

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
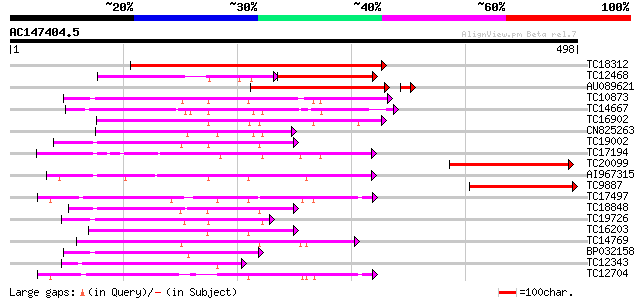
Query= AC147404.5 + phase: 0
(498 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC18312 similar to UP|Q9FIL1 (Q9FIL1) Protein kinase-like protei... 344 2e-95
TC12468 similar to UP|Q944A7 (Q944A7) AT4g35230/F23E12_210, part... 126 6e-60
AU089621 166 4e-42
TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partia... 159 8e-40
TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, com... 121 2e-28
TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-... 118 2e-27
CN825263 116 1e-26
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 114 4e-26
TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete 111 3e-25
TC20099 similar to UP|Q9FIL1 (Q9FIL1) Protein kinase-like protei... 109 1e-24
AI967315 107 6e-24
TC9887 similar to UP|Q944A7 (Q944A7) AT4g35230/F23E12_210, parti... 106 1e-23
TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2,... 105 1e-23
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 104 4e-23
TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase... 103 8e-23
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 100 4e-22
TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete 100 5e-22
BP032158 100 7e-22
TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine prote... 98 3e-21
TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3,... 96 1e-20
>TC18312 similar to UP|Q9FIL1 (Q9FIL1) Protein kinase-like protein
(AT5g59010/k19m22_210), partial (49%)
Length = 746
Score = 344 bits (883), Expect = 2e-95
Identities = 159/225 (70%), Positives = 198/225 (87%)
Frame = +1
Query: 107 EAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDKQPLPWEMRVRVAY 166
EA VG++R +R+ NL+GCC EG+ERLLVAE+MP +TLS+HLFHW+ QP+ W MR+RVA
Sbjct: 28 EARAVGQLRSERLANLVGCCCEGEERLLVAEFMPYETLSRHLFHWETQPMKWAMRLRVAL 207
Query: 167 HVAQALDHCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMKNSRDGKSYSTNLAYTPPE 226
++AQAL++CS++ R +YHDLNAYRILFD+DG+PRLS FGLMKNSRDG+SYSTNLA+TPPE
Sbjct: 208 YLAQALEYCSIKGRALYHDLNAYRILFDQDGNPRLSCFGLMKNSRDGRSYSTNLAFTPPE 387
Query: 227 FLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSHALDLIRGKNALLLMDSSLEGQYANDDA 286
+LR GRI AESV+YS+GT+LLDLLSGKHIPPSHALDLIRGKN L+LMDS LEG ++NDD
Sbjct: 388 YLRNGRITAESVVYSFGTLLLDLLSGKHIPPSHALDLIRGKNFLMLMDSGLEGHFSNDDR 567
Query: 287 TKLVELASKCLQFEARERPDIKFLLTAVTPLQKQKEVASHVLMGL 331
T+LV LAS+CLQ+E RERP+ K L+TA+TPLQK+ V S VL+G+
Sbjct: 568 TELVRLASRCLQYEPRERPNTKTLVTALTPLQKETSVPSSVLLGI 702
>TC12468 similar to UP|Q944A7 (Q944A7) AT4g35230/F23E12_210, partial (47%)
Length = 865
Score = 126 bits (317), Expect(2) = 6e-60
Identities = 60/88 (68%), Positives = 76/88 (86%)
Frame = +2
Query: 236 ESVIYSYGTVLLDLLSGKHIPPSHALDLIRGKNALLLMDSSLEGQYANDDATKLVELASK 295
+SVIYS+GTVLLDLLSGKHIPPSHALD+I+GKN +LLMDS LEG+++ ++AT +V LASK
Sbjct: 488 KSVIYSFGTVLLDLLSGKHIPPSHALDMIQGKNIMLLMDSHLEGKFSTEEATVVVNLASK 667
Query: 296 CLQFEARERPDIKFLLTAVTPLQKQKEV 323
CLQ+E RERPD K L+T +TPL + +V
Sbjct: 668 CLQYEPRERPDTKDLVTTLTPLHTKPDV 751
Score = 121 bits (304), Expect(2) = 6e-60
Identities = 75/178 (42%), Positives = 93/178 (52%), Gaps = 19/178 (10%)
Frame = +1
Query: 78 YKGKLENNRLVAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAE 137
YKG+L N R +AVK+FSK +WPD +QF+ EA+GVGK+RH R+ NLIG +GDERLLVAE
Sbjct: 10 YKGRLHNRRWIAVKKFSKAAWPDPKQFVEEASGVGKLRHPRLANLIGYSCDGDERLLVAE 189
Query: 138 YMPNDTLSKHLFHWDKQPLPWEMRVRVAYHVAQALDH----CSMENRKIYHDLNAYRILF 193
YMPNDTL+KHLFHW K DH C+ E I L +L
Sbjct: 190 YMPNDTLAKHLFHWXKS------------------DH*SGPCAQE*LSILLRLYIIALLR 315
Query: 194 DEDGDPR---LSSFGLMKNSR------------DGKSYSTNLAYTPPEFLRTGRIIAE 236
++ F + K R K AYTPPE+LR GR+ E
Sbjct: 316 VAPYTMT*MLIAFFSMRKVIRVFPVLV**XTAGMAKVIXPTWAYTPPEYLRNGRVTPE 489
>AU089621
Length = 460
Score = 166 bits (421), Expect(2) = 4e-42
Identities = 78/122 (63%), Positives = 101/122 (81%)
Frame = +2
Query: 212 DGKSYSTNLAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSHALDLIRGKNALL 271
DGKSYSTNLAYTPPE+LR GR+ ES+I+S+GTVLLDLLSGKHIPP+HALD+IRGKN L
Sbjct: 5 DGKSYSTNLAYTPPEYLRNGRVTPESIIFSFGTVLLDLLSGKHIPPTHALDMIRGKNIQL 184
Query: 272 LMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTAVTPLQKQKEVASHVLMGL 331
LMDS LEG ++ ++A + +LASKCLQ+E RERP K L+T + LQ +++V S+V++G+
Sbjct: 185 LMDSHLEGNFSTEEAIVVFDLASKCLQYEPRERPKTKDLVTTLLALQTKQDVPSYVMLGI 364
Query: 332 TK 333
K
Sbjct: 365 AK 370
Score = 21.9 bits (45), Expect(2) = 4e-42
Identities = 8/13 (61%), Positives = 11/13 (84%)
Frame = +1
Query: 344 LSPLGKACARMDL 356
LSP+G AC+R+ L
Sbjct: 403 LSPMGDACSRIGL 441
>TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partial (77%)
Length = 1478
Score = 159 bits (403), Expect = 8e-40
Identities = 104/316 (32%), Positives = 160/316 (49%), Gaps = 27/316 (8%)
Frame = +1
Query: 48 YGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQQFLAE 107
+G EL AT F ++ E G VYKG+L VAVK+ S Q+F+ E
Sbjct: 415 FGFRELADATRNFKEANLIGEGGF---GKVYKGRLTTGEAVAVKQLSHDGRQGFQEFVME 585
Query: 108 AAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFH--WDKQPLPWEMRVRVA 165
+ + H +V LIG C +GD+RLLV EYMP +L HLF DK+PL W R++VA
Sbjct: 586 VLMLSLLHHTNLVRLIGYCTDGDQRLLVYEYMPMGSLEDHLFELSHDKEPLNWSTRMKVA 765
Query: 166 YHVAQALD--HCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMK------NSRDGKSYS 217
A+ L+ HC+ + IY DL + IL D + +P+LS FGL K N+
Sbjct: 766 VGAARGLEYLHCTADPPVIYRDLKSANILLDNEFNPKLSDFGLAKLGPVGDNTHVSTRVM 945
Query: 218 TNLAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSHALDLIR--GKNALL---- 271
Y PE+ +G++ +S IYS+G VLL+LL+G+ A+D R G+ L+
Sbjct: 946 GTYGYCAPEYAMSGKLTLKSDIYSFGVVLLELLTGR-----RAIDTSRRPGEQNLVSWAR 1110
Query: 272 -----------LMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTAVTPLQKQ 320
++D L+G++ + + + + + CLQ + + RP I ++ A+ L Q
Sbjct: 1111PYFSDRRRFGHMVDPLLQGRFPSRCLHQAIAITAMCLQEQPKFRPLITDIVVALEYLASQ 1290
Query: 321 KEVASHVLMGLTKTPA 336
+H L G+ + P+
Sbjct: 1291GSPEAHYLYGVQQPPS 1338
>TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1591
Score = 121 bits (304), Expect = 2e-28
Identities = 94/323 (29%), Positives = 157/323 (48%), Gaps = 31/323 (9%)
Frame = +2
Query: 50 LNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQ-QFLAEA 108
L+EL+ T F ++ ++ GE + VY L + VAVK+ S P+ +FL +
Sbjct: 329 LDELKEKTDNFGSKALI---GEGSYGRVYYATLNDGNAVAVKKLDVSSEPETNNEFLTQV 499
Query: 109 AGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDK-----QPLP---WEM 160
+ V ++++ V L G C EG+ R+L E+ +L + H K QP P W
Sbjct: 500 SMVSRLKNDNFVELHGYCVEGNLRVLAYEFATMGSLHD-ILHGRKGVQGAQPGPTLDWIQ 676
Query: 161 RVRVAYHVAQALD--HCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMKNSRD--GKSY 216
RVR+A A+ L+ H ++ I+ D+ + +L ED +++ F L + D + +
Sbjct: 677 RVRIAVDAARGLEYLHEKVQPAIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAARLH 856
Query: 217 STNL----AYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSHALDLIRGKNALLL 272
ST + Y PE+ TG++ +S +YS+G VLL+LL+G+ P H + RG+ +L+
Sbjct: 857 STRVLGTFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRK-PVDHTMP--RGQQSLVT 1027
Query: 273 --------------MDSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTAVTPLQ 318
+D L+G+Y KL +A+ C+Q+EA RP++ ++ A+ P
Sbjct: 1028WATPRLSEDKVKQCVDPKLKGEYPPKGVAKLAAVAALCVQYEAEFRPNMSIVVKALQP-- 1201
Query: 319 KQKEVASHVLMGLTKTPAVLPLP 341
L KTPA P P
Sbjct: 1202------------LLKTPAAAPAP 1234
>TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-like
protein, partial (46%)
Length = 941
Score = 118 bits (296), Expect = 2e-27
Identities = 84/277 (30%), Positives = 136/277 (48%), Gaps = 22/277 (7%)
Frame = +3
Query: 77 VYKGKLENNRLVAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVA 136
VY G+++ VA+KR + S +F E + K+RH+ +V+LIG C E E +LV
Sbjct: 39 VYYGEVDGGTKVAIKRGNPLSEQGVHEFQTEIEMLSKLRHRHLVSLIGYCEENTEMILVY 218
Query: 137 EYMPNDTLSKHLFHWDKQPLPWEMRVRVAYHVAQALD--HCSMENRKIYHDLNAYRILFD 194
++M TL +HL+ K PLPW+ R+ + A+ L H + I+ D+ IL D
Sbjct: 219 DHMAYGTLREHLYKTQKPPLPWKQRLEICIGAARGLHYLHTGAKYTIIHRDVKTTNILLD 398
Query: 195 EDGDPRLSSFGLMKN--SRDGKSYST----NLAYTPPEFLRTGRIIAESVIYSYGTVLLD 248
E ++S FGL K + D ST + Y PE+ R ++ +S +YS+G VL +
Sbjct: 399 EKWVAKVSDFGLSKTGPTLDNTHVSTVVKGSFGYLDPEYFRRQQLTDKSDVYSFGVVLFE 578
Query: 249 LLSGK-HIPPSHALDLIR---------GKNAL-LLMDSSLEGQYANDDATKLVELASKCL 297
+L + + PS A + + K L ++D L+G+ A + K E A KC+
Sbjct: 579 ILCARPALNPSLAKEQVSLAEWASHCYNKGILDQILDPYLKGKIAPECFKKFAETAMKCV 758
Query: 298 QFEARERP---DIKFLLTAVTPLQKQKEVASHVLMGL 331
+ ERP D+ + L LQ+ E + + G+
Sbjct: 759 SDQGIERPSMGDVLWNLEFALQLQESAEESGNGFGGI 869
>CN825263
Length = 663
Score = 116 bits (290), Expect = 1e-26
Identities = 67/187 (35%), Positives = 105/187 (55%), Gaps = 10/187 (5%)
Frame = +1
Query: 76 VVYKGKLENNRLVAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLV 135
+VYKG L + R VAVK + ++FLAE + ++ H+ +V LIG C E R L+
Sbjct: 10 LVYKGILNDGRDVAVKILKRDDQRGGREFLAEVEMLSRLHHRNLVKLIGICIEKQTRCLI 189
Query: 136 AEYMPNDTLSKHLFHWDKQ--PLPWEMRVRVAYHVAQALDHCSMENRK--IYHDLNAYRI 191
E +PN ++ HL DK+ PL W R+++A A+ L + ++ I+ D + I
Sbjct: 190 YELVPNGSVESHLHGADKETGPLDWNARMKIALGAARGLAYLHEDSNPCVIHRDFKSSNI 369
Query: 192 LFDEDGDPRLSSFGLMKNSRD--GKSYSTNL----AYTPPEFLRTGRIIAESVIYSYGTV 245
L + D P++S FGL + + D K ST++ Y PE+ TG ++ +S +YSYG V
Sbjct: 370 LLECDFTPKVSDFGLARTALDEGNKHISTHVMGTFGYLAPEYAMTGHLLVKSDVYSYGVV 549
Query: 246 LLDLLSG 252
LL+LL+G
Sbjct: 550 LLELLTG 570
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 114 bits (285), Expect = 4e-26
Identities = 74/224 (33%), Positives = 115/224 (51%), Gaps = 9/224 (4%)
Frame = +1
Query: 39 ESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSW 98
+ + P ++ + L EL AT+ F+ + + E G + VY G+L + +AVKR S
Sbjct: 1 QKKQPAWRVFSLKELHSATNNFNYDNKLGEGGFGS---VYWGQLWDGSQIAVKRLKVWSN 171
Query: 99 PDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLF--HWDKQPL 156
+F E + +VRHK +++L G CAEG ERL+V +YMPN +L HL H + L
Sbjct: 172 KADMEFAVEVEILARVRHKNLLSLRGYCAEGQERLIVYDYMPNLSLLSHLHGQHSSECLL 351
Query: 157 PWEMRVRVAYHVAQALD--HCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMKNSRDGK 214
W R+ +A A+ + H I+ D+ A +L D D R++ FG K DG
Sbjct: 352 DWNRRMNIAIGSAEGIVYLHHQATPHIIHRDIKASNVLLDSDFQARVADFGFAKLIPDGA 531
Query: 215 SYST-----NLAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGK 253
++ T L Y PE+ G+ ++S+G +LL+L SGK
Sbjct: 532 THVTTRVKGTLGYLAPEYAMLGKANECCDVFSFGILLLELASGK 663
>TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete
Length = 2193
Score = 111 bits (277), Expect = 3e-25
Identities = 91/323 (28%), Positives = 145/323 (44%), Gaps = 24/323 (7%)
Frame = +1
Query: 24 SKKTDPGDDGGDGDQESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLE 83
S + PG G V E+ EL +AT+ FS + + + G A VY +L
Sbjct: 979 SGSSGPGTASATGLTSIMVAKSMEFSYQELAKATNNFSLDNKIGQGGFGA---VYYAELR 1149
Query: 84 NNRLVAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDT 143
+ A+K+ Q+ + +FL E + V H +V LIG C EG LV E++ N
Sbjct: 1150 GKK-TAIKKMDVQA---STEFLCELKVLTHVHHLNLVRLIGYCVEGS-LFLVYEHIDNGN 1314
Query: 144 LSKHLFHWDKQPLPWEMRVRVAYHVAQALDHCSMENRKIY--HDLNAYRILFDEDGDPRL 201
L ++L K+PLPW RV++A A+ L++ +Y D+ + IL D++ ++
Sbjct: 1315 LGQYLHGSGKEPLPWSSRVQIALDAARGLEYIHEHTVPVYIHRDVKSANILIDKNLRGKV 1494
Query: 202 SSFGLMKNSRDGKS-YSTNL----AYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKH-- 254
+ FGL K G S T L Y PPE+ + G I + +Y++G VL +L+S K+
Sbjct: 1495 ADFGLTKLIEVGNSTLQTRLVGTFGYMPPEYAQYGDISPKIDVYAFGVVLFELISAKNAV 1674
Query: 255 IPPSHALDLIRGKNALL---------------LMDSSLEGQYANDDATKLVELASKCLQF 299
+ + +G AL L+D L Y D K+ +L C +
Sbjct: 1675 LKTGELVAESKGLVALFEEALNKSDPCDALRKLVDPRLGENYPIDSVLKIAQLGRACTRD 1854
Query: 300 EARERPDIKFLLTAVTPLQKQKE 322
RP ++ L+ A+ L E
Sbjct: 1855 NPLLRPSMRSLVVALMTLSSLTE 1923
>TC20099 similar to UP|Q9FIL1 (Q9FIL1) Protein kinase-like protein
(AT5g59010/k19m22_210), partial (22%)
Length = 498
Score = 109 bits (272), Expect = 1e-24
Identities = 55/109 (50%), Positives = 72/109 (65%)
Frame = +2
Query: 387 VQDILNTKKFGDIAFRDKDFKNAIEYYSKLVVMMSVPSATVFARRAFAYLMNDQAELALR 446
+Q+ LN KK GD AFR KDF AI+ Y++ + ++ S TV+ARR +YLMN + AL
Sbjct: 2 MQETLNLKKHGDTAFRAKDFATAIKCYTQFIDGGTMVSPTVYARRCLSYLMNGMTQEALG 181
Query: 447 DAMQAQVCIPDWPTAFYLQALALSKLGMETDAQDMLNDGAAFEAKRSNS 495
DAMQAQ P+WPTA YLQA L LGME DA + L DG EA+++ +
Sbjct: 182 DAMQAQAVSPEWPTALYLQAACLFSLGMENDAPETLKDGTNMEAQKNQN 328
>AI967315
Length = 1308
Score = 107 bits (266), Expect = 6e-24
Identities = 86/317 (27%), Positives = 146/317 (45%), Gaps = 27/317 (8%)
Frame = +1
Query: 33 GGDGDQESQV----------PVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKL 82
GG+ D S V P +K + EL AT+ FS+E +V + G VYKG+L
Sbjct: 37 GGENDDSSLVVPPYEEHPPRPSWKCFSYEELFHATNGFSSENMVGKGGYAE---VYKGRL 207
Query: 83 ENNRLVAVKRFSKQSWPD--AQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMP 140
E+ +AVKR ++ + ++FL E +G V H ++ L+GCC + + LV E
Sbjct: 208 ESGDEIAVKRLTRTCRDERKEKEFLTEIGTIGHVCHSNVMPLLGCCID-NGLYLVFELST 384
Query: 141 NDTLSKHLFHWDKQPLPWEMRVRVAYHVAQALD--HCSMENRKIYHDLNAYRILFDEDGD 198
+++ + PL W+ R ++ A+ L H + R I+ D+ A IL ED +
Sbjct: 385 VGSVASLIHDEKMAPLDWKTRYKIVLGTARGLHYLHKGCQRRIIHRDIKASNILLTEDFE 564
Query: 199 PRLSSFGLMK------NSRDGKSYSTNLAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSG 252
P++S FGL K + PE+ G + ++ ++++G LL+++SG
Sbjct: 565 PQISDFGLAKWLPSQWTHHSIAPIEGTFGHLAPEYYMHGVVDEKTDVFAFGVFLLEVISG 744
Query: 253 -KHIPPSH------ALDLIRGKNALLLMDSSLEGQYANDDATKLVELASKCLQFEARERP 305
K + SH A ++ L+D LEG Y ++ AS C++ + RP
Sbjct: 745 RKPVDGSHQSLHTWAKPILSKWEIEKLVDPRLEGCYDVTQFNRVAFAASLCIRASSTWRP 924
Query: 306 DIKFLLTAVTPLQKQKE 322
+ +L + + KE
Sbjct: 925 TMSEVLEVMEEGEMDKE 975
>TC9887 similar to UP|Q944A7 (Q944A7) AT4g35230/F23E12_210, partial (19%)
Length = 614
Score = 106 bits (264), Expect = 1e-23
Identities = 52/94 (55%), Positives = 67/94 (70%)
Frame = +2
Query: 405 DFKNAIEYYSKLVVMMSVPSATVFARRAFAYLMNDQAELALRDAMQAQVCIPDWPTAFYL 464
DFK AI+ YS+ + + ++ S TVFARR+ YL+ DQ + ALRDAMQAQ PDWPTAFY+
Sbjct: 2 DFKTAIDNYSQFIDVGTMVSPTVFARRSLCYLLCDQPDPALRDAMQAQCVYPDWPTAFYM 181
Query: 465 QALALSKLGMETDAQDMLNDGAAFEAKRSNSWRG 498
Q++AL+KL M DA DMLN+ A E K+ RG
Sbjct: 182 QSVALAKLDMHKDAADMLNEATALEEKKQRGARG 283
>TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2, complete
Length = 2946
Score = 105 bits (263), Expect = 1e-23
Identities = 87/329 (26%), Positives = 152/329 (45%), Gaps = 30/329 (9%)
Frame = +1
Query: 25 KKTDPGDDGG--------DGDQESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNV 76
KK GD+G D + + + + + + AT+ FS + ++ GE
Sbjct: 1543 KKNKRGDEGEIGIINHWKDKRGDEDIDLATIFDFSTISSATNHFS---LSNKLGEGGFGP 1713
Query: 77 VYKGKLENNRLVAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVA 136
VYKG L N + +AVKR S S ++F E + +++H+ +V L GC DE
Sbjct: 1714 VYKGLLANGQEIAVKRLSNTSGQGMEEFKNEIKLIARLQHRNLVKLFGCSVHQDENSHAN 1893
Query: 137 EYMP--NDTLSKHLFHWDKQPLPWEMRVRVAYHVAQALDHCSMENRK--IYHDLNAYRIL 192
+ M D+ L W+K R+++ +A+ L + ++R I+ DL IL
Sbjct: 1894 KKMKILLDSTRSKLVDWNK-------RLQIIDGIARGLLYLHQDSRLRIIHRDLKTSNIL 2052
Query: 193 FDEDGDPRLSSFGLMK-------NSRDGKSYSTNLAYTPPEFLRTGRIIAESVIYSYGTV 245
D++ +P++S FGL + +R + T Y PPE+ G +S ++S+G +
Sbjct: 2053 LDDEMNPKISDFGLARIFIGDQVEARTKRVMGT-YGYMPPEYAVHGSFSIKSDVFSFGVI 2229
Query: 246 LLDLLSGKHI----PPSHALDLIR-------GKNALLLMDSSLEGQYANDDATKLVELAS 294
+L+++SGK I P H L+L+ + L L+D L+ + + + +A
Sbjct: 2230 VLEIISGKKIGRFYDPHHHLNLLSHAWRLWIEERPLELVDELLDDPVIPTEILRYIHVAL 2409
Query: 295 KCLQFEARERPDIKFLLTAVTPLQKQKEV 323
C+Q RPD +L+ V L +KE+
Sbjct: 2410 LCVQRRPENRPD---MLSIVLMLNGEKEL 2487
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 104 bits (259), Expect = 4e-23
Identities = 67/212 (31%), Positives = 111/212 (51%), Gaps = 10/212 (4%)
Frame = +2
Query: 52 ELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQQFLAEAAGV 111
EL AT FS + + E G + VY G+ + +AVK+ + +F E +
Sbjct: 278 ELHAATGGFSDDNKLGEGGFGS---VYWGRTSDGLQIAVKKLKAMNSKAEMEFAVEVEVL 448
Query: 112 GKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHL---FHWDKQPLPWEMRVRVAYHV 168
G+VRHK ++ L G C D+RL+V +YMPN +L HL F + Q L W+ R+++A
Sbjct: 449 GRVRHKNLLGLRGYCVGDDQRLIVYDYMPNLSLLSHLHGQFAVEVQ-LNWQKRMKIAIGS 625
Query: 169 AQAL--DHCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMKNSRDGKSYST-----NLA 221
A+ + H + I+ D+ A +L + D +P ++ FG K +G S+ T L
Sbjct: 626 AEGILYLHHEVTPHIIHRDIKASNVLLNSDFEPLVADFGFAKLIPEGVSHMTTRVKGTLG 805
Query: 222 YTPPEFLRTGRIIAESVIYSYGTVLLDLLSGK 253
Y PE+ G++ +YS+G +LL+L++G+
Sbjct: 806 YLAPEYAMWGKVSESCDVYSFGILLLELVTGR 901
>TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase, partial
(30%)
Length = 728
Score = 103 bits (256), Expect = 8e-23
Identities = 65/201 (32%), Positives = 102/201 (50%), Gaps = 14/201 (6%)
Frame = +3
Query: 46 KEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRL-VAVKRFSKQSWPDAQQF 104
+E+ EL++AT+ F ++ + + G VVY+G L +L VAVK FS+ F
Sbjct: 126 REFNYVELKKATNNFDEKHKLGQGGY---GVVYRGMLPKEKLEVAVKMFSRDKMKSTDDF 296
Query: 105 LAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHWD----KQPLPWEM 160
L+E + ++RHK +V L G C + LLV +YMPN +L H+F + PL W +
Sbjct: 297 LSELIIINRLRHKHLVRLQGWCHKNGVLLLVYDYMPNGSLDSHIFCEEGGTITTPLSWPL 476
Query: 161 RVRVAYHVAQALD--HCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMKNSRDGKSYST 218
R ++ VA AL+ H + + ++ DL A I+ D + + +L FGL + + K T
Sbjct: 477 RYKIISGVASALNYLHNEYDQKVVHRDLKASNIMLDSEFNAKLGDFGLARALENEKISYT 656
Query: 219 NL-------AYTPPEFLRTGR 232
L Y PE TG+
Sbjct: 657 ELEGVHGTMGYIAPECFHTGK 719
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 100 bits (250), Expect = 4e-22
Identities = 64/193 (33%), Positives = 101/193 (52%), Gaps = 9/193 (4%)
Frame = +2
Query: 70 GEKAPNVVYKGKLENNRLVAVKRFSKQ-SWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAE 128
G+ +VY+G + N VA+KR Q S + F AE +GK+RH+ ++ L+G +
Sbjct: 2213 GKGGAGIVYRGSMPNGTDVAIKRLVGQGSGRNDYGFRAEIETLGKIRHRNIMRLLGYVSN 2392
Query: 129 GDERLLVAEYMPNDTLSKHLFHWDKQPLPWEMRVRVAYHVAQAL--DHCSMENRKIYHDL 186
D LL+ EYMPN +L + L L WEMR ++A A+ L H I+ D+
Sbjct: 2393 KDTNLLLYEYMPNGSLGEWLHGAKGGHLRWEMRYKIAVEAARGLCYMHHDCSPLIIHRDV 2572
Query: 187 NAYRILFDEDGDPRLSSFGLMK------NSRDGKSYSTNLAYTPPEFLRTGRIIAESVIY 240
+ IL D D + ++ FGL K S+ S + + Y PE+ T ++ +S +Y
Sbjct: 2573 KSNNILLDADFEAHVADFGLAKFLYDPGASQSMSSIAGSYGYIAPEYAYTLKVDEKSDVY 2752
Query: 241 SYGTVLLDLLSGK 253
S+G VLL+L+ G+
Sbjct: 2753 SFGVVLLELIIGR 2791
>TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete
Length = 3495
Score = 100 bits (249), Expect = 5e-22
Identities = 75/270 (27%), Positives = 136/270 (49%), Gaps = 21/270 (7%)
Frame = +3
Query: 59 EFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQQFLAEAAGVGKVRHKR 118
E +TE + GE VY+G L + + VAVK S S ++F E + ++H+
Sbjct: 2151 EVATERYKTLIGEGGFGSVYRGTLNDGQEVAVKVRSSTSTQGTREFDNELNLLSAIQHEN 2330
Query: 119 MVNLIGCCAEGDERLLVAEYMPNDTLSKHLF--HWDKQPLPWEMRVRVAYHVAQALDHC- 175
+V L+G C E D+++LV +M N +L L+ ++ L W R+ +A A+ L +
Sbjct: 2331 LVPLLGYCNESDQQILVYPFMSNGSLQDRLYGEPAKRKILDWPTRLSIALGAARGLAYLH 2510
Query: 176 SMENRKIYH-DLNAYRILFDEDGDPRLSSFGLMKNS-RDGKSYST-----NLAYTPPEFL 228
+ R + H D+ + IL D +++ FG K + ++G SY + Y PE+
Sbjct: 2511 TFPGRSVIHRDIKSSNILLDHSMCAKVADFGFSKYAPQEGDSYVSLEVRGTAGYLDPEYY 2690
Query: 229 RTGRIIAESVIYSYGTVLLDLLSGKH-----IPPSH------ALDLIRGKNALLLMDSSL 277
+T ++ +S ++S+G VLL+++SG+ P + A IRG ++D +
Sbjct: 2691 KTQQLSEKSDVFSFGVVLLEIVSGREPLNIKRPRTEWSLVEWATPYIRGSKVDEIVDPGI 2870
Query: 278 EGQYANDDATKLVELASKCLQFEARERPDI 307
+G Y + ++VE+A +CL+ + RP +
Sbjct: 2871 KGGYHAEAMWRVVEVALQCLEPFSTYRPSM 2960
>BP032158
Length = 555
Score = 100 bits (248), Expect = 7e-22
Identities = 61/181 (33%), Positives = 101/181 (55%), Gaps = 5/181 (2%)
Frame = +2
Query: 48 YGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQQFLAE 107
+ L +++ AT+ F + E G VYKG L ++AVK+ S +S ++F+ E
Sbjct: 14 FSLRQIKAATNNFDPANKIGEGGF---GPVYKGVLSEGDVIAVKQLSSKSKQGNREFINE 184
Query: 108 AAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDKQP--LPWEMRVRVA 165
+ ++H +V L GCC EG++ LLV EYM N++L++ LF ++Q L W R+++
Sbjct: 185 IGMISALQHPNLVKLYGCCIEGNQLLLVYEYMENNSLARALFGNEEQKLNLNWRTRMKIC 364
Query: 166 YHVAQALDHCSMENR-KIYH-DLNAYRILFDEDGDPRLSSFGLMK-NSRDGKSYSTNLAY 222
+A+ L + E+R KI H D+ A +L D+D ++S FGL K + + ST +A
Sbjct: 365 VGIAKGLAYLHEESRLKIVHRDIKATNVLLDKDLKAKISDFGLAKLDEEENTHISTRIAG 544
Query: 223 T 223
T
Sbjct: 545 T 547
>TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine protein kinase
Cdk9 (Cyclin-dependent kinase Cdk9) , partial (7%)
Length = 723
Score = 98.2 bits (243), Expect = 3e-21
Identities = 60/166 (36%), Positives = 94/166 (56%), Gaps = 3/166 (1%)
Frame = +3
Query: 46 KEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQQFL 105
K + L AT F + V++ GE V+KGKL + R +AVK+ S++S QF+
Sbjct: 201 KTFSYETLVAATKNF---HAVNKLGEGGFGPVFKGKLNDGREIAVKKLSRRSNQGRTQFI 371
Query: 106 AEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFH-WDKQPLPWEMRVRV 164
EA + +V+H+ +V+L G CA G E+LLV EY+P ++L K LF K+ L W+ R +
Sbjct: 372 NEAKLLTRVQHRNVVSLFGYCAHGSEKLLVYEYVPRESLDKLLFRSQKKEQLDWKRRFDI 551
Query: 165 AYHVAQALDHCSMENRK--IYHDLNAYRILFDEDGDPRLSSFGLMK 208
VA+ L + ++ I+ D+ A IL DE P+++ FGL +
Sbjct: 552 ISGVARGLLYLHEDSHDCIIHRDIKAANILLDEKWVPKIADFGLAR 689
>TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3, complete
Length = 2481
Score = 95.9 bits (237), Expect = 1e-20
Identities = 86/350 (24%), Positives = 145/350 (40%), Gaps = 51/350 (14%)
Frame = +1
Query: 25 KKTDPGDDGG----------DGDQESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKAP 74
KK + D+GG D + + + + + + T+ FS ++ GE
Sbjct: 1318 KKNEREDEGGIETRIINHWKDKRGDEDIDLATIFDFSTISSTTNHFSES---NKLGEGGF 1488
Query: 75 NVVYKGKLENNRLVAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLL 134
VYKG L N + +AVKR S S ++F E + +++H+ +V L+GC DE LL
Sbjct: 1489 GPVYKGVLANGQEIAVKRLSNTSGQGMEEFKNEVKLIARLQHRNLVKLLGCSIHHDEMLL 1668
Query: 135 VAEYMPNDTLSKHLFHWDKQPLPWEMRVRVAYHVAQALDHCSMENRKIYHDLNAYRILFD 194
+ E+M N +L +F + R+R+ I+ DL IL D
Sbjct: 1669 IYEFMHNRSLDYFIF---------DSRLRI-----------------IHRDLKTSNILLD 1770
Query: 195 EDGDPRLSSFGLMK------NSRDGKSYSTNLAYTPPEFLRTGRIIAESVIYSYGTVLLD 248
+ +P++S FGL + K Y PE+ G +S ++S+G ++L+
Sbjct: 1771 SEMNPKISDFGLARIFTGDQVEAKTKRVMGTYGYMSPEYAVHGSFSVKSDVFSFGVIVLE 1950
Query: 249 LLSGKHI----PPSH-----------ALDLIRG--------------------KNALLLM 273
++SGK I P H A+ LI+ + L L+
Sbjct: 1951 IISGKKIGRFCDPHHHRNLLSHSSNFAVFLIKALRICMFENVKNRKAWRLWIEERPLELV 2130
Query: 274 DSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTAVTPLQKQKEV 323
D L+G + + + +A C+Q RPD +L+ V L +KE+
Sbjct: 2131 DELLDGLAIPTEILRYIHIALLCVQQRPEYRPD---MLSVVLMLNGEKEL 2271
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.134 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,621,256
Number of Sequences: 28460
Number of extensions: 92680
Number of successful extensions: 744
Number of sequences better than 10.0: 196
Number of HSP's better than 10.0 without gapping: 681
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 686
length of query: 498
length of database: 4,897,600
effective HSP length: 94
effective length of query: 404
effective length of database: 2,222,360
effective search space: 897833440
effective search space used: 897833440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC147404.5


