

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
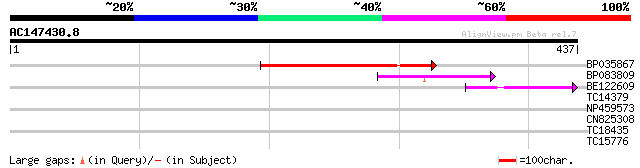
Query= AC147430.8 + phase: 1 /pseudo
(437 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP035867 116 8e-27
BP083809 66 1e-11
BE122609 41 4e-04
TC14379 homologue to UP|O23254 (O23254) Serine hydroxymethyltran... 32 0.27
NP459573 NADH dehydrogenase ND4L [Lotus japonicus] 29 1.7
CN825308 28 3.0
TC18435 28 3.9
TC15776 homologue to UP|PSD6_ORYSA (Q8W425) 26S proteasome non-A... 27 8.7
>BP035867
Length = 482
Score = 116 bits (290), Expect = 8e-27
Identities = 63/136 (46%), Positives = 84/136 (61%)
Frame = -2
Query: 194 KKITTFIYARTSLISILHSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEW 253
+KIT +I +RT+LI++L HT RDL+RPG TRFATSYL LG ++N A RMF S+E
Sbjct: 481 RKITPYICSRTTLITLLQFHTHGRDLIRPGLTRFATSYLNLG*FHDNWGAFERMFASEE* 302
Query: 254 KSGKCAKLRDGKALEDIILDKKFWKDIVICLKGATPLMKVLRLADLDKKTSHGIHL*SNG 313
+ AK RDGK + D+I+DK FWKDI C+KG T + SH + L* N
Sbjct: 301 QKRTYAKTRDGKKICDMIMDKLFWKDITTCVKGCTSFYESASCG*F-*YASHEVDL*GNR 125
Query: 314 SSKGANSKEF*WCQKE 329
G ++K F WC ++
Sbjct: 124 QV*GDDTKSFSWCHEK 77
>BP083809
Length = 369
Score = 66.2 bits (160), Expect = 1e-11
Identities = 39/94 (41%), Positives = 56/94 (59%), Gaps = 3/94 (3%)
Frame = +2
Query: 284 LKGATPLMKVLRLADLDKKTSHGIHL*SNGSSKGA--NSKEF*WCQKEPVWNIIDDRWDR 341
LK PL+ VLRLAD +KK + G + +K A S + + +W+IID+RW+
Sbjct: 83 LKVMGPLVCVLRLADSEKKPAMGYIYEAMDKAKEAIRASLHDDKSKYQKIWDIIDNRWNS 262
Query: 342 MLHRPLHAAAYYLNPQLHYRPG-FKADIEVRKGL 374
LHRPLHAA ++LNP+ +Y D EV+KG+
Sbjct: 263 QLHRPLHAAGHFLNPEYYYNDSELDYDFEVQKGV 364
>BE122609
Length = 356
Score = 40.8 bits (94), Expect = 4e-04
Identities = 26/87 (29%), Positives = 41/87 (46%), Gaps = 1/87 (1%)
Frame = +1
Query: 352 YYLNPQLHY-RPGFKADIEVRKGLMDCITRMVEDPEEQAKIEVQMHDFKKQIGYFGTKIA 410
++LNP+ Y P D E+ +GL + ++E E KI ++ + G+FG K +
Sbjct: 19 HFLNPEFFYANPTMDLDAEIMRGLYE----IIESDENTDKIPNELLVY*MVGGFFGMKAS 186
Query: 411 KLTLDKSTPADWWESVVFEYLELQRFA 437
+PA+WW S LQ FA
Sbjct: 187 IKRRTTISPAEWWRSF*QHTPNLQSFA 267
>TC14379 homologue to UP|O23254 (O23254) Serine hydroxymethyltransferase
(Serine methylase) (Glycine hydroxymethyltransferase)
(SHMT) , partial (37%)
Length = 605
Score = 31.6 bits (70), Expect = 0.27
Identities = 16/46 (34%), Positives = 28/46 (60%)
Frame = -3
Query: 88 RRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVG 133
RRRT + VN+PRG V ++ ++++ + + + E+I AVV G
Sbjct: 405 RRRTGIRLDVNTPRGGVEVEGLESARAAEILDLVDELIAAVVAVTG 268
>NP459573 NADH dehydrogenase ND4L [Lotus japonicus]
Length = 306
Score = 28.9 bits (63), Expect = 1.7
Identities = 17/69 (24%), Positives = 33/69 (47%)
Frame = -2
Query: 153 EMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLISILHS 212
+ L+R+ ++ P AA + + +EK P + K +K+T FI+ + +I
Sbjct: 269 DFLLREIEETIAGPIAASAAAIAITKIEKIFPFNWRLSKKSEKVTRFIF---TAFNISSR 99
Query: 213 HTKNRDLVR 221
H + R + R
Sbjct: 98 HIRARTIFR 72
>CN825308
Length = 600
Score = 28.1 bits (61), Expect = 3.0
Identities = 18/66 (27%), Positives = 25/66 (37%)
Frame = +2
Query: 11 CKVLLQQCHIIQRSKEFNIMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKR 70
C V C + + M S G+ PP Y+D + KL +E ST
Sbjct: 26 CSVFPCFCSVFTPKSLAISLDSMASTEGSAAPPPLYFDEKWKLSKKEGSSTRSRSSSSTN 205
Query: 71 EWKKTS 76
KK+S
Sbjct: 206FMKKSS 223
>TC18435
Length = 505
Score = 27.7 bits (60), Expect = 3.9
Identities = 18/50 (36%), Positives = 24/50 (48%), Gaps = 6/50 (12%)
Frame = -2
Query: 161 KLYWTPCAAHCLDLMLEDLEKKIPI------HGETIPKGKKITTFIYART 204
K YW P A LE+++KKI I G+ G K+ T IY+ T
Sbjct: 432 KDYWAPTLALRWPTSLENIKKKIHIVSIIGGQGKVHILGNKLITSIYSHT 283
>TC15776 homologue to UP|PSD6_ORYSA (Q8W425) 26S proteasome non-ATPase
regulatory subunit 6 (26S proteasome regulatory particle
non-ATPase subunit 7) (OsRPN7), partial (35%)
Length = 686
Score = 26.6 bits (57), Expect = 8.7
Identities = 11/25 (44%), Positives = 14/25 (56%)
Frame = -3
Query: 154 MLMRKRKKLYWTPCAAHCLDLMLED 178
ML R K++WT C +C L L D
Sbjct: 525 MLARPTSKMHWTKCP*NCQYLCLSD 451
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.138 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,675,180
Number of Sequences: 28460
Number of extensions: 100044
Number of successful extensions: 633
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 630
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 632
length of query: 437
length of database: 4,897,600
effective HSP length: 93
effective length of query: 344
effective length of database: 2,250,820
effective search space: 774282080
effective search space used: 774282080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC147430.8


