

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147404.4 - phase: 0
(583 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
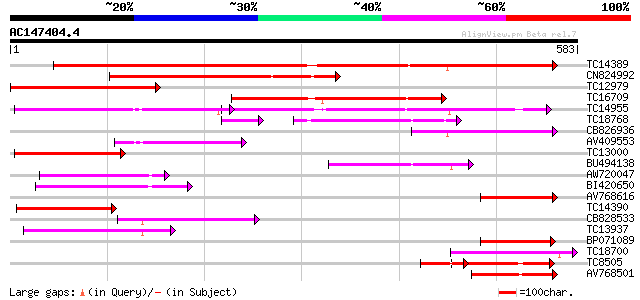
Score E
Sequences producing significant alignments: (bits) Value
TC14389 weakly similar to UP|Q9FNL8 (Q9FNL8) Peptide transporter... 377 e-105
CN824992 317 3e-87
TC12979 similar to UP|Q9LQL2 (Q9LQL2) F5D14.23 protein, partial ... 273 6e-74
TC16709 weakly similar to PIR|T47573|T47573 peptide transport-li... 178 2e-45
TC14955 UP|O22305 (O22305) Peptide transporter, complete 177 4e-45
TC18768 107 3e-26
CB826936 109 1e-24
AV409553 103 8e-23
TC13000 similar to PIR|T47573|T47573 peptide transport-like prot... 102 2e-22
BU494138 97 9e-21
AW720047 97 9e-21
BI420650 94 5e-20
AV768616 92 3e-19
TC14390 weakly similar to UP|Q9FNL8 (Q9FNL8) Peptide transporter... 87 7e-18
CB828533 87 7e-18
TC13937 weakly similar to UP|Q9LVE0 (Q9LVE0) Nitrate transporter... 85 4e-17
BP071089 84 6e-17
TC18700 similar to UP|Q9LYR6 (Q9LYR6) Peptide transporter-like p... 82 2e-16
TC8505 similar to UP|Q8LG02 (Q8LG02) Nitrate transporter, partia... 82 2e-16
AV768501 82 2e-16
>TC14389 weakly similar to UP|Q9FNL8 (Q9FNL8) Peptide transporter, partial
(72%)
Length = 1990
Score = 377 bits (967), Expect = e-105
Identities = 205/527 (38%), Positives = 321/527 (60%), Gaps = 9/527 (1%)
Frame = +3
Query: 46 LAFFGVGVNLVLFLTRVLGQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIF 105
+A++G+ NLV+FL L + A++NNVS W G V+ LVGA+++D+Y GRY T I
Sbjct: 45 MAYYGISSNLVVFLGSKLHEGTVASSNNVSNWAGVVWTTPLVGAYIADAYLGRYWTFIIA 224
Query: 106 QGIFVTGLVSLSVTTYLALLRPKGCGNG--KLECGEHSSLEMGMFYLSIYLIALGNGGYQ 163
I++ G+ L++T L LRP C G +C + S G+FYL++Y+IALG GG +
Sbjct: 225 SCIYLLGMCLLTLTVSLPALRPPPCAQGVENQDCPQASPWTRGIFYLALYIIALGAGGTK 404
Query: 164 PNIATFGADQFDEDHSKESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWAS 223
PNI+T GADQFDE KE+ K++FF+++ ++ +G LFS T+L Y +D W+LG+
Sbjct: 405 PNISTMGADQFDEYEPKENTYKLSFFNWWVFSILVGVLFSTTVLVYIQDNLSWSLGYGLP 584
Query: 224 AGSAFLALVLFLVGTPKYRHFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKE 283
++++FLVGTP YRH P G+PL+R QVF AA K + + L+ + +E
Sbjct: 585 TVGLAFSILVFLVGTPFYRHKLPSGSPLTRMLQVFVAAGIKWKAHVPHDPKQLHELSMEE 764
Query: 284 SSNNSNRKILHTHGFKFLDRAAYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPI 343
+NNS +I HT +FLD+AA K G+ +PW LC +TQVEE K + +++PI
Sbjct: 765 YANNSRNRIDHTSTLRFLDKAAV---------KTGKTSPWRLCTVTQVEETKQMTKMVPI 917
Query: 344 WLCTIIYSVVFTQMASLFVEQGAAMKTTIS-SFKIPPASMSSFDILSVAIFIFFYRRVLD 402
+ T+I + Q +LF++QG + ++ +F+IPPA +S+ I+S+ I Y RV
Sbjct: 918 LITTLIPCTMLIQAHTLFIKQGTTLDRSMGPNFEIPPAGLSAILIISMLTSIPIYDRVFV 1097
Query: 403 PLVGKLKKSSSKGLTELQRMGIGLVIAIIAMVTAGIVECYRLKYAK------QGDTSSLS 456
P++ + K + +G+T LQR+G GL++ ++ MV A + E RL+ A+ Q D LS
Sbjct: 1098PVIRRYTK-NPRGITMLQRLGAGLLMYVLIMVIAWLTERKRLRVAREKHLLGQHDIIPLS 1274
Query: 457 IFWQIPQYALIGASEVFMYVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIV 516
IF IPQYAL G +E F + +++FF Q P+G+KS G + TS++LG ++SS ++S V
Sbjct: 1275IFILIPQYALTGVAENFAEIAKMDFFYYQAPEGMKSLGISYSTTSVALGCFLSSFLLSTV 1454
Query: 517 MKISTEDHMPGWIPGNLNRGHLDRFFFLLAVLTSLDLIAYIACAKWF 563
I+ + GWI NLN LD ++ + +L+ L+ + ++ AK+F
Sbjct: 1455ADITKKHSHQGWILDNLNISRLDYYYAFMVILSFLNFLCFLVTAKFF 1595
>CN824992
Length = 708
Score = 317 bits (813), Expect = 3e-87
Identities = 160/238 (67%), Positives = 190/238 (79%)
Frame = +2
Query: 103 AIFQGIFVTGLVSLSVTTYLALLRPKGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGY 162
AIFQ IFV GL SLS+TTYL LL P GCG+ +L C HSS + +FY+SIYLIALGNGGY
Sbjct: 2 AIFQVIFVVGLASLSLTTYLFLLEPNGCGDKELPCTSHSSYQTALFYVSIYLIALGNGGY 181
Query: 163 QPNIATFGADQFDEDHSKESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWA 222
QPNIATFGADQFDE+ KE +SK++FFSYFYLALNLGSLFSNTIL YFED+GLW LGFWA
Sbjct: 182 QPNIATFGADQFDEEDPKEQHSKISFFSYFYLALNLGSLFSNTILNYFEDDGLWTLGFWA 361
Query: 223 SAGSAFLALVLFLVGTPKYRHFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEK 282
SAGSAFLALVLFL GTP YR+FKP GNP+ RFCQVF AA RK V++ S G++ + + +
Sbjct: 362 SAGSAFLALVLFLCGTPWYRYFKPGGNPVPRFCQVFVAAMRKWKVKV-SPGEEDQLHEVE 538
Query: 283 ESSNNSNRKILHTHGFKFLDRAAYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRL 340
E S N RK+ HT GF+FLD+AA ITS+ E ++ + + W+L +TQVEEVKCILRL
Sbjct: 539 ECSTNGGRKMFHTEGFRFLDKAALITSQ--ESKQENECSLWHLSTVTQVEEVKCILRL 706
>TC12979 similar to UP|Q9LQL2 (Q9LQL2) F5D14.23 protein, partial (25%)
Length = 601
Score = 273 bits (698), Expect = 6e-74
Identities = 133/155 (85%), Positives = 142/155 (90%)
Frame = +3
Query: 1 GKFKEETEELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLT 60
GKFKEE EE+TLDGSVDWHGRP+IRA SGRW AGTIILLNQGLATLAFFGVGVNLVLFLT
Sbjct: 135 GKFKEEAEEVTLDGSVDWHGRPAIRAKSGRWVAGTIILLNQGLATLAFFGVGVNLVLFLT 314
Query: 61 RVLGQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTT 120
RVLGQDNA AANNVSKWTGTVY+FSLVGAFLSDSYWGRYKTCAIFQGIFV GLV LS+++
Sbjct: 315 RVLGQDNADAANNVSKWTGTVYLFSLVGAFLSDSYWGRYKTCAIFQGIFVLGLVFLSLSS 494
Query: 121 YLALLRPKGCGNGKLECGEHSSLEMGMFYLSIYLI 155
YL+LLRPKGCG+ L CG+HSSLEMGMF YLI
Sbjct: 495 YLSLLRPKGCGSELLHCGKHSSLEMGMFSYQSYLI 599
>TC16709 weakly similar to PIR|T47573|T47573 peptide transport-like protein
- Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(41%)
Length = 712
Score = 178 bits (452), Expect = 2e-45
Identities = 103/226 (45%), Positives = 141/226 (61%), Gaps = 5/226 (2%)
Frame = +1
Query: 229 LALVLFLVGTPKYRHFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNS 288
+A+V F GT YR+ KP G+PL+R CQV A+ RK GVQ + LY I + ES+
Sbjct: 19 VAVVSFFSGTRLYRNQKPGGSPLTRMCQVVVASMRKCGVQAPDDKSLLYEIADTESAIKG 198
Query: 289 NRKILHTHGFKFLDRAAYITSRDLEVQKGGQH---NPWYLCPITQVEEVKCILRLLPIWL 345
+RK+ HT+ F D+AA + VQ Q +PW LC TQVEE+K ILR+LP+W
Sbjct: 199 SRKLDHTNELSFFDKAAVV------VQSSDQKDSVDPWRLCTETQVEELKSILRMLPVWA 360
Query: 346 CTIIYSVVFTQMASLFVEQGAAMKTTI--SSFKIPPASMSSFDILSVAIFIFFYRRVLDP 403
II++ V+ QM++LFV QG M T + S+FKIPPA++S FD LSV ++ Y ++ P
Sbjct: 361 TGIIFATVYGQMSTLFVLQGQTMNTHVGNSNFKIPPAALSIFDTLSVIFWVPVYDWIIVP 540
Query: 404 LVGKLKKSSSKGLTELQRMGIGLVIAIIAMVTAGIVECYRLKYAKQ 449
+ K GLT+LQRMGIGL I+I AMV A I+E RL+ ++
Sbjct: 541 IARKF-SGHKNGLTQLQRMGIGLFISIFAMVAAAILEVIRLRMVRR 675
>TC14955 UP|O22305 (O22305) Peptide transporter, complete
Length = 2300
Score = 177 bits (449), Expect = 4e-45
Identities = 112/349 (32%), Positives = 184/349 (52%), Gaps = 9/349 (2%)
Frame = +1
Query: 218 LGFWASAGSAFLALVLFLVGTPKYRH-FKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDL 276
L + SA L+ ++F G+P YRH + +P F +V A R +Q+ N +L
Sbjct: 667 LAYSLSAIGFLLSSIIFFWGSPVYRHKSRQARSPSMNFIRVPLVAFRNRKLQLPCNPSEL 846
Query: 277 YVIDEKESSNNSNRKILHTHGFKFLDRAAYITSR-DLEVQKGGQHNPWYLCPITQVEEVK 335
+ ++ RKI HT F FLDRAA S DL NP C +TQVE K
Sbjct: 847 HEFQLNYYISSGARKIHHTSHFSFLDRAAIRESNTDLS-------NP--PCTVTQVEGTK 999
Query: 336 CILRLLPIWLCTIIYSVVFTQMASLFVEQGAAMKTTIS-SFKIPPASMSSFDILSVAIFI 394
+L + IWL +I + + +++FV QG M T+ F++P AS+ F +L+ I +
Sbjct: 1000LVLGMFQIWLLMLIPTNCWALESTIFVRQGTTMDRTLGPKFRLPAASLWCFIVLTTLICL 1179
Query: 395 FFYRRVLDPLVGKLKKSSSKGLTELQRMGIGLVIAIIAMVTAGIVECYRLKYAKQG---- 450
Y P + + + + +G+ LQR+GIG+ I +IAM VE R+ K+
Sbjct: 1180PIYDHYFIPFMRR-RTGNHRGIKLLQRVGIGMAIQVIAMAVTYAVETQRMSVIKKHHIVG 1356
Query: 451 --DTSSLSIFWQIPQYALIGASEVFMYVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYV 508
+T +SIFW +PQ ++G S F+ G LEFF Q+P+ +K G+ LC + ++ G+Y+
Sbjct: 1357PEETVPMSIFWLLPQNIILGVSNAFLATGMLEFFYDQSPEEMKGLGTTLCTSCVAAGSYI 1536
Query: 509 SSLIVSIVMKISTEDHMPGWIPGNLNRGHLDRFFFLLAVLTSLDLIAYI 557
++ +V+++ K++ WI NLN HLD ++ L V+++L+ ++
Sbjct: 1537NTFLVTMIDKLN-------WIGNNLNDSHLDYYYAFLFVISALNFGVFL 1662
Score = 152 bits (385), Expect = 1e-37
Identities = 87/231 (37%), Positives = 138/231 (59%), Gaps = 5/231 (2%)
Frame = +3
Query: 6 ETEELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQ 65
E + TLDG+VD GRP + + +G+ A T IL+ + L A++GVG NLV F+T L +
Sbjct: 39 EGKGYTLDGTVDLAGRPVLSSLTGKQKACTYILVYRVLERFAYYGVGANLVNFMTTQLNK 218
Query: 66 DNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALL 125
D ++ + + W+G + ++GA+++D+Y GR+ T I+ GLV L +TT L L
Sbjct: 219 DVVSSITSFNNWSGLATLTPILGAYIADTYTGRFWTITFSLLIYAIGLVLLVLTTTLKSL 398
Query: 126 RPKGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYSK 185
RP C NG C E S+L++ +FY S+Y IA+G+G +PN++TFGADQFD+ +E K
Sbjct: 399 RP-ACENG--ICREASNLQVALFYTSLYTIAVGSGAVKPNMSTFGADQFDDFRHEEKEQK 569
Query: 186 VAFFSYFYLALNLGSLFSNTILGYFEDE-----GLWALGFWASAGSAFLAL 231
V+FF+++ GSL + + Y +++ GL + +W S +L L
Sbjct: 570 VSFFNWWAFNGACGSLMATLFVVYIQEKNGXGFGLQFICYWFSTFLYYLFL 722
>TC18768
Length = 738
Score = 107 bits (268), Expect(2) = 3e-26
Identities = 62/173 (35%), Positives = 100/173 (56%), Gaps = 1/173 (0%)
Frame = +2
Query: 293 LHTHGFKFLDRAAYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSV 352
+HT F+FLD+A R E + + W LC + QVE+VK +L ++PI+ CTI+++
Sbjct: 233 VHTDKFRFLDKACI---RVEEAGSNTKKSSWRLCSVGQVEQVKILLSVIPIFSCTIVFNT 403
Query: 353 VFTQMASLFVEQGAAMKTTIS-SFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKS 411
+ Q+ + V+QG+AM T ++ SF IPPAS+ S + + I + Y P ++
Sbjct: 404 ILAQLQTFSVQQGSAMDTHLTKSFHIPPASLQSIPYILLIIVVPLYDTFFVPFARRITGH 583
Query: 412 SSKGLTELQRMGIGLVIAIIAMVTAGIVECYRLKYAKQGDTSSLSIFWQIPQY 464
S G++ L+R+G GL +A +MV+A ++E R A +LSIFW PQ+
Sbjct: 584 ES-GISPLRRIGFGLFLATFSMVSAALLEKKRRDEA-LNHNKTLSIFWITPQF 736
Score = 28.1 bits (61), Expect(2) = 3e-26
Identities = 16/44 (36%), Positives = 23/44 (51%)
Frame = +1
Query: 218 LGFWASAGSAFLALVLFLVGTPKYRHFKPCGNPLSRFCQVFFAA 261
+GF SA + L+ + GT YR+ P G+ L+ QV AA
Sbjct: 28 VGFGISAAVMAMGLISLISGTFYYRNKPPQGSILTPIAQVLVAA 159
>CB826936
Length = 596
Score = 109 bits (272), Expect = 1e-24
Identities = 59/156 (37%), Positives = 92/156 (58%), Gaps = 6/156 (3%)
Frame = +3
Query: 414 KGLTELQRMGIGLVIAIIAMVTAGIVECYRLKYAK------QGDTSSLSIFWQIPQYALI 467
+G+ LQR+G GLV+ II MVTA + E RL A+ Q D L+IF +PQ+AL
Sbjct: 30 RGIAMLQRLGAGLVMYIITMVTAWLSERKRLSVAREYNLLGQHDKIPLTIFILLPQFALT 209
Query: 468 GASEVFMYVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPG 527
G + F+ + L+FF Q P+G+KS G + TS +LG + SS ++S V ++ + G
Sbjct: 210 GVANNFVDIAMLDFFYDQAPEGMKSLGISYSTTSTALGCFFSSFLLSTVADLTKKHGHKG 389
Query: 528 WIPGNLNRGHLDRFFFLLAVLTSLDLIAYIACAKWF 563
W+ NLN HLD ++ +A+L+ + + ++ AK F
Sbjct: 390 WVLDNLNVSHLDYYYAFMAILSVANFLCFLVAAKLF 497
>AV409553
Length = 433
Score = 103 bits (257), Expect = 8e-23
Identities = 53/136 (38%), Positives = 82/136 (59%)
Frame = +2
Query: 108 IFVTGLVSLSVTTYLALLRPKGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIA 167
++V G++ L+V L L+P C NG C + S+ ++ FY ++Y A+G GG +PNI+
Sbjct: 23 VYVMGMILLTVAVSLKSLKPT-CTNGI--CNKASTSQIVFFYTALYTTAIGAGGTKPNIS 193
Query: 168 TFGADQFDEDHSKESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSA 227
TFGADQFD+ + E +K +FF+++ LG+L + L Y ++ W LG+
Sbjct: 194 TFGADQFDDFNPHEKQTKASFFNWWMFTSFLGALIATLGLVYIQENLGWGLGYGIPTSGL 373
Query: 228 FLALVLFLVGTPKYRH 243
L+LV+F VGTP YRH
Sbjct: 374 LLSLVIFYVGTPMYRH 421
>TC13000 similar to PIR|T47573|T47573 peptide transport-like protein -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(21%)
Length = 446
Score = 102 bits (254), Expect = 2e-22
Identities = 51/114 (44%), Positives = 73/114 (63%)
Frame = +3
Query: 6 ETEELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQ 65
E + T DG+VD+ G P+ + +G W A IL N+ LA++G+ NLVL+ + L Q
Sbjct: 99 EDDNYTKDGTVDYLGNPANKKQTGTWKACPFILGNECCERLAYYGMSTNLVLYFKKQLNQ 278
Query: 66 DNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVT 119
+A A+ NVS W+GT YI L+GAFL+DSY GRY T A F I+V G+ L+++
Sbjct: 279 HSATASKNVSNWSGTCYITPLIGAFLADSYLGRYWTIACFSIIYVIGMTLLTLS 440
>BU494138
Length = 518
Score = 96.7 bits (239), Expect = 9e-21
Identities = 53/155 (34%), Positives = 82/155 (52%), Gaps = 5/155 (3%)
Frame = +2
Query: 328 ITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGAAMKTTI-SSFKIPPASMSSFD 386
I QVEE+KC+ R+ PIW I+ Q + V Q M I S F+IP S+
Sbjct: 23 IQQVEEIKCLARIFPIWAAGILGFTAMAQQGTFTVSQAMKMDRHIGSKFQIPAGSLGVIS 202
Query: 387 ILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMGIGLVIAIIAMVTAGIVECYRLKY 446
+++ +++ FY R P V ++ K G+T LQR+GIG+V ++++M+ AG+VE R
Sbjct: 203 FITIGLWVPFYDRFFVPAVRRITKHEG-GITLLQRIGIGMVFSVLSMIVAGLVEKVRRGV 379
Query: 447 AKQGDT----SSLSIFWQIPQYALIGASEVFMYVG 477
A + +S+ W PQ L+G E F +G
Sbjct: 380 ANSNPNPLGIAPMSVMWLAPQLVLMGLCEAFNAIG 484
>AW720047
Length = 419
Score = 96.7 bits (239), Expect = 9e-21
Identities = 49/134 (36%), Positives = 77/134 (56%)
Frame = +3
Query: 31 WFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDNAAAANNVSKWTGTVYIFSLVGAF 90
W A IL + LA++G+ NLV++LT L + +AA NV+ + GT Y+ L+G+F
Sbjct: 6 WKACPFILGSFFCERLAYYGIATNLVIYLTTKLNEGTVSAAKNVNTFHGTCYLTPLIGSF 185
Query: 91 LSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRPKGCGNGKLECGEHSSLEMGMFYL 150
L+D+YWGRY T +F +++ G+ L ++ + L+P C N S+ F+L
Sbjct: 186 LADAYWGRYWTITVFYAVYLIGMCILILSATIPALKPAECVNSVCTATLAQSV---XFFL 356
Query: 151 SIYLIALGNGGYQP 164
+Y IALG GG +P
Sbjct: 357 GLYSIALGTGGIKP 398
>BI420650
Length = 576
Score = 94.4 bits (233), Expect = 5e-20
Identities = 57/162 (35%), Positives = 84/162 (51%)
Frame = +1
Query: 27 TSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDNAAAANNVSKWTGTVYIFSL 86
T G + A + L + F V+LVL+ V+ D A++AN ++ + G+ ++ SL
Sbjct: 103 TFGGFRASMFVFALSALDNMGFVANMVSLVLYFIMVMHFDLASSANTLTNFMGSTFLLSL 282
Query: 87 VGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRPKGCGNGKLECGEHSSLEMG 146
VG F+SD+Y R TC +F + V LV L+V L L P+ CG G +
Sbjct: 283 VGGFISDTYLNRLTTCLLFGSLEVLALVMLTVQAGLDSLHPEACGKSSCIKGGIAV---- 450
Query: 147 MFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYSKVAF 188
M Y S+ L+ALG GG + + FGADQFDE E+ + F
Sbjct: 451 MLYTSLGLLALGLGGVRGAMVAFGADQFDEKDPVEAKALATF 576
>AV768616
Length = 396
Score = 91.7 bits (226), Expect = 3e-19
Identities = 44/79 (55%), Positives = 60/79 (75%)
Frame = -1
Query: 485 QTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGHLDRFFFL 544
Q PD ++S SAL + ++SLG Y+SSL+V+IV KIST++ GWIP NLN GH+D FF+L
Sbjct: 393 QAPDAMRSLSSALSLLTVSLGQYLSSLLVTIVTKISTKNGGSGWIPDNLNYGHVDYFFWL 214
Query: 545 LAVLTSLDLIAYIACAKWF 563
LAVL+ L+LI Y+ AK +
Sbjct: 213 LAVLSVLNLIVYLPVAKLY 157
>TC14390 weakly similar to UP|Q9FNL8 (Q9FNL8) Peptide transporter, partial
(18%)
Length = 622
Score = 87.0 bits (214), Expect = 7e-18
Identities = 43/103 (41%), Positives = 65/103 (62%)
Frame = +1
Query: 8 EELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDN 67
E+ T DG+VD GRP +R+ +G W A T I+ + +A++G+ NLV+FL L +
Sbjct: 76 EDYTQDGTVDLKGRPVLRSKTGSWKACTFIVGYELFERMAYYGISSNLVVFLGSKLHEGT 255
Query: 68 AAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFV 110
A++NNVS W G V+ LVGA+++D+Y GRY T I I++
Sbjct: 256 VASSNNVSNWAGVVWTTPLVGAYIADAYLGRYWTFIIASCIYL 384
>CB828533
Length = 557
Score = 87.0 bits (214), Expect = 7e-18
Identities = 49/149 (32%), Positives = 81/149 (53%), Gaps = 3/149 (2%)
Frame = +1
Query: 112 GLVSLSVTTYLALLRPKGCGNGKL---ECGEHSSLEMGMFYLSIYLIALGNGGYQPNIAT 168
G+ L+ T + ++ C + + EC E S ++ + +++Y IA+G GG + +++
Sbjct: 103 GMCLLTGATTIPSMKAPPCSSVQRQHHECIEASGKQVALLLVALYTIAVGAGGIKSSVSG 282
Query: 169 FGADQFDEDHSKESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAF 228
FG+DQFD +E V FF+ FY ++ GSLFS +L Y +D GF AG
Sbjct: 283 FGSDQFDTTDPREEKKMVFFFNRFYFFVSTGSLFSVLVLVYVQDNIGRGWGFGIPAGIML 462
Query: 229 LALVLFLVGTPKYRHFKPCGNPLSRFCQV 257
++L L G P YR+ +P G+PL+ +V
Sbjct: 463 VSLAFLLYGKPLYRYKRPQGSPLTLIWKV 549
>TC13937 weakly similar to UP|Q9LVE0 (Q9LVE0) Nitrate transporter
(AT3g21670/MIL23_23), partial (27%)
Length = 558
Score = 84.7 bits (208), Expect = 4e-17
Identities = 50/159 (31%), Positives = 86/159 (53%), Gaps = 3/159 (1%)
Frame = +2
Query: 15 SVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDNAAAANNV 74
+VD+ GRP ++ +G W A +IL + + G+ +N+V +L VL A +A V
Sbjct: 80 AVDYLGRPPDKSKTGGWTAAWLILGVELAQKIMVLGISMNIVTYLVGVLNLPTAYSATIV 259
Query: 75 SKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRPKGCGNGK 134
+ GT+ + SL+G FL+D GRY T AI I G+ L+ T + ++ C + +
Sbjct: 260 TNCLGTMNLVSLLGGFLADVKLGRYLTIAISATISAVGMCLLTGATTIPSMKAPPCSSVQ 439
Query: 135 L---ECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFG 170
EC E S ++ + +++Y IA+G GG + +++ FG
Sbjct: 440 RQHHECIEASGKQVALLLVALYTIAVGAGGIKSSVSGFG 556
>BP071089
Length = 393
Score = 84.0 bits (206), Expect = 6e-17
Identities = 39/77 (50%), Positives = 56/77 (72%)
Frame = -3
Query: 485 QTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGHLDRFFFL 544
+ PD ++S SAL +T+ +LGNYVS+L+V IV K++T GWIP N+NRGHLD F++L
Sbjct: 385 EAPDAMRSLCSALSLTTNALGNYVSTLLVIIVTKVTTSSGSLGWIPDNMNRGHLDYFYWL 206
Query: 545 LAVLTSLDLIAYIACAK 561
L VL+ L+ + Y+ AK
Sbjct: 205 LTVLSLLNFLVYLWIAK 155
>TC18700 similar to UP|Q9LYR6 (Q9LYR6) Peptide transporter-like protein,
partial (18%)
Length = 615
Score = 82.4 bits (202), Expect = 2e-16
Identities = 44/134 (32%), Positives = 74/134 (54%), Gaps = 4/134 (2%)
Frame = +1
Query: 454 SLSIFWQIPQYALIGASEVFMYVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIV 513
+LS +W + QY LIG +EVF VG LEF + PD +KS GSA + LG + +++I
Sbjct: 43 NLSAYWLLIQYCLIGVAEVFCIVGLLEFLYEEAPDAMKSIGSAYAALAGGLGCFAATIIN 222
Query: 514 SIVMKISTEDHMPGWIPGNLNRGHLDRFFFLLAVLTSLDLIAYIACAKWF----QNIQMA 569
SI+ ++ ++ W+ N+N G D F+++L L+ ++ +I A + Q +
Sbjct: 223 SIIKSLTGKEGKESWLAQNINTGRFDYFYWILTALSLVNFCIFIYSAHRYKYRTQQVYEM 402
Query: 570 CKYDNNDEPSSCKV 583
K+D + SS +V
Sbjct: 403 EKHDVANNVSSTRV 444
>TC8505 similar to UP|Q8LG02 (Q8LG02) Nitrate transporter, partial (23%)
Length = 1495
Score = 82.4 bits (202), Expect = 2e-16
Identities = 46/106 (43%), Positives = 65/106 (60%)
Frame = +3
Query: 455 LSIFWQIPQYALIGASEVFMYVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVS 514
LS FW + F YVGQLEFF + P+ +KS + L + ++S+G +VSSL+VS
Sbjct: 216 LSFFWW-------EQGKAFAYVGQLEFFIREAPERMKSMSTGLFLAALSMGYFVSSLLVS 374
Query: 515 IVMKISTEDHMPGWIPGNLNRGHLDRFFFLLAVLTSLDLIAYIACA 560
IV K+S + W+ NLN+G LD F++LLAVL L+ I +I A
Sbjct: 375 IVDKLSKKK----WLKSNLNKGRLDYFYWLLAVLGVLNFILFIVLA 500
Score = 43.5 bits (101), Expect = 9e-05
Identities = 22/49 (44%), Positives = 33/49 (66%)
Frame = +2
Query: 423 GIGLVIAIIAMVTAGIVECYRLKYAKQGDTSSLSIFWQIPQYALIGASE 471
GIGLV + +AM+ + IVE R + A + T+ +S FW +PQ+ L+GA E
Sbjct: 101 GIGLVFSFVAMMVSAIVEKERRENAVKKHTN-ISAFWLVPQFFLVGAGE 244
>AV768501
Length = 442
Score = 82.0 bits (201), Expect = 2e-16
Identities = 36/88 (40%), Positives = 63/88 (70%)
Frame = -1
Query: 476 VGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNR 535
+GQL+FF + P G+K+ L ++++SLG + SS++V++V K++ GW+ NLN+
Sbjct: 442 IGQLDFFLRECPKGMKTMSPGLFLSTLSLGFFFSSMLVTLVHKMTGPRQ--GWLADNLNQ 269
Query: 536 GHLDRFFFLLAVLTSLDLIAYIACAKWF 563
G LD F++LLAVL+ ++L+ ++ CAKW+
Sbjct: 268 GKLDDFYWLLAVLSGVNLVVFLFCAKWY 185
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.139 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,664,368
Number of Sequences: 28460
Number of extensions: 153845
Number of successful extensions: 1046
Number of sequences better than 10.0: 74
Number of HSP's better than 10.0 without gapping: 1012
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1017
length of query: 583
length of database: 4,897,600
effective HSP length: 95
effective length of query: 488
effective length of database: 2,193,900
effective search space: 1070623200
effective search space used: 1070623200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 58 (26.9 bits)
Medicago: description of AC147404.4


