

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147406.4 - phase: 0
(310 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
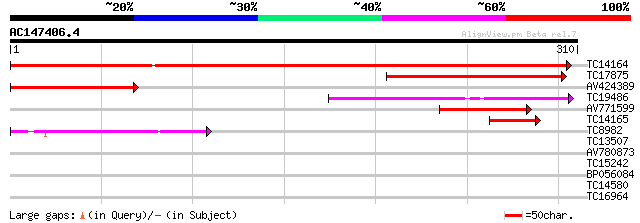
TC14164 weakly similar to UP|Q8VYH6 (Q8VYH6) AT4g17280/dl4675c, ... 436 e-123
TC17875 weakly similar to UP|Q8VYH6 (Q8VYH6) AT4g17280/dl4675c, ... 160 3e-40
AV424389 118 1e-27
TC19486 similar to UP|Q9SJ74 (Q9SJ74) Expressed protein, partial... 97 5e-21
AV771599 48 2e-06
TC14165 45 2e-05
TC8982 similar to UP|Q84QW4 (Q84QW4) OJ1191_A10.2 protein, parti... 40 7e-04
TC13507 35 0.021
AV780873 28 2.6
TC15242 similar to UP|ASSY_ARATH (Q9SZX3) Argininosuccinate synt... 27 5.8
BP056084 26 7.5
TC14580 weakly similar to UP|RK21_SPIOL (P24613) 50S ribosomal p... 26 9.8
TC16964 similar to UP|Q9SV20 (Q9SV20) Beta-COP-like protein, par... 26 9.8
>TC14164 weakly similar to UP|Q8VYH6 (Q8VYH6) AT4g17280/dl4675c, partial
(44%)
Length = 1606
Score = 436 bits (1122), Expect = e-123
Identities = 209/307 (68%), Positives = 253/307 (82%)
Frame = +2
Query: 1 MIGAQALVAIPQASGSPKAYTSNIVDTSTRLQEGTISYPVSGLSATYQNNKVTIFATLTL 60
M+GAQALVAIPQ+SGSPK YT++I + + EG ISYP +GLSATY N+++TI+ATLTL
Sbjct: 386 MVGAQALVAIPQSSGSPKVYTTSIANYDPNMAEGNISYPHTGLSATYSNSELTIYATLTL 565
Query: 61 PNGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIGSRQRRRNT 120
P+GTTSLVH+WQDG +S ST Q H+ +++ +KE LDL SGT+QA S S +RRRN
Sbjct: 566 PSGTTSLVHLWQDGAVSG-STLQAHARGTANLQAKESLDLASGTTQAGSSGNSVRRRRNV 742
Query: 121 HGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSAYIVGLSGFGTGLKLGS 180
HGVLN +SWGILMP GAVIARYLKVFKSADPAWFYLH+TCQ +AYIVG++G+GTGLKLGS
Sbjct: 743 HGVLNTVSWGILMPLGAVIARYLKVFKSADPAWFYLHVTCQTAAYIVGVAGWGTGLKLGS 922
Query: 181 DSEGITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNIYHHVVGYVTISISIVN 240
DS GI Y THRAL I L L TLQVFAL LRPNKDHK+RFYWN+YH +GY TI ISIVN
Sbjct: 923 DSAGIEYSTHRALGITLFCLGTLQVFALLLRPNKDHKIRFYWNLYHWGIGYATIIISIVN 1102
Query: 241 VFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLEAYTWMVCMKRKKADNKTSDGVN 300
+FKGF+A+ VGDRY +WK+AYIGII ALGGIAVLLE YTW++ +KR+K++N+ + G N
Sbjct: 1103IFKGFDAMEKSVGDRYDDWKNAYIGIIAALGGIAVLLEVYTWIIVLKRRKSENQMAHGAN 1282
Query: 301 GANGHGS 307
G NG+GS
Sbjct: 1283GVNGYGS 1303
>TC17875 weakly similar to UP|Q8VYH6 (Q8VYH6) AT4g17280/dl4675c, partial
(19%)
Length = 567
Score = 160 bits (405), Expect = 3e-40
Identities = 71/98 (72%), Positives = 88/98 (89%)
Frame = +1
Query: 207 ALFLRPNKDHKLRFYWNIYHHVVGYVTISISIVNVFKGFEALGDFVGDRYKNWKHAYIGI 266
ALFLR NKDHK RFYWNIYHH++GY TI ISI+NVFKGF+ L ++VGDRY NWK+AYIGI
Sbjct: 1 ALFLRLNKDHKHRFYWNIYHHLIGYSTIIISIINVFKGFDTLENYVGDRYDNWKNAYIGI 180
Query: 267 IGALGGIAVLLEAYTWMVCMKRKKADNKTSDGVNGANG 304
I ALGGIAVLLEAYTW++ +KR+++++KTS G+NGA+G
Sbjct: 181 IAALGGIAVLLEAYTWIIVLKRRQSESKTSHGINGAHG 294
>AV424389
Length = 309
Score = 118 bits (295), Expect = 1e-27
Identities = 60/71 (84%), Positives = 67/71 (93%), Gaps = 1/71 (1%)
Frame = +2
Query: 1 MIGAQALVAIPQAS-GSPKAYTSNIVDTSTRLQEGTISYPVSGLSATYQNNKVTIFATLT 59
M+GAQALVAIPQ+S GSP+AYTS+I T+T+LQEGTISYPVSGLSA YQNN+VTIFATLT
Sbjct: 95 MVGAQALVAIPQSSDGSPRAYTSSIASTNTQLQEGTISYPVSGLSAEYQNNEVTIFATLT 274
Query: 60 LPNGTTSLVHV 70
LPNGTTSLVHV
Sbjct: 275 LPNGTTSLVHV 307
>TC19486 similar to UP|Q9SJ74 (Q9SJ74) Expressed protein, partial (31%)
Length = 510
Score = 96.7 bits (239), Expect = 5e-21
Identities = 49/134 (36%), Positives = 74/134 (54%)
Frame = +1
Query: 175 GLKLGSDSEGITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNIYHHVVGYVTI 234
G++LG S G+ Y HR L I + L +Q AL RPN+ ++ R YW YHH VGY +
Sbjct: 4 GIRLGELSPGVEYRLHRKLGIAVFCLGAMQTLALLFRPNERNRFRKYWKSYHHFVGYSCV 183
Query: 235 SISIVNVFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLEAYTWMVCMKRKKADNK 294
+ VNVF+GFE +G Y K +Y + + G+ + LE +W+V ++ K D
Sbjct: 184 VLGFVNVFQGFEVMG--ASRSYA--KLSYCLGLSTMIGVCIALEVNSWVVFCRKSKEDKM 351
Query: 295 TSDGVNGANGHGSS 308
+G+ G++ GSS
Sbjct: 352 RREGLIGSSDKGSS 393
>AV771599
Length = 535
Score = 48.1 bits (113), Expect = 2e-06
Identities = 26/52 (50%), Positives = 35/52 (67%), Gaps = 2/52 (3%)
Frame = -3
Query: 236 ISIVNVFKGFEALGDFVGDRYKNWKHAYI-GIIGALGGIAVLLEAYT-WMVC 285
ISIVN+FKGF+A+ VGDRY +WK+AYI + IAVL+ + W+ C
Sbjct: 533 ISIVNIFKGFDAMEKSVGDRYDDWKYAYIRDYCCSWVAIAVLVGRFIHWIPC 378
>TC14165
Length = 431
Score = 44.7 bits (104), Expect = 2e-05
Identities = 18/28 (64%), Positives = 24/28 (85%)
Frame = +3
Query: 263 YIGIIGALGGIAVLLEAYTWMVCMKRKK 290
YIGII ALGG+AVLLE YTW+ C++ ++
Sbjct: 3 YIGIIAALGGLAVLLEVYTWIPCLEEEE 86
>TC8982 similar to UP|Q84QW4 (Q84QW4) OJ1191_A10.2 protein, partial (10%)
Length = 901
Score = 39.7 bits (91), Expect = 7e-04
Identities = 31/114 (27%), Positives = 50/114 (43%), Gaps = 4/114 (3%)
Frame = +2
Query: 1 MIGAQALVAIPQASGSPK----AYTSNIVDTSTRLQEGTISYPVSGLSATYQNNKVTIFA 56
M G QALVA G K + NI S + + S+ LSA + +TIFA
Sbjct: 263 MAGTQALVAY---KGGDKNVVGVHLYNITSYSELAEVKSFSFETWDLSAEESSAAITIFA 433
Query: 57 TLTLPNGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASG 110
+ +P ++ +WQ G + S +H + + +K L +V+ T+ G
Sbjct: 434 AVKVPENADNISQIWQVGPVVSGKI-NKHDFKPENLAAKAPLSVVATTAVVGGG 592
>TC13507
Length = 413
Score = 34.7 bits (78), Expect = 0.021
Identities = 16/52 (30%), Positives = 30/52 (56%)
Frame = +3
Query: 257 KNWKHAYIGIIGALGGIAVLLEAYTWMVCMKRKKADNKTSDGVNGANGHGSS 308
+ ++ Y+ ++ LG + ++LE TW+V +KRK S+ +G N +G S
Sbjct: 9 EKYRAIYLDVLLILGALVLVLEVLTWIVVLKRKYCSK--SNTYDGYNNNGQS 158
>AV780873
Length = 487
Score = 27.7 bits (60), Expect = 2.6
Identities = 20/60 (33%), Positives = 26/60 (43%), Gaps = 4/60 (6%)
Frame = +3
Query: 147 KSADPAWFYLHITCQVSAYIVGLSGFGTGLKLGSDSEGIT---YDTHR-ALAIVLVTLAT 202
K P W +L C+ I+ + L LG SEGI YD A A++LV T
Sbjct: 207 KKGRPFWMFLWDACKDLTLIILMVAAAASLALGIKSEGIEEGWYDGGSIAFAVLLVIFVT 386
>TC15242 similar to UP|ASSY_ARATH (Q9SZX3) Argininosuccinate synthase,
chloroplast precursor (Citrulline--aspartate ligase) ,
partial (57%)
Length = 1296
Score = 26.6 bits (57), Expect = 5.8
Identities = 10/23 (43%), Positives = 17/23 (73%)
Frame = -3
Query: 86 SHESSHQNSKEVLDLVSGTSQAA 108
++ +SH+ K++L L+SGTS A
Sbjct: 712 NYRASHRGPKQILPLISGTSTQA 644
>BP056084
Length = 522
Score = 26.2 bits (56), Expect = 7.5
Identities = 8/12 (66%), Positives = 10/12 (82%)
Frame = +3
Query: 211 RPNKDHKLRFYW 222
+ N DHKL+FYW
Sbjct: 258 KKNSDHKLKFYW 293
>TC14580 weakly similar to UP|RK21_SPIOL (P24613) 50S ribosomal protein L21,
chloroplast precursor (CL21) (CS-L7), partial (42%)
Length = 938
Score = 25.8 bits (55), Expect = 9.8
Identities = 12/34 (35%), Positives = 20/34 (58%)
Frame = +3
Query: 209 FLRPNKDHKLRFYWNIYHHVVGYVTISISIVNVF 242
F N+ H LRF+ N HH ++ + SI+++F
Sbjct: 36 FCNCNRFHTLRFFRNPLHH---FLHLQTSILSIF 128
>TC16964 similar to UP|Q9SV20 (Q9SV20) Beta-COP-like protein, partial (15%)
Length = 593
Score = 25.8 bits (55), Expect = 9.8
Identities = 19/71 (26%), Positives = 34/71 (47%), Gaps = 4/71 (5%)
Frame = +3
Query: 53 TIFATLTLPNGTTSLVHVWQDGVLSSDSTPQ----EHSHESSHQNSKEVLDLVSGTSQAA 108
T+F++L P ++ + + +S+ S PQ ++ Q + +GTSQ
Sbjct: 66 TLFSSLVFPGSSSPI----PNSTISTPSLPQLLIRSETNGEIVQFGGALRQGNAGTSQRD 233
Query: 109 SGIGSRQRRRN 119
GI R+RRR+
Sbjct: 234 QGISRRKRRRS 266
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,237,548
Number of Sequences: 28460
Number of extensions: 70621
Number of successful extensions: 428
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 424
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 425
length of query: 310
length of database: 4,897,600
effective HSP length: 90
effective length of query: 220
effective length of database: 2,336,200
effective search space: 513964000
effective search space used: 513964000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC147406.4


