

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147406.5 - phase: 0 /pseudo
(99 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
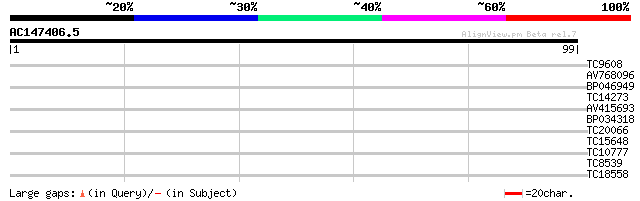
Score E
Sequences producing significant alignments: (bits) Value
TC9608 similar to UP|Q94AZ4 (Q94AZ4) At1g12310/F5O11_2, partial ... 33 0.006
AV768096 31 0.039
BP046949 26 0.97
TC14273 similar to UP|DCE1_ARATH (Q42521) Glutamate decarboxylas... 26 0.97
AV415693 26 1.3
BP034318 25 2.2
TC20066 similar to GB|AAM91455.1|22137220|AY133625 AT3g11400/F24... 23 6.3
TC15648 23 6.3
TC10777 similar to UP|Q9SCL3 (Q9SCL3) Pre-mRNA splicing factor S... 23 6.3
TC8539 similar to UP|Q94KE3 (Q94KE3) AT3g52990/F8J2_160 (Pyruvat... 23 8.2
TC18558 similar to UP|Q7XAE5 (Q7XAE5) S1 self-incompatibility lo... 23 8.2
>TC9608 similar to UP|Q94AZ4 (Q94AZ4) At1g12310/F5O11_2, partial (97%)
Length = 817
Score = 33.5 bits (75), Expect = 0.006
Identities = 15/40 (37%), Positives = 24/40 (59%)
Frame = -3
Query: 27 MQFVVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGF 66
++ V KL + LG+H +QK+ +GG EI+ D+GF
Sbjct: 371 LEGVTKLAIEGLGFHVFGHEIQKSREIEGGGEILLGDDGF 252
>AV768096
Length = 358
Score = 30.8 bits (68), Expect = 0.039
Identities = 15/70 (21%), Positives = 33/70 (46%)
Frame = +3
Query: 29 FVVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWLI 88
F K+L K A+++ + + W ++ D + ++ F +A +++ GPW I
Sbjct: 3 FTYKVLNKL----AVKQTIVRAWGNLQDLQVGDLNLNTFLFTFSDANLATRILAEGPWNI 170
Query: 89 FDHCLAVTHW 98
+ L++ W
Sbjct: 171MGNLLSLQFW 200
>BP046949
Length = 408
Score = 26.2 bits (56), Expect = 0.97
Identities = 16/44 (36%), Positives = 23/44 (51%), Gaps = 5/44 (11%)
Frame = +1
Query: 48 QKTWRTQG----GFEIMDNDNGFYMVKFDEAADKEKVIT-GGPW 86
+K ++QG G+EIMD D +V F K+ V+ GG W
Sbjct: 40 KKIHKSQGPLKNGYEIMDGDTEIQLVLFR*KGGKQYVLKFGGAW 171
>TC14273 similar to UP|DCE1_ARATH (Q42521) Glutamate decarboxylase 1 (GAD 1)
, partial (89%)
Length = 1826
Score = 26.2 bits (56), Expect = 0.97
Identities = 14/29 (48%), Positives = 18/29 (61%)
Frame = +3
Query: 43 MRERLQKTWRTQGGFEIMDNDNGFYMVKF 71
++E LQKT G FEI+ DNG +V F
Sbjct: 1125 LKEGLQKT----GRFEIVSKDNGVPLVAF 1199
>AV415693
Length = 394
Score = 25.8 bits (55), Expect = 1.3
Identities = 13/42 (30%), Positives = 24/42 (56%)
Frame = -3
Query: 15 NQSFNNYVRHGRMQFVVKLLGKNLGYHAMRERLQKTWRTQGG 56
NQ F N+ + + KL ++L HA+R + ++ W+ +GG
Sbjct: 212 NQPFCNWKPEEALYW*TKLQKQHLP-HALRRKNERRWKKRGG 90
>BP034318
Length = 559
Score = 25.0 bits (53), Expect = 2.2
Identities = 13/38 (34%), Positives = 21/38 (55%)
Frame = -2
Query: 12 ISNNQSFNNYVRHGRMQFVVKLLGKNLGYHAMRERLQK 49
I N SFN GR QF+++ + K+ +R RL++
Sbjct: 195 IGGNVSFNYTSSGGRKQFIIRSIRKDSIDGLVRSRLKE 82
>TC20066 similar to GB|AAM91455.1|22137220|AY133625 AT3g11400/F24K9_7
{Arabidopsis thaliana;}, partial (45%)
Length = 557
Score = 23.5 bits (49), Expect = 6.3
Identities = 7/11 (63%), Positives = 8/11 (72%)
Frame = -1
Query: 83 GGPWLIFDHCL 93
G PWL +HCL
Sbjct: 80 GAPWLSIEHCL 48
>TC15648
Length = 536
Score = 23.5 bits (49), Expect = 6.3
Identities = 8/18 (44%), Positives = 12/18 (66%), Gaps = 1/18 (5%)
Frame = -2
Query: 82 TGGPWLIFD-HCLAVTHW 98
T GP+ ++ HCL + HW
Sbjct: 202 THGPFTVYPWHCLQLVHW 149
>TC10777 similar to UP|Q9SCL3 (Q9SCL3) Pre-mRNA splicing factor SF2-like
protein, partial (46%)
Length = 597
Score = 23.5 bits (49), Expect = 6.3
Identities = 11/27 (40%), Positives = 13/27 (47%)
Frame = +1
Query: 66 FYMVKFDEAADKEKVITGGPWLIFDHC 92
+ V+FD A D E I G FD C
Sbjct: 376 YCFVEFDNARDAEDAIRGRDGYNFDGC 456
>TC8539 similar to UP|Q94KE3 (Q94KE3) AT3g52990/F8J2_160 (Pyruvate
kinase-like protein), partial (81%)
Length = 1430
Score = 23.1 bits (48), Expect = 8.2
Identities = 8/19 (42%), Positives = 10/19 (52%)
Frame = -3
Query: 45 ERLQKTWRTQGGFEIMDND 63
ER + WRT + DND
Sbjct: 870 ERNSRAWRTSSACRVYDND 814
>TC18558 similar to UP|Q7XAE5 (Q7XAE5) S1 self-incompatibility locus-linked
pollen 3.15 protein (Fragment), partial (18%)
Length = 639
Score = 23.1 bits (48), Expect = 8.2
Identities = 10/24 (41%), Positives = 15/24 (61%), Gaps = 2/24 (8%)
Frame = +2
Query: 77 KEKVITGG--PWLIFDHCLAVTHW 98
K V T G +LI+D C+ +TH+
Sbjct: 77 KSAVTTSGLSHYLIYDSCINITHY 148
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.141 0.460
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,734,878
Number of Sequences: 28460
Number of extensions: 17391
Number of successful extensions: 127
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 127
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 127
length of query: 99
length of database: 4,897,600
effective HSP length: 75
effective length of query: 24
effective length of database: 2,763,100
effective search space: 66314400
effective search space used: 66314400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 47 (22.7 bits)
Medicago: description of AC147406.5


