

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147405.6 - phase: 0
(416 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
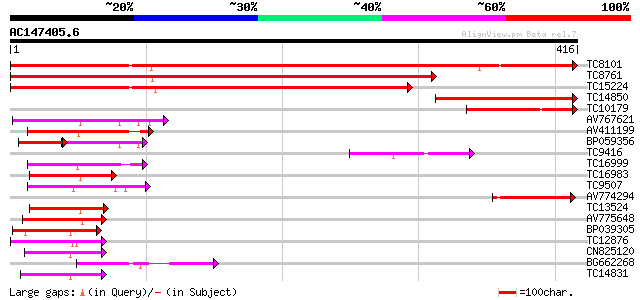
Score E
Sequences producing significant alignments: (bits) Value
TC8101 similar to UP|O24074 (O24074) DNAJ-like protein, partial ... 623 e-179
TC8761 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein P... 587 e-168
TC15224 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein ... 575 e-165
TC14850 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein ... 200 4e-52
TC10179 similar to UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM... 133 5e-32
AV767621 98 2e-21
AV411199 89 1e-18
BP059356 56 3e-15
TC9416 similar to UP|Q9FEW7 (Q9FEW7) DnaJ like protein, partial ... 73 8e-14
TC16999 similar to UP|Q9T024 (Q9T024) DNAJ-like protein (AT4g391... 69 1e-12
TC16983 similar to UP|Q9ZSY2 (Q9ZSY2) ALTERED response to gravit... 68 2e-12
TC9507 similar to UP|Q8VZB6 (Q8VZB6) DnaJ-like protein, partial ... 65 2e-11
AV774294 65 2e-11
TC13524 similar to UP|O48678 (O48678) F3I6.4 protein, partial (19%) 65 3e-11
AV775648 60 5e-10
BP039305 58 3e-09
TC12876 55 2e-08
CN825120 55 2e-08
BG662268 54 5e-08
TC14831 homologue to UP|P93499 (P93499) DnaJ-like protein (Fragm... 52 1e-07
>TC8101 similar to UP|O24074 (O24074) DNAJ-like protein, partial (90%)
Length = 1737
Score = 623 bits (1607), Expect = e-179
Identities = 303/423 (71%), Positives = 361/423 (84%), Gaps = 7/423 (1%)
Frame = +1
Query: 1 MFGRAPKKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEV 60
MFGRAP+KSD+T++YE+LGV K+AS+D++KKAY+KAA+KNHPDKGGDPEKFKEL QAYEV
Sbjct: 148 MFGRAPRKSDNTKFYEVLGVPKSASEDEIKKAYRKAAMKNHPDKGGDPEKFKELGQAYEV 327
Query: 61 LSDPEKREIYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGG---GGGGSSRGRRQRRGE 117
LSDPEK+E+YD YGEDALKEGMGGG H+PFDIF +FFGG GGGGSSRGRRQR+GE
Sbjct: 328 LSDPEKKELYDQYGEDALKEGMGGGESF-HNPFDIFETFFGGAGFGGGGSSRGRRQRQGE 504
Query: 118 DVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMKCAGCQGTGMKVSIRHL 177
DVVH +KVSLED+Y GT+KKLSLSRNVLC KC GKGSKSG + +C GCQGTGMKV R +
Sbjct: 505 DVVHSIKVSLEDVYNGTTKKLSLSRNVLCLKCKGKGSKSGTAGRCFGCQGTGMKVIRRQI 684
Query: 178 GPSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFP 237
G M+QQMQH C++C+GTGE I+++DRCPQCKG K+ QEKKVLEVHVEKGM+ QKI F
Sbjct: 685 GLGMVQQMQHVCSDCRGTGEVISERDRCPQCKGNKISQEKKVLEVHVEKGMRQGQKIVFE 864
Query: 238 GEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQL 297
G+ADEAPDT+TGDIVFV+Q KEHPKFKR+ +DL+++H LSLTEALCGFQF + HLDGRQL
Sbjct: 865 GQADEAPDTITGDIVFVVQVKEHPKFKRELDDLYIDHNLSLTEALCGFQFAVKHLDGRQL 1044
Query: 298 LIKSNPGEVVKPDSYKAINDEGMPMYQRPFMKGKLYIHFTVEFPDT--LSLDQVKGLEAV 355
LIKSNPGEV+KP +KAINDEGMP + RPF+KG+LYI F V+FPD+ +S DQ + LE +
Sbjct: 1045LIKSNPGEVIKPGQHKAINDEGMPQHGRPFIKGRLYIKFNVDFPDSGFISPDQCQLLEKI 1224
Query: 356 LPAKPSSQLTDMEIDECEETTLHDV-NMEEENRRKQQQQQQEAYDED-DDMPGGAQRVQC 413
LP K SS+ DME+D+CEET LHDV N+EEE RR++QQ+ +EAYDED DD QRVQC
Sbjct: 1225LPQK-SSKKADMELDDCEETILHDVNNLEEEMRRRKQQRSREAYDEDEDDDEPSTQRVQC 1401
Query: 414 AQQ 416
AQQ
Sbjct: 1402AQQ 1410
>TC8761 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37,
partial (74%)
Length = 1050
Score = 587 bits (1512), Expect = e-168
Identities = 288/318 (90%), Positives = 300/318 (93%), Gaps = 5/318 (1%)
Frame = +3
Query: 1 MFGRAP-KKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYE 59
MFGRAP KKSDSTRYY+ILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYE
Sbjct: 78 MFGRAPPKKSDSTRYYDILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYE 257
Query: 60 VLSDPEKREIYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGGG----GGGSSRGRRQRR 115
VL+DPEKREIYD YGEDALKEGMGGGGGGGHDPFDIFSSFFGG GGG SRGRRQRR
Sbjct: 258 VLNDPEKREIYDNYGEDALKEGMGGGGGGGHDPFDIFSSFFGGSPFSSGGGGSRGRRQRR 437
Query: 116 GEDVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMKCAGCQGTGMKVSIR 175
GEDVV+PLKVSLEDLYLG +KKLSLSRNVLCSKC+GKGSKSGAS+KC GCQG+GMKVSIR
Sbjct: 438 GEDVVYPLKVSLEDLYLGAAKKLSLSRNVLCSKCNGKGSKSGASLKCPGCQGSGMKVSIR 617
Query: 176 HLGPSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKIT 235
H+GPSMIQQMQHPCNECKGTGETIND+DRCPQCKG+KV QEKKVLEV VEKGMQN QKIT
Sbjct: 618 HIGPSMIQQMQHPCNECKGTGETINDRDRCPQCKGDKVSQEKKVLEVFVEKGMQNQQKIT 797
Query: 236 FPGEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGR 295
FPGEADEAPDT TGDIVFVLQ KEHPKFKRK+EDLFVEHTLSLTEALCGFQFVLTHLDGR
Sbjct: 798 FPGEADEAPDTTTGDIVFVLQLKEHPKFKRKAEDLFVEHTLSLTEALCGFQFVLTHLDGR 977
Query: 296 QLLIKSNPGEVVKPDSYK 313
QLLIKSNPGEVVK +K
Sbjct: 978 QLLIKSNPGEVVKLPLHK 1031
Score = 34.3 bits (77), Expect = 0.039
Identities = 18/29 (62%), Positives = 18/29 (62%)
Frame = -2
Query: 321 PMYQRPFMKGKLYIHFTVEFPDTLSLDQV 349
PMYQRPFMKGKLY V F L QV
Sbjct: 1049 PMYQRPFMKGKLYNFPRV*FDQELPTIQV 963
>TC15224 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37,
partial (71%)
Length = 1031
Score = 575 bits (1483), Expect = e-165
Identities = 284/299 (94%), Positives = 287/299 (95%), Gaps = 4/299 (1%)
Frame = +3
Query: 1 MFGRAPKKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEV 60
MFGRAPKKSDSTRYYEILGV K ASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEV
Sbjct: 138 MFGRAPKKSDSTRYYEILGVPKNASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEV 317
Query: 61 LSDPEKREIYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGGGGG----GSSRGRRQRRG 116
LSDPEKREIYD YGEDALKEGMGGGGGG HDPFDIF SFFGGGGG GSSRGRRQRRG
Sbjct: 318 LSDPEKREIYDQYGEDALKEGMGGGGGG-HDPFDIFQSFFGGGGGPFSGGSSRGRRQRRG 494
Query: 117 EDVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMKCAGCQGTGMKVSIRH 176
EDVVHPLKVSLEDLY GT+KKLSLSRNVLCSKC+GKGSKSGASMKCAGCQGTGMKVSIRH
Sbjct: 495 EDVVHPLKVSLEDLYSGTAKKLSLSRNVLCSKCNGKGSKSGASMKCAGCQGTGMKVSIRH 674
Query: 177 LGPSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITF 236
LGPSMIQQMQHPCNECKGTGETIND+DRCPQCKGEKVVQEKKVLEV VEKGMQN QKITF
Sbjct: 675 LGPSMIQQMQHPCNECKGTGETINDRDRCPQCKGEKVVQEKKVLEVIVEKGMQNGQKITF 854
Query: 237 PGEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGR 295
PGEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGR
Sbjct: 855 PGEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGR 1031
>TC14850 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37,
partial (25%)
Length = 577
Score = 200 bits (508), Expect = 4e-52
Identities = 95/104 (91%), Positives = 100/104 (95%)
Frame = +1
Query: 313 KAINDEGMPMYQRPFMKGKLYIHFTVEFPDTLSLDQVKGLEAVLPAKPSSQLTDMEIDEC 372
KAINDEGMPMYQRPFMKGKLYIHFTVEFPD+L+ +QVK LEA LPAKPSSQLTDME+DEC
Sbjct: 1 KAINDEGMPMYQRPFMKGKLYIHFTVEFPDSLNPNQVKDLEAALPAKPSSQLTDMELDEC 180
Query: 373 EETTLHDVNMEEENRRKQQQQQQEAYDEDDDMPGGAQRVQCAQQ 416
EETTLHDVNMEEENRRK+QQ QQEAYDEDDDMPGGAQRVQCAQQ
Sbjct: 181 EETTLHDVNMEEENRRKEQQAQQEAYDEDDDMPGGAQRVQCAQQ 312
>TC10179 similar to UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37, partial
(19%)
Length = 518
Score = 133 bits (335), Expect = 5e-32
Identities = 68/82 (82%), Positives = 74/82 (89%), Gaps = 1/82 (1%)
Frame = +3
Query: 336 FTVEFPDTLSLDQVKGLEAVLPAKPSSQLTDMEIDECEETTLHDVNMEEENRRKQQQQQQ 395
FTVEFP++LS +QVK LEAVLP KPSSQLTDME+DECEETTLHDVN+EEE RR+ QQ QQ
Sbjct: 3 FTVEFPESLSAEQVKALEAVLPPKPSSQLTDMELDECEETTLHDVNIEEETRRR-QQAQQ 179
Query: 396 EAYDEDDDM-PGGAQRVQCAQQ 416
EAYDEDDDM GGAQRVQCAQQ
Sbjct: 180 EAYDEDDDMHGGGAQRVQCAQQ 245
>AV767621
Length = 560
Score = 98.2 bits (243), Expect = 2e-21
Identities = 60/139 (43%), Positives = 77/139 (55%), Gaps = 25/139 (17%)
Frame = +1
Query: 3 GRAPKKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEK-----FKELAQA 57
G +++ YY+IL V + AS DDLKKAY+K A+K HPDK + +K FK++++A
Sbjct: 31 GEGEEEAMGVDYYKILQVDRGASDDDLKKAYRKLAMKWHPDKNPNNKKEAEAKFKQISEA 210
Query: 58 YEVLSDPEKREIYDTYGEDALK--------EGMGGGGGGGHDPF--------DIFSSFFG 101
Y+VLSDP+KR +YD YGE+ LK G GG GG F DIFS FFG
Sbjct: 211 YDVLSDPQKRAVYDQYGEEGLKGQVPPPGAGGFPGGSNGGSASFRFNPRSADDIFSEFFG 390
Query: 102 ----GGGGGSSRGRRQRRG 116
GG G GR G
Sbjct: 391 FSSPFGGMGDMGGRAGASG 447
>AV411199
Length = 423
Score = 89.0 bits (219), Expect = 1e-18
Identities = 49/97 (50%), Positives = 61/97 (62%), Gaps = 5/97 (5%)
Frame = +2
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPE-----KFKELAQAYEVLSDPEKRE 68
YYEIL V A+ ++LKKAY++ A+K HPDK D + KFK ++++YEVLSDP+KR
Sbjct: 71 YYEILEVDHHATDEELKKAYRRLAMKWHPDKNPDNKNDAETKFKLISESYEVLSDPQKRA 250
Query: 69 IYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGGGGG 105
IYD YGED LK GM G SFF GGG
Sbjct: 251 IYDRYGEDRLKGGMPSPDAG-------MDSFFRTGGG 340
>BP059356
Length = 436
Score = 56.2 bits (134), Expect(2) = 3e-15
Identities = 32/79 (40%), Positives = 42/79 (52%), Gaps = 16/79 (20%)
Frame = +1
Query: 39 KNHPDKGGDPEKFKELAQAYEVLSDPEKREIYDTYGEDALK--------EGMGGGGGGGH 90
KN +K FK++++AY+VLSDP+KR +YD +GE+ L G GGG GG
Sbjct: 187 KNPNNKKEAEANFKKISEAYDVLSDPQKRAVYDQFGEEGLNGQVPPPGAGGFSGGGDGGP 366
Query: 91 DPF--------DIFSSFFG 101
F DI S FFG
Sbjct: 367 ASFRFNPRSADDILSEFFG 423
Score = 42.0 bits (97), Expect(2) = 3e-15
Identities = 18/36 (50%), Positives = 26/36 (72%)
Frame = +3
Query: 7 KKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHP 42
+K+ YY++L V ++A DDLKKAY+K A+K HP
Sbjct: 75 EKAMGVDYYKVLQVDRSAKDDDLKKAYRKLAMKWHP 182
>TC9416 similar to UP|Q9FEW7 (Q9FEW7) DnaJ like protein, partial (26%)
Length = 558
Score = 73.2 bits (178), Expect = 8e-14
Identities = 44/94 (46%), Positives = 53/94 (55%), Gaps = 2/94 (2%)
Frame = +2
Query: 250 DIVFVLQQKEHPKFKRKSEDLFVEHTLSLTE--ALCGFQFVLTHLDGRQLLIKSNPGEVV 307
DIVF+ +K H F R DL V +SLTE AL GF F LT LDGR L I N G
Sbjct: 2 DIVFIFDEKPHNVFTRDGNDLVVTQKISLTEDEALTGFTFPLTTLDGRCLPIAINNG--T 175
Query: 308 KPDSYKAINDEGMPMYQRPFMKGKLYIHFTVEFP 341
P+ K I EGMP+ + P +G L I F ++FP
Sbjct: 176 DPNYEKVITGEGMPISKDPSKRGNLRIKFNIKFP 277
>TC16999 similar to UP|Q9T024 (Q9T024) DNAJ-like protein
(AT4g39150/T22F8_50), partial (35%)
Length = 648
Score = 69.3 bits (168), Expect = 1e-12
Identities = 42/92 (45%), Positives = 53/92 (56%), Gaps = 4/92 (4%)
Frame = +3
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKG-GDP---EKFKELAQAYEVLSDPEKREI 69
YY++LGV+ AS D+KKAY A HPDK GDP E F+ L +AY+VLSDPEKRE
Sbjct: 129 YYDVLGVNVDASAADIKKAYYIKARIVHPDKNPGDPKAAENFQMLGEAYQVLSDPEKREA 308
Query: 70 YDTYGEDALKEGMGGGGGGGHDPFDIFSSFFG 101
YD G+ + + DP +F FG
Sbjct: 309 YDKNGKAGIPQDT------MLDPTAVFGMLFG 386
>TC16983 similar to UP|Q9ZSY2 (Q9ZSY2) ALTERED response to gravity (ARG1
protein), partial (42%)
Length = 716
Score = 68.2 bits (165), Expect = 2e-12
Identities = 34/68 (50%), Positives = 48/68 (70%), Gaps = 4/68 (5%)
Frame = +2
Query: 15 YEILGVSKTASQDDLKKAYKKAAIKNHPDKG-GDPEK---FKELAQAYEVLSDPEKREIY 70
YE+L VS+ ++ ++K AY+K A+K HPDK +PE FKE+A +Y +LSDPEKR Y
Sbjct: 245 YEVLSVSRDSTDQEIKTAYRKLALKYHPDKNVNNPEASELFKEVAYSYSILSDPEKRRQY 424
Query: 71 DTYGEDAL 78
D+ G +AL
Sbjct: 425 DSAGFEAL 448
>TC9507 similar to UP|Q8VZB6 (Q8VZB6) DnaJ-like protein, partial (40%)
Length = 905
Score = 65.5 bits (158), Expect = 2e-11
Identities = 39/107 (36%), Positives = 59/107 (54%), Gaps = 17/107 (15%)
Frame = +1
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKG---GDPEKFKELAQAYEVLSDPEKREIY 70
YYEILG+ K+ + +D++K+Y+K ++K HPDK G E FK +++A++ L + E + Y
Sbjct: 520 YYEILGLEKSCTVEDVRKSYRKLSLKVHPDKNKAPGAEEAFKAVSKAFQCLGNEESKRKY 699
Query: 71 DTYGED--------ALKEGMG------GGGGGGHDPFDIFSSFFGGG 103
D GED A + G G G D +IF +FF GG
Sbjct: 700 DLSGEDESVYERRAAARPGPGAARAYNGYYEADFDADEIFRNFFFGG 840
>AV774294
Length = 498
Score = 65.5 bits (158), Expect = 2e-11
Identities = 35/63 (55%), Positives = 45/63 (70%), Gaps = 2/63 (3%)
Frame = -1
Query: 355 VLPAKPSSQLTDMEIDECEETTLHDVN-MEEENRRKQQQQQQEAYDED-DDMPGGAQRVQ 412
+LP PSS+ DME+D+CEET HDVN +EEE RR++Q + +EAYDED DD RV
Sbjct: 495 ILPL-PSSKKADMELDDCEETIFHDVNHLEEEMRRRKQPRSREAYDEDEDDDEPSTPRVP 319
Query: 413 CAQ 415
CA+
Sbjct: 318 CAR 310
>TC13524 similar to UP|O48678 (O48678) F3I6.4 protein, partial (19%)
Length = 485
Score = 64.7 bits (156), Expect = 3e-11
Identities = 29/62 (46%), Positives = 44/62 (70%), Gaps = 4/62 (6%)
Frame = +1
Query: 15 YEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEK----FKELAQAYEVLSDPEKREIY 70
YE+LGVS++++ ++K AY+K A+K HPDK + K FKE+ +Y +LSDP+KR Y
Sbjct: 298 YEVLGVSRSSTDQEIKTAYRKMALKYHPDKNANDPKAADMFKEVTFSYNILSDPDKRRQY 477
Query: 71 DT 72
D+
Sbjct: 478 DS 483
>AV775648
Length = 371
Score = 60.5 bits (145), Expect = 5e-10
Identities = 29/65 (44%), Positives = 42/65 (64%), Gaps = 3/65 (4%)
Frame = +1
Query: 10 DSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFK---ELAQAYEVLSDPEK 66
D Y+++GVS+ A+ ++KKAY K ++K+HPD DPE K ++A AYE+L D
Sbjct: 130 DEDDCYDLVGVSQNANSSEIKKAYYKLSLKHHPDTNPDPESKKLLVKVANAYEILKDEAT 309
Query: 67 REIYD 71
RE YD
Sbjct: 310 REQYD 324
>BP039305
Length = 555
Score = 57.8 bits (138), Expect = 3e-09
Identities = 32/73 (43%), Positives = 46/73 (62%), Gaps = 8/73 (10%)
Frame = +3
Query: 3 GRAPKKSD-----STRYYEILGVSKTASQDDLKKAYKKAAIKNHPD---KGGDPEKFKEL 54
GR ++S S YY LGV K+A+ ++K AY++ A + HPD + G +KFKE+
Sbjct: 336 GRTTRRSHTVFAASADYYNTLGVPKSATVKEIKAAYRRLARQYHPDVYKEPGATDKFKEI 515
Query: 55 AQAYEVLSDPEKR 67
+ AYEVLSD +KR
Sbjct: 516 SNAYEVLSDDKKR 554
>TC12876
Length = 820
Score = 55.5 bits (132), Expect = 2e-08
Identities = 31/82 (37%), Positives = 48/82 (57%), Gaps = 11/82 (13%)
Frame = -1
Query: 1 MFGRAPKKSD-STRYYEILGVSKTASQDDLKKAYKKAAIKNHPDK---GGD-------PE 49
+ GR D S +Y +LG++K ++ +L+ AYKK A+K HPD+ G+ +
Sbjct: 274 ILGRMANGGDKSNDFYAVLGLNKECTESELRNAYKKLALKWHPDRCSASGNLKFVEEAKK 95
Query: 50 KFKELAQAYEVLSDPEKREIYD 71
KF+ + +AY VLSD KR +YD
Sbjct: 94 KFQSIQEAYSVLSDANKRLMYD 29
>CN825120
Length = 667
Score = 55.1 bits (131), Expect = 2e-08
Identities = 28/66 (42%), Positives = 37/66 (55%), Gaps = 6/66 (9%)
Frame = +3
Query: 12 TRYYEILGVSKTASQDDLKKAYKKAAIKNHPD------KGGDPEKFKELAQAYEVLSDPE 65
T YEILG++ AS ++K AY++ A HPD K +F ++ AY LSDPE
Sbjct: 201 TSLYEILGIAAAASDQEIKAAYRRLARVRHPDVAAVDRKDSSTNEFMKIHAAYSTLSDPE 380
Query: 66 KREIYD 71
KR YD
Sbjct: 381 KRASYD 398
>BG662268
Length = 298
Score = 53.9 bits (128), Expect = 5e-08
Identities = 36/109 (33%), Positives = 53/109 (48%), Gaps = 5/109 (4%)
Frame = +1
Query: 50 KFKELAQAYEVLSDPEKREIYDTYGEDALKEGMGGGGGGGHDPFD-----IFSSFFGGGG 104
KF+E++ AYEVL D KR+ YD G DA GG G +PF+ +F+ FGG
Sbjct: 16 KFQEVSIAYEVLKDEVKRQEYDQVGHDAYINPQAGGFEG--NPFEDFVKGMFNQKFGG-- 183
Query: 105 GGSSRGRRQRRGEDVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKG 153
+DV +++S + G +K L+ +V C+ C G G
Sbjct: 184 ------------DDVKTSIELSFMEAVQGCAKTLTYQTDVRCNTCGGSG 294
>TC14831 homologue to UP|P93499 (P93499) DnaJ-like protein (Fragment),
partial (63%)
Length = 851
Score = 52.4 bits (124), Expect = 1e-07
Identities = 28/69 (40%), Positives = 37/69 (53%), Gaps = 6/69 (8%)
Frame = +3
Query: 9 SDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPD------KGGDPEKFKELAQAYEVLS 62
S + YEILG+ AS ++K AY++ A HPD K + F ++ AY LS
Sbjct: 231 SPCSSLYEILGIPAGASSQEIKAAYRRLARVCHPDVAAIDRKNSSADDFMKIHAAYSTLS 410
Query: 63 DPEKREIYD 71
DPEKR YD
Sbjct: 411 DPEKRATYD 437
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.134 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,492,927
Number of Sequences: 28460
Number of extensions: 91195
Number of successful extensions: 1673
Number of sequences better than 10.0: 180
Number of HSP's better than 10.0 without gapping: 1031
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1365
length of query: 416
length of database: 4,897,600
effective HSP length: 93
effective length of query: 323
effective length of database: 2,250,820
effective search space: 727014860
effective search space used: 727014860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC147405.6


