

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
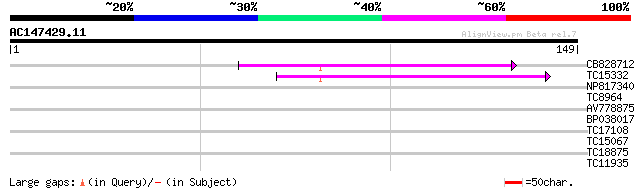
Query= AC147429.11 - phase: 0
(149 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CB828712 42 3e-05
TC15332 similar to UP|PPA1_LYCES (P27061) Acid phosphatase precu... 39 4e-04
NP817340 galactomannan galactosyltransferase [Lotus japonicus] 28 0.51
TC8964 weakly similar to UP|Q94F54 (Q94F54) AT3g15070/K15M2_22, ... 26 2.5
AV778875 26 3.3
BP038017 25 4.3
TC17108 homologue to UP|Q84UD8 (Q84UD8) Apyrase-like protein, pa... 25 5.6
TC15067 25 5.6
TC18875 similar to UP|CTNS_ARATH (P57758) Cystinosin homolog, pa... 25 5.6
TC11935 homologue to UP|S61G_ORYSA (P38385) Protein transport pr... 24 9.5
>CB828712
Length = 532
Score = 42.4 bits (98), Expect = 3e-05
Identities = 24/74 (32%), Positives = 37/74 (49%), Gaps = 1/74 (1%)
Frame = +2
Query: 61 NDYSYCKIHSLHAKLNNLEE-HNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVK 119
+D Y L ++NN+ VP+ C + Y+ GGQY RDL I Y + +
Sbjct: 236 DDERYGLSWRLAVEMNNVRPWRTVPDRCYNHVKNYMTGGQYERDLQLVTEQILLYASEIT 415
Query: 120 PSEDGFDVVLIDID 133
+ DGFD ++D+D
Sbjct: 416 RAGDGFDAWILDVD 457
>TC15332 similar to UP|PPA1_LYCES (P27061) Acid phosphatase precursor 1 ,
partial (89%)
Length = 1040
Score = 38.9 bits (89), Expect = 4e-04
Identities = 23/73 (31%), Positives = 33/73 (44%), Gaps = 1/73 (1%)
Frame = +2
Query: 71 LHAKLNNLEE-HNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSEDGFDVVL 129
L A+ NNL VP C + +Y+ G Y DL+ +Y V+ +DG D +
Sbjct: 254 LAAEANNLTPWKTVPEECAEHVKEYMTGKGYVYDLEMVSKEAGEYAKSVELGDDGKDAWI 433
Query: 130 IDIDSLFQWNPPH 142
DID N P+
Sbjct: 434 FDIDETLLSNLPY 472
>NP817340 galactomannan galactosyltransferase [Lotus japonicus]
Length = 1314
Score = 28.5 bits (62), Expect = 0.51
Identities = 21/61 (34%), Positives = 27/61 (43%)
Frame = +1
Query: 21 FIAGFAALGLLLVTLLVSMAMMLQSCQNSNGGIIELRNINNDYSYCKIHSLHAKLNNLEE 80
F AG A+ +LLVT + C N G + LR N YC+IH + NN
Sbjct: 337 FAAG-ASDRILLVT-----GSQPKRCHNPIGDHLLLRFFKNKVDYCRIHDIDIIYNNALL 498
Query: 81 H 81
H
Sbjct: 499 H 501
>TC8964 weakly similar to UP|Q94F54 (Q94F54) AT3g15070/K15M2_22, partial
(19%)
Length = 594
Score = 26.2 bits (56), Expect = 2.5
Identities = 8/26 (30%), Positives = 17/26 (64%)
Frame = +3
Query: 41 MMLQSCQNSNGGIIELRNINNDYSYC 66
+ ++S + GG +E+ +NN+ S+C
Sbjct: 315 LQIRSFDSWRGGCVEMSEVNNNISFC 392
>AV778875
Length = 464
Score = 25.8 bits (55), Expect = 3.3
Identities = 14/41 (34%), Positives = 20/41 (48%)
Frame = +2
Query: 21 FIAGFAALGLLLVTLLVSMAMMLQSCQNSNGGIIELRNINN 61
+I + +V +L ++ SCQ S GII LRN N
Sbjct: 227 WIGSLTFFRICMVKILGKRFKIVMSCQLSMKGIISLRNWRN 349
>BP038017
Length = 570
Score = 25.4 bits (54), Expect = 4.3
Identities = 10/28 (35%), Positives = 19/28 (67%)
Frame = -3
Query: 39 MAMMLQSCQNSNGGIIELRNINNDYSYC 66
+AMM+ SC+ S ++ +R+I+ +YC
Sbjct: 400 LAMMIDSCKWSKWNLVLIRSIHCVKNYC 317
>TC17108 homologue to UP|Q84UD8 (Q84UD8) Apyrase-like protein, partial
(16%)
Length = 573
Score = 25.0 bits (53), Expect = 5.6
Identities = 9/18 (50%), Positives = 13/18 (72%)
Frame = +1
Query: 77 NLEEHNVPNICKDLALQY 94
++E+ N+P IC DL QY
Sbjct: 55 HIEDGNLPYICMDLVYQY 108
>TC15067
Length = 542
Score = 25.0 bits (53), Expect = 5.6
Identities = 11/37 (29%), Positives = 17/37 (45%), Gaps = 7/37 (18%)
Frame = +2
Query: 68 IHSLHAKLNNLEEHNVPN-------ICKDLALQYIKG 97
+HSLH L + + VPN +C L ++G
Sbjct: 164 LHSLHLALRKIHSYEVPNPRSKELDVCLSLTTSIVQG 274
>TC18875 similar to UP|CTNS_ARATH (P57758) Cystinosin homolog, partial (58%)
Length = 567
Score = 25.0 bits (53), Expect = 5.6
Identities = 15/51 (29%), Positives = 22/51 (42%)
Frame = +1
Query: 20 IFIAGFAALGLLLVTLLVSMAMMLQSCQNSNGGIIELRNINNDYSYCKIHS 70
+F+A A + L + L S NSN ++ Y YCK+HS
Sbjct: 436 VFVAWLTAAICFFIALSDHSWLWLLSIFNSN---------SSSYDYCKVHS 561
>TC11935 homologue to UP|S61G_ORYSA (P38385) Protein transport protein SEC61
gamma subunit, complete
Length = 550
Score = 24.3 bits (51), Expect = 9.5
Identities = 20/81 (24%), Positives = 32/81 (38%)
Frame = -3
Query: 56 LRNINNDYSYCKIHSLHAKLNNLEEHNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYF 115
L N+D LH K N HN N + L ++ LD +V+ ++
Sbjct: 320 LTRSNDDVVNWNEDELHEKPNE-SHHNETNCGTNSNLGEFFAIRFVATLDEPDAVLGEFS 144
Query: 116 NGVKPSEDGFDVVLIDIDSLF 136
G++ +GF + SLF
Sbjct: 143 EGIEHRINGFHSLSFSSLSLF 81
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.138 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,610,008
Number of Sequences: 28460
Number of extensions: 31702
Number of successful extensions: 141
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 141
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 141
length of query: 149
length of database: 4,897,600
effective HSP length: 82
effective length of query: 67
effective length of database: 2,563,880
effective search space: 171779960
effective search space used: 171779960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 51 (24.3 bits)
Medicago: description of AC147429.11


