

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147364.8 + phase: 0
(954 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
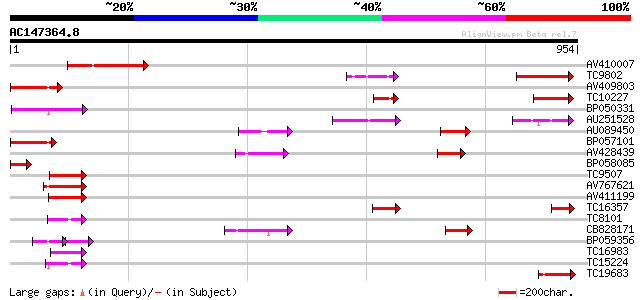
Score E
Sequences producing significant alignments: (bits) Value
AV410007 180 1e-45
TC9802 weakly similar to UP|AAS49059 (AAS49059) At5g53150, parti... 119 2e-27
AV409803 99 4e-21
TC10227 UP|Q7Q2V3 (Q7Q2V3) EbiP1409 (Fragment), partial (10%) 95 5e-20
BP050331 81 7e-16
AU251528 76 2e-14
AU089450 72 5e-13
BP057101 69 5e-12
AV428439 66 3e-11
BP058085 62 6e-10
TC9507 similar to UP|Q8VZB6 (Q8VZB6) DnaJ-like protein, partial ... 61 7e-10
AV767621 61 7e-10
AV411199 57 1e-08
TC16357 weakly similar to UP|Q8DZP0 (Q8DZP0) Bacterial luciferas... 57 2e-08
TC8101 similar to UP|O24074 (O24074) DNAJ-like protein, partial ... 51 8e-07
CB828171 50 2e-06
BP059356 36 4e-06
TC16983 similar to UP|Q9ZSY2 (Q9ZSY2) ALTERED response to gravit... 49 5e-06
TC15224 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein ... 47 1e-05
TC19683 similar to UP|Q9S7L6 (Q9S7L6) AT2G05250 protein (AT2G052... 47 2e-05
>AV410007
Length = 425
Score = 180 bits (456), Expect = 1e-45
Identities = 95/141 (67%), Positives = 108/141 (76%), Gaps = 5/141 (3%)
Frame = +3
Query: 98 KNKFAGAEAAFKLIGEAQRVLSDREKRTRYDMKLNV--NKTAMPPRSNQPKVPTNFNSAT 155
KNKFAGAE+AFKLIGEAQRVL D+EKRT+ DMK V +KTAMP R N K PTNFNSAT
Sbjct: 3 KNKFAGAESAFKLIGEAQRVLLDKEKRTQLDMKRRVPTSKTAMP-RHNPQKAPTNFNSAT 179
Query: 156 K-NNVRTNFTNSNTQQP--PQQQNKQPPQQQNGVRRTFWTACPFCSVKYEYYREILNKSL 212
K NNV+ FTN+ QQ QQQ +Q QQQNG R TFWT CPFCSVKY+YYRE+LNK L
Sbjct: 180 KTNNVKPKFTNTKAQQQRQQQQQRQQAKQQQNGSRPTFWTHCPFCSVKYQYYREVLNKPL 359
Query: 213 RCQQCHRLFVAYILDMQGTSP 233
RCQ C + F+A ++QGTSP
Sbjct: 360 RCQHCQKPFIACDANVQGTSP 422
>TC9802 weakly similar to UP|AAS49059 (AAS49059) At5g53150, partial (3%)
Length = 593
Score = 119 bits (299), Expect = 2e-27
Identities = 59/96 (61%), Positives = 73/96 (75%)
Frame = +3
Query: 853 YRKWSNKIKCSDLKNWDYDIVEVLEVADLFIETSILEHVTGFSSVFRGKSIEGSSGNLRI 912
YRKW+ KIK SDL+ W+YDIV++L+V DL I LE V GF SVF+ + GS+ ++I
Sbjct: 3 YRKWTAKIKPSDLQTWEYDIVQILDVNDLRIAVVHLELVKGFPSVFKVQKNGGSAVTMKI 182
Query: 913 PKKELLRFSHQIPAFKLTEEHGDLRGFWELDPGALP 948
PK ELLRFSHQIPA KLTE+HG ++G ELDPGALP
Sbjct: 183 PKAELLRFSHQIPAIKLTEKHGSVKGCCELDPGALP 290
Score = 54.7 bits (130), Expect = 7e-08
Identities = 35/88 (39%), Positives = 51/88 (57%), Gaps = 2/88 (2%)
Frame = +3
Query: 568 YEYEFVEILSDYVEGEGVHVAYLGKLKGFVSIFIQIMKEDNQP--FQIPSAELFRFSHRI 625
+EY+ V+IL V + V +L +KGF S+F ++ K +IP AEL RFSH+I
Sbjct: 48 WEYDIVQILD--VNDLRIAVVHLELVKGFPSVF-KVQKNGGSAVTMKIPKAELLRFSHQI 218
Query: 626 PSFKMTGQEGVDVHLGYLEFDPASLPMN 653
P+ K+T + G G E DP +LPM+
Sbjct: 219 PAIKLTEKHG--SVKGCCELDPGALPMH 296
>AV409803
Length = 413
Score = 98.6 bits (244), Expect = 4e-21
Identities = 48/88 (54%), Positives = 66/88 (74%)
Frame = +1
Query: 1 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQK 60
MDCNK+EA RAKEIAEKK R++AGA+KFALKA LYP LE + Q L + +V+ SAE K
Sbjct: 148 MDCNKDEAARAKEIAEKKFSLREYAGAKKFALKALNLYPALEGVPQFLSILNVYISAENK 327
Query: 61 VFGDEINWYGILQLERTAGDAMIKKQFR 88
+ G +++WYGIL ++ A + I+K++R
Sbjct: 328 ING-QMDWYGILGVDPFADEETIRKKYR 408
>TC10227 UP|Q7Q2V3 (Q7Q2V3) EbiP1409 (Fragment), partial (10%)
Length = 624
Score = 95.1 bits (235), Expect = 5e-20
Identities = 51/70 (72%), Positives = 56/70 (79%), Gaps = 2/70 (2%)
Frame = +2
Query: 881 LFIETSILEHVTGFSSVFRGKSIEGSSGNLR-IPKKELLRFSHQIPAFKLTEE-HGDLRG 938
+FI+ ILEHV FSSVFR KS G S NLR I KELLRFSH+IP F+LTEE HG+LRG
Sbjct: 2 VFIDVLILEHVPDFSSVFRAKSNLGPSVNLRKISTKELLRFSHRIPGFRLTEEEHGNLRG 181
Query: 939 FWELDPGALP 948
FWELDPGALP
Sbjct: 182 FWELDPGALP 211
Score = 41.6 bits (96), Expect = 6e-04
Identities = 19/42 (45%), Positives = 30/42 (71%)
Frame = +2
Query: 612 QIPSAELFRFSHRIPSFKMTGQEGVDVHLGYLEFDPASLPMN 653
+I + EL RFSHRIP F++T +E ++ G+ E DP +LP++
Sbjct: 95 KISTKELLRFSHRIPGFRLTEEEHGNLR-GFWELDPGALPVH 217
>BP050331
Length = 519
Score = 81.3 bits (199), Expect = 7e-16
Identities = 52/139 (37%), Positives = 81/139 (57%), Gaps = 11/139 (7%)
Frame = +3
Query: 4 NKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQKVFG 63
+K E R EIAE++ + RD G+R+ AL AQ P+LE Q+L + DV +AE+ +
Sbjct: 51 SKSEGERLLEIAEQQXQKRDLKGSRQLALIAQENEPLLEGSDQILAIIDVLEAAEKPLTT 230
Query: 64 ---------DEINWYGILQLE-RTAGDA-MIKKQFRKFALQLHPDKNKFAGAEAAFKLIG 112
+ +++Y ILQ++ R + D +IK+Q+R+ AL LHPDKN F+ + F L+
Sbjct: 231 TASTTTNNRNNLDFYAILQVDHRDSQDLNLIKRQYRRLALLLHPDKNLFSFSHHTFDLVS 410
Query: 113 EAQRVLSDREKRTRYDMKL 131
A +LSD ++ YD L
Sbjct: 411 HAWALLSDPAQKEIYDAGL 467
>AU251528
Length = 388
Score = 76.3 bits (186), Expect = 2e-14
Identities = 37/114 (32%), Positives = 63/114 (54%)
Frame = +3
Query: 544 KGETWALFKNWDITWYKDEESHRQYEYEFVEILSDYVEGEGVHVAYLGKLKGFVSIFIQI 603
KG+ WA+++NW W + ++++ VE+L D+ E GV + L K+ GF ++F
Sbjct: 3 KGDIWAIYRNWSPDWNELTADEVMHKFDVVEVLEDFSEEHGVTIIPLVKVAGFKTVFHHH 182
Query: 604 MKEDNQPFQIPSAELFRFSHRIPSFKMTGQEGVDVHLGYLEFDPASLPMNLEEI 657
+ + + IP E+ RFSH++PS +TG E + G DPA+ P L ++
Sbjct: 183 I-DPREIRVIPREEMLRFSHQVPSCLLTGLEAQNAPKGCRVLDPAATPFELHQV 341
Score = 59.3 bits (142), Expect = 3e-09
Identities = 43/112 (38%), Positives = 63/112 (55%), Gaps = 9/112 (8%)
Frame = +3
Query: 846 KGEVWALYRKWS---NKIKCSDLKNWDYDIVEVLEVADLFIETSI----LEHVTGFSSVF 898
KG++WA+YR WS N++ ++ + +D+VEVLE D E + L V GF +VF
Sbjct: 3 KGDIWAIYRNWSPDWNELTADEVMH-KFDVVEVLE--DFSEEHGVTIIPLVKVAGFKTVF 173
Query: 899 RGKSIEGSSGNLRIPKKELLRFSHQIPAFKLT--EEHGDLRGFWELDPGALP 948
I+ + IP++E+LRFSHQ+P+ LT E +G LDP A P
Sbjct: 174 H-HHIDPREIRV-IPREEMLRFSHQVPSCLLTGLEAQNAPKGCRVLDPAATP 323
>AU089450
Length = 383
Score = 71.6 bits (174), Expect = 5e-13
Identities = 40/95 (42%), Positives = 54/95 (56%), Gaps = 4/95 (4%)
Frame = +3
Query: 386 VKQKQEEKIRAGG-KEAAEGSKQMDKT---FEHSSPGSTSKTSNCPNAYVYPDAEFSDFD 441
V ++++ + AG K + + +D + FEH G S T PD +F DFD
Sbjct: 108 VNEREKSQAEAGQVKSGSHTNTVLDVSGNHFEHGKIGPVSMT--------VPDPDFHDFD 263
Query: 442 KDRKKECFAPGQIWAIYDSIDGMPRFYALIRKVLS 476
KDR +ECF P QIWA+ D DGMPR Y LIR+V+S
Sbjct: 264 KDRSEECFRPKQIWAL*DEEDGMPRLYCLIREVIS 368
Score = 46.6 bits (109), Expect = 2e-05
Identities = 20/50 (40%), Positives = 32/50 (64%)
Frame = +3
Query: 725 PEAFEIPDAQFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYYGQIKKVVT 774
P + +PD F +F+ RS + F+ QIWA +EDGMP+ Y I++V++
Sbjct: 219 PVSMTVPDPDFHDFDKDRSEECFRPKQIWAL*DEEDGMPRLYCLIREVIS 368
>BP057101
Length = 546
Score = 68.6 bits (166), Expect = 5e-12
Identities = 36/78 (46%), Positives = 56/78 (71%)
Frame = +3
Query: 1 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQK 60
+ NKEEAL+A EIAE++ RDFAGA+ +A+KA+ L P +E I+QM+ +V+ ++E K
Sbjct: 300 VQANKEEALQALEIAERRFAQRDFAGAKSYAVKAKTLCPEVEGISQMVATFEVYIASEIK 479
Query: 61 VFGDEINWYGILQLERTA 78
G E+++Y IL L+ +A
Sbjct: 480 HNG-ELDYYAILGLKPSA 530
>AV428439
Length = 292
Score = 65.9 bits (159), Expect = 3e-11
Identities = 36/90 (40%), Positives = 49/90 (54%), Gaps = 1/90 (1%)
Frame = +3
Query: 380 KNEHEKVKQKQEEKIRAGGKEAAEGSKQMDKTFEHSSPGSTSKTSNC-PNAYVYPDAEFS 438
K E K+ EK +A + + ++ + S PG + S P + PD++F
Sbjct: 30 KLSSEAATLKEREKAQA---DVGQVKRETCRRAVLSVPGLELEHSKAGPISITVPDSDFH 200
Query: 439 DFDKDRKKECFAPGQIWAIYDSIDGMPRFY 468
DFDKDR +ECF P QIWA+YD DGMPR Y
Sbjct: 201 DFDKDRAEECFRPKQIWALYDEEDGMPRLY 290
Score = 47.0 bits (110), Expect = 1e-05
Identities = 19/46 (41%), Positives = 30/46 (64%)
Frame = +3
Query: 721 SASTPEAFEIPDAQFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYY 766
S + P + +PD+ F +F+ R+ + F+ QIWA Y +EDGMP+ Y
Sbjct: 153 SKAGPISITVPDSDFHDFDKDRAEECFRPKQIWALYDEEDGMPRLY 290
>BP058085
Length = 288
Score = 61.6 bits (148), Expect = 6e-10
Identities = 29/36 (80%), Positives = 33/36 (91%)
Frame = +3
Query: 1 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQR 36
M+CNKEEALRAK IAEKKME++DF GARK ALKAQ+
Sbjct: 180 MECNKEEALRAKVIAEKKMESKDFTGARKIALKAQQ 287
>TC9507 similar to UP|Q8VZB6 (Q8VZB6) DnaJ-like protein, partial (40%)
Length = 905
Score = 61.2 bits (147), Expect = 7e-10
Identities = 27/63 (42%), Positives = 43/63 (67%)
Frame = +1
Query: 67 NWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSDREKRTR 126
N+Y IL LE++ ++K +RK +L++HPDKNK GAE AFK + +A + L + E + +
Sbjct: 517 NYYEILGLEKSCTVEDVRKSYRKLSLKVHPDKNKAPGAEEAFKAVSKAFQCLGNEESKRK 696
Query: 127 YDM 129
YD+
Sbjct: 697 YDL 705
>AV767621
Length = 560
Score = 61.2 bits (147), Expect = 7e-10
Identities = 35/73 (47%), Positives = 47/73 (63%), Gaps = 2/73 (2%)
Frame = +1
Query: 58 EQKVFGDEINWYGILQLERTAGDAMIKKQFRKFALQLHPDK--NKFAGAEAAFKLIGEAQ 115
E++ G +++Y ILQ++R A D +KK +RK A++ HPDK N AEA FK I EA
Sbjct: 40 EEEAMG--VDYYKILQVDRGASDDDLKKAYRKLAMKWHPDKNPNNKKEAEAKFKQISEAY 213
Query: 116 RVLSDREKRTRYD 128
VLSD +KR YD
Sbjct: 214 DVLSDPQKRAVYD 252
>AV411199
Length = 423
Score = 57.0 bits (136), Expect = 1e-08
Identities = 29/65 (44%), Positives = 42/65 (64%), Gaps = 2/65 (3%)
Frame = +2
Query: 66 INWYGILQLERTAGDAMIKKQFRKFALQLHPDKN--KFAGAEAAFKLIGEAQRVLSDREK 123
+++Y IL+++ A D +KK +R+ A++ HPDKN AE FKLI E+ VLSD +K
Sbjct: 65 VDYYEILEVDHHATDEELKKAYRRLAMKWHPDKNPDNKNDAETKFKLISESYEVLSDPQK 244
Query: 124 RTRYD 128
R YD
Sbjct: 245 RAIYD 259
>TC16357 weakly similar to UP|Q8DZP0 (Q8DZP0) Bacterial luciferase family
protein, partial (6%)
Length = 572
Score = 56.6 bits (135), Expect = 2e-08
Identities = 27/47 (57%), Positives = 33/47 (69%)
Frame = +1
Query: 611 FQIPSAELFRFSHRIPSFKMTGQEGVDVHLGYLEFDPASLPMNLEEI 657
F IP EL+RFSH+IPS+KMTG E V G E DPA LP++L E+
Sbjct: 16 FCIPPNELYRFSHQIPSYKMTGDEREGVPRGSFELDPAGLPLSLFEV 156
Score = 46.6 bits (109), Expect = 2e-05
Identities = 24/41 (58%), Positives = 28/41 (67%), Gaps = 2/41 (4%)
Frame = +1
Query: 912 IPKKELLRFSHQIPAFKLT--EEHGDLRGFWELDPGALPPS 950
IP EL RFSHQIP++K+T E G RG +ELDP LP S
Sbjct: 22 IPPNELYRFSHQIPSYKMTGDEREGVPRGSFELDPAGLPLS 144
>TC8101 similar to UP|O24074 (O24074) DNAJ-like protein, partial (90%)
Length = 1737
Score = 51.2 bits (121), Expect = 8e-07
Identities = 27/65 (41%), Positives = 39/65 (59%)
Frame = +1
Query: 64 DEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSDREK 123
D +Y +L + ++A + IKK +RK A++ HPDK G FK +G+A VLSD EK
Sbjct: 175 DNTKFYEVLGVPKSASEDEIKKAYRKAAMKNHPDK---GGDPEKFKELGQAYEVLSDPEK 345
Query: 124 RTRYD 128
+ YD
Sbjct: 346 KELYD 360
>CB828171
Length = 570
Score = 50.1 bits (118), Expect = 2e-06
Identities = 20/47 (42%), Positives = 30/47 (63%)
Frame = +3
Query: 733 AQFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYYGQIKKVVTSPTIE 779
A ++F+ +S + Q QIWA Y D D MP YY QIKK+ ++P ++
Sbjct: 285 ASCYDFKKEKSREMLQCDQIWAIYGDRDNMPHYYAQIKKIESTPLLD 425
Score = 45.1 bits (105), Expect = 5e-05
Identities = 36/120 (30%), Positives = 50/120 (41%), Gaps = 5/120 (4%)
Frame = +3
Query: 362 SYRETAKTNDQNGLAADPKNEHEKVKQKQEEKIRAGGKEAAEGSK-QMDKTFEHSSPGST 420
S E +T+D+ L D N H K RA K+ G T ++
Sbjct: 54 SQPEGFQTHDKTPLKHDTNN-HGNETLKVRRSTRAFSKKTHLGDAGDCSDTDKNMKDSVF 230
Query: 421 SKTSNCPNAYVYPD----AEFSDFDKDRKKECFAPGQIWAIYDSIDGMPRFYALIRKVLS 476
S+ A+ D A DF K++ +E QIWAIY D MP +YA I+K+ S
Sbjct: 231 SEPDGSDRAFSKKDCSVGASCYDFKKEKSREMLQCDQIWAIYGDRDNMPHYYAQIKKIES 410
>BP059356
Length = 436
Score = 36.2 bits (82), Expect(2) = 4e-06
Identities = 17/58 (29%), Positives = 34/58 (58%)
Frame = +3
Query: 39 PVLENIAQMLVVCDVHCSAEQKVFGDEINWYGILQLERTAGDAMIKKQFRKFALQLHP 96
PV +++ + + +K G +++Y +LQ++R+A D +KK +RK A++ HP
Sbjct: 15 PVTQSLTHIAPAPPQNSG*SEKAMG--VDYYKVLQVDRSAKDDDLKKAYRKLAMKWHP 182
Score = 32.0 bits (71), Expect(2) = 4e-06
Identities = 21/49 (42%), Positives = 25/49 (50%), Gaps = 2/49 (4%)
Frame = +1
Query: 94 LHPDKNKFAGAEAAFKLIGEAQRVLSDREKRTRYDM--KLNVNKTAMPP 140
L + N AEA FK I EA VLSD +KR YD + +N PP
Sbjct: 181 LDKNPNNKKEAEANFKKISEAYDVLSDPQKRAVYDQFGEEGLNGQVPPP 327
>TC16983 similar to UP|Q9ZSY2 (Q9ZSY2) ALTERED response to gravity (ARG1
protein), partial (42%)
Length = 716
Score = 48.5 bits (114), Expect = 5e-06
Identities = 26/61 (42%), Positives = 35/61 (56%), Gaps = 1/61 (1%)
Frame = +2
Query: 69 YGILQLERTAGDAMIKKQFRKFALQLHPDKN-KFAGAEAAFKLIGEAQRVLSDREKRTRY 127
Y +L + R + D IK +RK AL+ HPDKN A FK + + +LSD EKR +Y
Sbjct: 245 YEVLSVSRDSTDQEIKTAYRKLALKYHPDKNVNNPEASELFKEVAYSYSILSDPEKRRQY 424
Query: 128 D 128
D
Sbjct: 425 D 427
>TC15224 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37,
partial (71%)
Length = 1031
Score = 47.4 bits (111), Expect = 1e-05
Identities = 29/75 (38%), Positives = 40/75 (52%), Gaps = 6/75 (8%)
Frame = +3
Query: 60 KVFG------DEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGE 113
K+FG D +Y IL + + A +KK ++K A++ HPDK G FK + +
Sbjct: 135 KMFGRAPKKSDSTRYYEILGVPKNASQDDLKKAYKKAAIKNHPDKG---GDPEKFKELAQ 305
Query: 114 AQRVLSDREKRTRYD 128
A VLSD EKR YD
Sbjct: 306 AYEVLSDPEKREIYD 350
>TC19683 similar to UP|Q9S7L6 (Q9S7L6) AT2G05250 protein
(AT2G05250/F5G3.15), partial (9%)
Length = 555
Score = 46.6 bits (109), Expect = 2e-05
Identities = 26/63 (41%), Positives = 38/63 (60%), Gaps = 1/63 (1%)
Frame = +1
Query: 891 VTGFSSVFRGKSIEGSSGNLRIPKKELLRFSHQIPAFKLTEEHGDL-RGFWELDPGALPP 949
++GF +V+ KS S IP++E+LRFSHQ+P++ L E +L W+LDP A P
Sbjct: 55 LSGFKTVY--KSNADKSAIKWIPRREMLRFSHQVPSWLLKGEASNLPERCWDLDPAATPD 228
Query: 950 SFL 952
L
Sbjct: 229 DLL 237
Score = 38.5 bits (88), Expect = 0.005
Identities = 27/79 (34%), Positives = 42/79 (52%), Gaps = 1/79 (1%)
Frame = +1
Query: 584 GVHVAYLGKLKGFVSIFIQIMKEDNQPFQ-IPSAELFRFSHRIPSFKMTGQEGVDVHLGY 642
GV V L KL GF +++ D + IP E+ RFSH++PS+ + G E ++
Sbjct: 28 GVCVTPLIKLSGFKTVYKS--NADKSAIKWIPRREMLRFSHQVPSWLLKG-EASNLPERC 198
Query: 643 LEFDPASLPMNLEEIAVTQ 661
+ DPA+ P +L A T+
Sbjct: 199 WDLDPAATPDDLLHAAATE 255
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.132 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,670,694
Number of Sequences: 28460
Number of extensions: 235898
Number of successful extensions: 1588
Number of sequences better than 10.0: 75
Number of HSP's better than 10.0 without gapping: 1507
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1550
length of query: 954
length of database: 4,897,600
effective HSP length: 99
effective length of query: 855
effective length of database: 2,080,060
effective search space: 1778451300
effective search space used: 1778451300
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC147364.8


