (45%)
Frame = +3
Query: 63 LKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKMDQV 120
++ EE+ +IED N I E N+E D++ H SS + R+ D++
Sbjct: 3 IRHEEILPDIEDSNTKTIRLEPNQESSTSDEE-HLSRPSSSHNVNPIRMVAETSTDEI 173
>AV408600
Length = 418
Score = 26.9 bits (58), Expect = 1.5
Identities = 18/68 (26%), Positives = 35/68 (51%)
Frame = -3
Query: 33 SARETTALANTRVDWKETQEAHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKD 92
+A ET + +E + HVF GL+ EE++ I++G +++ ++ +EEK
Sbjct: 413 NAGETARIGGEIPGGEELRTRHVFEGLTNGLQSEEIR-GIQNGEDMKMKVRLHEAEEEKQ 237
Query: 93 DKWHRVER 100
H ++R
Sbjct: 236 S--HALQR 219
>BI420607
Length = 408
Score = 26.2 bits (56), Expect = 2.5
Identities = 12/22 (54%), Positives = 16/22 (72%)
Frame = -2
Query: 71 EIEDGNVLQISGERNKEQEEKD 92
E E G L+ SGERNK ++EK+
Sbjct: 194 EEESGPRLKKSGERNKREKEKE 129
>TC15019 UP|Q9FY16 (Q9FY16) Ferredoxin-nitrite reductase precursor ,
complete
Length = 2098
Score = 26.2 bits (56), Expect = 2.5
Identities = 18/54 (33%), Positives = 26/54 (47%), Gaps = 4/54 (7%)
Frame = +3
Query: 66 EEVKVEIEDGNVLQISGERNKEQEEKDDKW----HRVERSSGKFMRRFRLPENV 115
E KV IE+ + S + K+ + KW HR + G+FM R +LP V
Sbjct: 264 ELAKVSIEELD----SSKLTKDDVDVRLKWLGLFHRRKHQYGRFMMRLKLPNGV 413
>TC9574
Length = 980
Score = 25.8 bits (55), Expect = 3.3
Identities = 12/31 (38%), Positives = 18/31 (57%)
Frame = +1
Query: 60 LPGLKKEEVKVEIEDGNVLQISGERNKEQEE 90
LPGL+ E + L++ GER++E EE
Sbjct: 364 LPGLRSELEALRRRHSAALELMGERDEELEE 456
>BP042940
Length = 432
Score = 25.8 bits (55), Expect = 3.3
Identities = 19/51 (37%), Positives = 24/51 (46%)
Frame = -2
Query: 94 KWHRVERSSGKFMRRFRLPENVKMDQVKAGMENGVLTVTVPKEEEKKSEVK 144
KW RVE S K R R EN+ D A +E + T T + E KS +
Sbjct: 263 KWRRVEISMHKTRTRSRKCENLNRD-*NANLEAALKTRTRYQIREHKSRTR 114
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 25.4 bits (54), Expect = 4.3
Identities = 11/35 (31%), Positives = 22/35 (62%)
Frame = +1
Query: 65 KEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVE 99
+EEV+ E+E+ + E +E+EE++++ VE
Sbjct: 208 EEEVEEEVEEEEEEEEEEEEGEEEEEEEEEEEEVE 312
>BP032664
Length = 561
Score = 25.0 bits (53), Expect = 5.7
Identities = 10/44 (22%), Positives = 26/44 (58%)
Frame = +3
Query: 63 LKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFM 106
++ EEV E ++ IS + ++ ++ ++D+W V++ +F+
Sbjct: 72 IENEEVLDEEQNYMEKSISNQGSEMRDMEEDEWQMVQKKGSQFV 203
>TC14318 similar to UP|O64739 (O64739) Ran-binding protein (AtRanBP1b),
partial (73%)
Length = 1099
Score = 25.0 bits (53), Expect = 5.7
Identities = 12/31 (38%), Positives = 18/31 (57%)
Frame = +1
Query: 64 KKEEVKVEIEDGNVLQISGERNKEQEEKDDK 94
KK+EVK E + SGE++K +K D+
Sbjct: 625 KKDEVKTESKAEEQEPASGEKSKADADKKDE 717
>TC11310 similar to UP|BAD11331 (BAD11331) BRI1-KD interacting protein 102
(Fragment), partial (13%)
Length = 667
Score = 25.0 bits (53), Expect = 5.7
Identities = 21/87 (24%), Positives = 41/87 (46%)
Frame = +1
Query: 58 VDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKM 117
V++ G KKE+ K + +D +++ ++ E++ K H+V+ G E K
Sbjct: 64 VEVEGEKKEKKKKKKKDQENGEVASSDEEKAEKEKKKKHKVKVEDGS--PDLDQSEKKKK 237
Query: 118 DQVKAGMENGVLTVTVPKEEEKKSEVK 144
+ E+ ++ K E++KSE K
Sbjct: 238 KKKDQDAEDNAAEISNGK-EDRKSEKK 315
>CN825107
Length = 254
Score = 25.0 bits (53), Expect = 5.7
Identities = 19/42 (45%), Positives = 27/42 (64%), Gaps = 8/42 (19%)
Frame = +3
Query: 66 EEVKVEIEDGNVLQISGER------NKE--QEEKDDKWHRVE 99
EEV+ EDG+VLQ+S E+ N+E +EE++DK VE
Sbjct: 96 EEVE---EDGDVLQVSEEKYVVDGENEEEGEEEEEDKKVGVE 212
>BP037026
Length = 574
Score = 25.0 bits (53), Expect = 5.7
Identities = 11/42 (26%), Positives = 17/42 (40%)
Frame = -3
Query: 20 SMDIWDPLQGFPSSARETTALANTRVDWKETQEAHVFSVDLP 61
SM + L G+ S+ + WKE+ A +D P
Sbjct: 233 SMMVMGTLSGYKSTGKRVGGSRGGTASWKESDAAAAMRLDFP 108
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.312 0.131 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,024,789
Number of Sequences: 28460
Number of extensions: 20913
Number of successful extensions: 96
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 94
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 95
length of query: 150
length of database: 4,897,600
effective HSP length: 82
effective length of query: 68
effective length of database: 2,563,880
effective search space: 174343840
effective search space used: 174343840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 51 (24.3 bits)
Medicago: description of AC147434.11


