

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
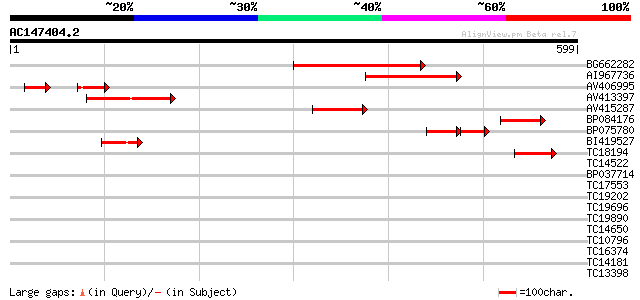
Query= AC147404.2 + phase: 0
(599 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG662282 205 2e-53
AI967736 119 1e-27
AV406995 59 2e-17
AV413397 80 1e-15
AV415287 77 8e-15
BP084176 76 1e-14
BP075780 49 5e-12
BI419527 52 2e-07
TC18194 weakly similar to UP|Q9FF76 (Q9FF76) Similarity to inosi... 40 8e-04
TC14522 similar to UP|Q9FJ79 (Q9FJ79) DNA topoisomerase I, parti... 32 0.29
BP037714 30 0.86
TC17553 weakly similar to UP|AAP20176 (AAP20176) Kinase substrat... 30 1.1
TC19202 similar to UP|Q9LFZ9 (Q9LFZ9) F20N2.11, partial (5%) 28 4.3
TC19696 similar to UP|Q9ZQT8 (Q9ZQT8) WREBP-2 protein, partial (... 28 5.6
TC19890 27 7.3
TC14650 similar to UP|IF3A_TOBAC (Q40554) Eukaryotic translation... 27 7.3
TC10796 similar to GB|AAO64916.1|29029074|BT005981 At5g53430 {Ar... 27 7.3
TC16374 similar to UP|Q7LZA6 (Q7LZA6) Protamine II (Fragment), p... 27 7.3
TC14181 similar to UP|Q9FE06 (Q9FE06) Phi-1-like protein (AT5g64... 27 9.5
TC13398 weakly similar to UP|Q8AWB2 (Q8AWB2) Keratin gamma3, par... 27 9.5
>BG662282
Length = 440
Score = 205 bits (521), Expect = 2e-53
Identities = 102/141 (72%), Positives = 123/141 (86%), Gaps = 1/141 (0%)
Frame = +1
Query: 300 DVSDMLSDEDNDTFSELANNEDANGII-SVKSHPKYVRIVSKQMVGIYVSVWVQRKLRRH 358
DV + ED+DTF +L +N+D + ++ + P+YVRIVSKQMVGIYVSVWVQR+LRRH
Sbjct: 16 DVCGVSPPEDDDTFLDLPDNQDDIELGGAMDTRPRYVRIVSKQMVGIYVSVWVQRRLRRH 195
Query: 359 VHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSRMCFVCSHLASGQKDGAEQRRNSDVHEI 418
+++LKVS VGVG+MGY+GNKGSVSVSMS+FQSRMCFVCSHL+SGQKDGAEQRRNS+VHEI
Sbjct: 196 INNLKVSPVGVGIMGYIGNKGSVSVSMSLFQSRMCFVCSHLSSGQKDGAEQRRNSNVHEI 375
Query: 419 LQRTRFSSVFDTDQPQKIPSH 439
L+RT FSSVFD+DQP IPSH
Sbjct: 376 LRRTCFSSVFDSDQPLTIPSH 438
>AI967736
Length = 334
Score = 119 bits (299), Expect = 1e-27
Identities = 55/101 (54%), Positives = 77/101 (75%)
Frame = +2
Query: 377 NKGSVSVSMSVFQSRMCFVCSHLASGQKDGAEQRRNSDVHEILQRTRFSSVFDTDQPQKI 436
++GSVSVSMS++Q+ CFVCSHL SG+K+G + +RN+DVHEI +RT F S+ P+ I
Sbjct: 8 HEGSVSVSMSIYQTLFCFVCSHLTSGEKEGDDLKRNADVHEIHRRTHFQSLSYIGLPKTI 187
Query: 437 PSHDKIFWFGDLNYRINMSDGEIRKLVDLKKWNELMKFDQL 477
H++I W GDLNYRIN+S+ E R LV K+W++L++ DQL
Sbjct: 188 LDHERIIWLGDLNYRINLSNVETRALVSKKQWSKLIEKDQL 310
>AV406995
Length = 281
Score = 58.5 bits (140), Expect(2) = 2e-17
Identities = 28/34 (82%), Positives = 30/34 (87%)
Frame = +2
Query: 72 KDYEMKRHRRRKSETLRAQYINTKEVRVTIGTWN 105
K Y+M RHRR SETLRAQYINTKEVRVT+GTWN
Sbjct: 182 KSYKM-RHRRGNSETLRAQYINTKEVRVTVGTWN 280
Score = 47.8 bits (112), Expect(2) = 2e-17
Identities = 22/28 (78%), Positives = 25/28 (88%)
Frame = +1
Query: 16 KKWLNVKQKVYDFSEDEANTETDESEDD 43
KKWLN+K KVYDFSEDE +TET ESED+
Sbjct: 1 KKWLNIKPKVYDFSEDEVDTET-ESEDE 81
>AV413397
Length = 382
Score = 79.7 bits (195), Expect = 1e-15
Identities = 41/94 (43%), Positives = 57/94 (60%)
Frame = +1
Query: 82 RKSETLRAQYINTKEVRVTIGTWNVAGKHPCNDLEIEGWLCTEEEPSDIYIIGFQEVVPL 141
R SE + ++ RV TWNV GK P +L++ +L EP D+Y++GFQE+VPL
Sbjct: 10 RGSEPSMSSSEALQDFRVFAATWNVGGKCPSENLDLSDFLQVHNEP-DMYVLGFQEIVPL 186
Query: 142 NAGNVFGAEDSKPIPKWDALIRRTLNKSSEPGTK 175
NAGNV ED++P KW LI +LN S+ +K
Sbjct: 187 NAGNVLVLEDNEPAAKWLTLINHSLNGPSDLASK 288
>AV415287
Length = 296
Score = 77.0 bits (188), Expect = 8e-15
Identities = 33/58 (56%), Positives = 46/58 (78%)
Frame = +2
Query: 321 DANGIISVKSHPKYVRIVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNK 378
D ++ K YV++VSKQMVGI+++VWV+R LR+H+ +LKVS VGVG+MGY+GNK
Sbjct: 122 DLGALMKRKRRSSYVKLVSKQMVGIFITVWVRRSLRKHIQNLKVSTVGVGVMGYIGNK 295
>BP084176
Length = 353
Score = 76.3 bits (186), Expect = 1e-14
Identities = 31/48 (64%), Positives = 41/48 (84%)
Frame = -3
Query: 519 EKRRAPAWCDRILWLGKGIKQLKYQSAENQLSDHRPVSSIFLVDVEVI 566
EKRRAPAWCDRI+W GKG+KQL+Y +E++LSDHRPV ++F +V V+
Sbjct: 312 EKRRAPAWCDRIVWYGKGLKQLQYARSESKLSDHRPVKAVFSAEVRVL 169
>BP075780
Length = 400
Score = 48.9 bits (115), Expect(2) = 5e-12
Identities = 21/31 (67%), Positives = 22/31 (70%)
Frame = +2
Query: 477 LSNELCKGHVFEGWKEGLINFPPTYKYEFNS 507
L EL +GHVF GW EG I FPPTYKY NS
Sbjct: 305 LKKELREGHVFRGWHEGAIEFPPTYKYHPNS 397
Score = 38.9 bits (89), Expect(2) = 5e-12
Identities = 15/37 (40%), Positives = 23/37 (61%)
Frame = +3
Query: 441 KIFWFGDLNYRINMSDGEIRKLVDLKKWNELMKFDQL 477
++ W GDLNYRI M D E + + ++W +K DQ+
Sbjct: 111 RVVWLGDLNYRIYMPDSETQSSIKRREWET*LKQDQV 221
>BI419527
Length = 393
Score = 52.4 bits (124), Expect = 2e-07
Identities = 21/43 (48%), Positives = 31/43 (71%)
Frame = +3
Query: 98 RVTIGTWNVAGKHPCNDLEIEGWLCTEEEPSDIYIIGFQEVVP 140
R+ +GTWNV GK P +L ++ WL P++IY+IGFQE++P
Sbjct: 267 RMFVGTWNVGGKSPNEELNLKDWLML-PSPAEIYVIGFQEIIP 392
>TC18194 weakly similar to UP|Q9FF76 (Q9FF76) Similarity to inositol
polyphosphate 5'-phosphatase, partial (9%)
Length = 676
Score = 40.4 bits (93), Expect = 8e-04
Identities = 19/44 (43%), Positives = 28/44 (63%)
Frame = +1
Query: 534 GKGIKQLKYQSAENQLSDHRPVSSIFLVDVEVIDHRKLERAIYF 577
G G++QL Y E + SDHRPV + FLV+VEV+ + ++ F
Sbjct: 1 GTGLQQLSYVRKEFKFSDHRPVCATFLVEVEVMFRGQKKKVFTF 132
>TC14522 similar to UP|Q9FJ79 (Q9FJ79) DNA topoisomerase I, partial (12%)
Length = 635
Score = 32.0 bits (71), Expect = 0.29
Identities = 21/78 (26%), Positives = 37/78 (46%)
Frame = +2
Query: 169 SSEPGTKKKSNSAPPSPIRRISSFNTQINPLDSALDKKEEIKTIISIEKNLQLSKIYDID 228
S P KKK N + + ++IS + +I + + KE++KT+ L SKI +D
Sbjct: 86 SKSPDGKKKKNLSSEALEKKISQTHAKIEKMQRDMKTKEDLKTVA-----LGTSKINYLD 250
Query: 229 LQTILDWPELRLDPIHHV 246
+ + W + PI +
Sbjct: 251 PRITVAWCKRHEVPIEKI 304
>BP037714
Length = 463
Score = 30.4 bits (67), Expect = 0.86
Identities = 21/72 (29%), Positives = 36/72 (49%), Gaps = 2/72 (2%)
Frame = +1
Query: 27 DFSEDEANTETDESEDDVKLIEIETNHTSKPQANTTKPTQLNTSCKDYEMKRHRRRKSET 86
D SE+E+ ET +++ +IEIE + KP+ + + K E+ R R + E
Sbjct: 253 DESEEESEEETKKNKGTQGIIEIENPNLVKPKNVKARDVDIE---KTTELSRREREEIEK 423
Query: 87 LRA--QYINTKE 96
RA +Y+ +E
Sbjct: 424 QRAHERYMRLQE 459
>TC17553 weakly similar to UP|AAP20176 (AAP20176) Kinase substrate HASPP28
(Fragment), partial (61%)
Length = 754
Score = 30.0 bits (66), Expect = 1.1
Identities = 25/91 (27%), Positives = 45/91 (48%), Gaps = 5/91 (5%)
Frame = +3
Query: 11 PSTVMKKWLNVKQKVYD---FSEDEANTETDESEDDVKLIEIETNHTSKPQANTTKPTQL 67
P T +K + +++V + S DE+ ET +++ +IEIE + KP++ + L
Sbjct: 171 PRTFKQKEVEDEKEVSEEEATSGDESEEETTKTKGTQGVIEIENPNLVKPKSLKARDVDL 350
Query: 68 NTSCKDYEMKRHRRRKSETLRA--QYINTKE 96
K E+ R R + E RA +Y+ +E
Sbjct: 351 E---KTTELSRREREEIEKQRAHERYMRLQE 434
>TC19202 similar to UP|Q9LFZ9 (Q9LFZ9) F20N2.11, partial (5%)
Length = 526
Score = 28.1 bits (61), Expect = 4.3
Identities = 20/62 (32%), Positives = 28/62 (44%), Gaps = 4/62 (6%)
Frame = -2
Query: 96 EVRVTIGTWNVAGKHPCNDLE----IEGWLCTEEEPSDIYIIGFQEVVPLNAGNVFGAED 151
+V VT WN+ GKH +L I+ LCT P ++G V+P + V
Sbjct: 411 KVLVTYLLWNMRGKHLT*NLNYIFIIQSELCTVYAPQVSVLLGLDLVLPASGFLVSNLRV 232
Query: 152 SK 153
SK
Sbjct: 231 SK 226
>TC19696 similar to UP|Q9ZQT8 (Q9ZQT8) WREBP-2 protein, partial (12%)
Length = 496
Score = 27.7 bits (60), Expect = 5.6
Identities = 15/41 (36%), Positives = 19/41 (45%)
Frame = +3
Query: 2 KGKRSEVFWPSTVMKKWLNVKQKVYDFSEDEANTETDESED 42
KG P V+KK VKQ+ YD +E + D ED
Sbjct: 78 KGSAPTPSAPLNVIKKPWPVKQETYDEDSEETEEDRDNVED 200
>TC19890
Length = 477
Score = 27.3 bits (59), Expect = 7.3
Identities = 21/90 (23%), Positives = 42/90 (46%), Gaps = 12/90 (13%)
Frame = -2
Query: 249 SPKMRRVQSTSDSASLYGFEMKSLHQSSGNFSLLWSE---KQQEI---------VPQVFD 296
SPKM S + + SL +++G FSL W++ K++ + +P +
Sbjct: 317 SPKM*ATPKQPTSLMIKSIKFNSLSKANGRFSLQWTQPKAKEEALKIRTWWRGELPAEKE 138
Query: 297 SHLDVSDMLSDEDNDTFSELANNEDANGII 326
S + +++ DE +T + N +D+ +I
Sbjct: 137 SLIK*PELVLDEHINTRLAMKNAQDSPVLI 48
>TC14650 similar to UP|IF3A_TOBAC (Q40554) Eukaryotic translation initiation
factor 3 subunit 10 (eIF-3 theta) (Eukaryotic
translation initiation factor 3 large subunit) (eIF3a)
(PNLA-35), partial (11%)
Length = 978
Score = 27.3 bits (59), Expect = 7.3
Identities = 8/29 (27%), Positives = 18/29 (61%)
Frame = +3
Query: 425 SSVFDTDQPQKIPSHDKIFWFGDLNYRIN 453
+SV + P + HD+++W G+ N+ ++
Sbjct: 822 TSVLLRNMPTRFSVHDRVYWVGNRNFNLD 908
>TC10796 similar to GB|AAO64916.1|29029074|BT005981 At5g53430 {Arabidopsis
thaliana;}, partial (5%)
Length = 519
Score = 27.3 bits (59), Expect = 7.3
Identities = 11/37 (29%), Positives = 20/37 (53%)
Frame = +1
Query: 54 TSKPQANTTKPTQLNTSCKDYEMKRHRRRKSETLRAQ 90
++ P +QLN SC Y+ K+ R++K + + Q
Sbjct: 229 STNPSPPLQGKSQLNQSCNFYQTKKERKKKQKKIGEQ 339
>TC16374 similar to UP|Q7LZA6 (Q7LZA6) Protamine II (Fragment), partial
(63%)
Length = 1130
Score = 27.3 bits (59), Expect = 7.3
Identities = 21/96 (21%), Positives = 43/96 (43%)
Frame = +3
Query: 5 RSEVFWPSTVMKKWLNVKQKVYDFSEDEANTETDESEDDVKLIEIETNHTSKPQANTTKP 64
RSE + +K LN+ ++ + + E +T+++E++V+ K
Sbjct: 543 RSEKLYKQGRLKAMLNINGEIENDEDSETEIQTEDTEENVQ-----------------KK 671
Query: 65 TQLNTSCKDYEMKRHRRRKSETLRAQYINTKEVRVT 100
+ NT + KR ++KS+ NT+++R T
Sbjct: 672 KKKNTKGEKQNKKRLGKKKSDK-----SNTRKMRST 764
>TC14181 similar to UP|Q9FE06 (Q9FE06) Phi-1-like protein
(AT5g64260/MSJ1_10), partial (84%)
Length = 1244
Score = 26.9 bits (58), Expect = 9.5
Identities = 15/51 (29%), Positives = 27/51 (52%), Gaps = 2/51 (3%)
Frame = -2
Query: 44 VKLIEIETNHTSKPQANTTKPTQLNTSCKDYEMKRHR--RRKSETLRAQYI 92
V+ + IET + + T+ T++ + K+ +RHR RR+ TL+ I
Sbjct: 892 VRAVRIETRPRNPIHQHLTRITRIRPAPKNPSARRHRLQRRRCRTLKVSII 740
>TC13398 weakly similar to UP|Q8AWB2 (Q8AWB2) Keratin gamma3, partial (4%)
Length = 526
Score = 26.9 bits (58), Expect = 9.5
Identities = 20/76 (26%), Positives = 33/76 (43%)
Frame = +3
Query: 21 VKQKVYDFSEDEANTETDESEDDVKLIEIETNHTSKPQANTTKPTQLNTSCKDYEMKRHR 80
++Q F E EA SED ++ +EI+ + A T K +L ++YE K+
Sbjct: 153 IEQLAQAFLELEARKGA--SEDKIQWVEIKQHFLDLETALTKKHEELEAREREYEQKQSE 326
Query: 81 RRKSETLRAQYINTKE 96
R + + KE
Sbjct: 327 MNTLLAERKEEVARKE 374
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.132 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,392,093
Number of Sequences: 28460
Number of extensions: 135014
Number of successful extensions: 814
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 804
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 812
length of query: 599
length of database: 4,897,600
effective HSP length: 95
effective length of query: 504
effective length of database: 2,193,900
effective search space: 1105725600
effective search space used: 1105725600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC147404.2


