

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
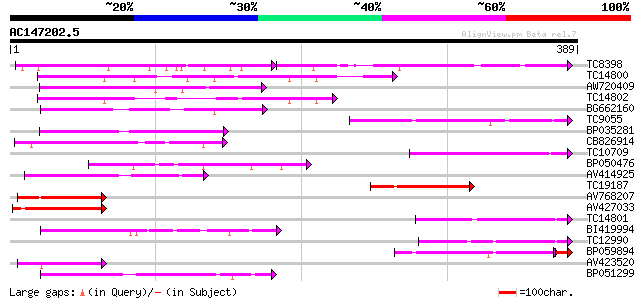
Query= AC147202.5 - phase: 0
(389 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8398 weakly similar to UP|Q84KK1 (Q84KK1) S haplotype-specific... 74 6e-30
TC14800 weakly similar to UP|Q9FHP3 (Q9FHP3) Genomic DNA, chromo... 125 1e-29
AW720409 97 5e-21
TC14802 weakly similar to UP|CAD56661 (CAD56661) S locus F-box (... 96 1e-20
BG662160 85 2e-17
TC9055 84 4e-17
BP035281 80 8e-16
CB826914 71 4e-13
TC10709 68 2e-12
BP050476 68 3e-12
AV414925 67 7e-12
TC19187 65 2e-11
AV768207 63 1e-10
AV427033 62 1e-10
TC14801 62 2e-10
BI419994 60 8e-10
TC12990 59 1e-09
BP059894 57 3e-09
AV423520 55 2e-08
BP051299 54 3e-08
>TC8398 weakly similar to UP|Q84KK1 (Q84KK1) S haplotype-specific F-box
protein a, partial (9%)
Length = 1602
Score = 73.9 bits (180), Expect(2) = 6e-30
Identities = 66/217 (30%), Positives = 99/217 (45%), Gaps = 14/217 (6%)
Frame = +1
Query: 184 YLQKGMTGVFSLGDNVWRNIESFP--------LGYYLD-NRVHLRDSLNWLGLRSYVDDC 234
+ + + V+++GD WR I+SFP G D N V+L ++NW+ LR +
Sbjct: 715 HCMQSVANVYNMGDGCWRRIQSFPDFTREHWGRGTAPDQNGVYLNGTINWV-LRLNIA-- 885
Query: 235 DDYDCEYITSIEQFMIVALDLRTETCKELLLP----RGFDEVPCYEPSLCVLMDCICFSH 290
YD I++LDL +E C L LP G V P L VL D +C H
Sbjct: 886 --YD-----------IISLDLGSEVCSRLSLPCFPPPGPKCVYDLPPILGVLKDSLCVLH 1026
Query: 291 LVKKTHLVIWKMMDYGDDDSWTQLLEINLQILKKIDEWSAWVPLHLSKNYDTLILGSTLE 350
VIWKM ++G +SWT L + K ++ + S+N + L++ T +
Sbjct: 1027NNNDKTFVIWKMNEFGAHESWTPLFNFDF----KKEDLYRFDKFFFSENDEVLLMTLTKQ 1194
Query: 351 HEFAVYNLRDGSIE-KTRITNGESSWIYINDYVESLV 386
+YN D + + R+ N W I +YVESLV
Sbjct: 1195---VLYNHSDITFKFNDRVHNYIHDWNNIENYVESLV 1296
Score = 73.6 bits (179), Expect(2) = 6e-30
Identities = 72/225 (32%), Positives = 107/225 (47%), Gaps = 46/225 (20%)
Frame = +3
Query: 5 PPK--SQQPPSNVASS---PIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFV 59
PP+ + PP+ VA + P IL +LI E+LS LPVKSL++FRCVC SWK+ IS+ F+
Sbjct: 12 PPQHDAATPPTLVADTEPPPPILIDELIWEILSLLPVKSLVRFRCVCKSWKLTISNPQFM 191
Query: 60 KLHLQRS-------IQNPRLAVTQESRECSMDVIVSPTSLSH----LLENPSKPTTLT-- 106
KLHL+RS + ++ V + + ++S + H + P L
Sbjct: 192 KLHLRRSSAKAAADFADSQVVVMTKRDHIAHSPLLSKFHIVHDETFTADEPWPVHALREP 371
Query: 107 -NDPYYSL-----------NDK---DCRSVAGSCNGLLCLL-----GLSEDREMWLRFWN 146
N P+ S+ NDK CR + G+CNGL+ ++ ++ N
Sbjct: 372 FNQPHVSICYAPSLFSSQNNDKLKGHCRLI-GACNGLVSVINEGYNSAESATKLSCCNLN 548
Query: 147 PATR---AISDKLG----HYPADFTGGFEV-AFGYDNSTDTYKVV 183
PAT+ +S L + + F + FGYD DTYKVV
Sbjct: 549 PATKLRSQVSPTLSLCYKQFHSLIHSVFRIFGFGYDPLNDTYKVV 683
>TC14800 weakly similar to UP|Q9FHP3 (Q9FHP3) Genomic DNA, chromosome 5, TAC
clone:K14B20, partial (7%)
Length = 782
Score = 125 bits (314), Expect = 1e-29
Identities = 96/266 (36%), Positives = 134/266 (50%), Gaps = 19/266 (7%)
Frame = +3
Query: 20 IILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHL--QRSIQNPRLAVTQE 77
+ LP +LI E+LS+LPVKSLLQFR V +WK ISD FVKLHL + S +N T
Sbjct: 42 LCLPDELIIEILSWLPVKSLLQFRVVSKTWKSFISDPQFVKLHLLHRLSFRNADFEHTSL 221
Query: 78 SRECSMD----VIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRS---VAGSCNGLLC 130
+C D +S ++S LLE+PS + + C S G+CNGL+
Sbjct: 222 LIKCHTDDFGRPYISSRTVSSLLESPSA----------IVASRSCISGYDFIGTCNGLVS 371
Query: 131 LLGLSEDRE-----MWLRFWNPATRAISDKLGHYPADFTGGFEVAFGYDNSTDTYKVVYL 185
L L+ D +RFWNPATR +S P+ + FGYD S+DTYKVV +
Sbjct: 372 LRKLNYDESNTNNFSQVRFWNPATRTMSQ--DSPPSWSPRNLHLGFGYDCSSDTYKVVGM 545
Query: 186 QKGMT--GVFSLGDNVWRNIESFPLG--YYLDNRVHLRDSLNWLGLRSYVDDCDDYDCEY 241
G+T V+++GDN WR I+ P + + V++ ++LNWL
Sbjct: 546 IPGLTMVNVYNMGDNCWRTIQISPHAPMHLQGSAVYVSNTLNWL---------------- 677
Query: 242 ITSIEQFMIVALDLRTETC-KELLLP 266
+ IV+ DL E C ++L LP
Sbjct: 678 -AGTNPYFIVSFDLEKEECAQQLSLP 752
>AW720409
Length = 575
Score = 97.1 bits (240), Expect = 5e-21
Identities = 60/162 (37%), Positives = 85/162 (52%), Gaps = 6/162 (3%)
Frame = +3
Query: 21 ILPLDLITELLSFLPVKSLLQFRCVCMSWKILI-SDSFFVKLHLQRSIQNPRLAVTQESR 79
+LP +L E+LS+LPVK+L++F CV SWK LI D F KLHL RS +N + +
Sbjct: 90 VLPFELWIEILSWLPVKTLMRFSCVSKSWKSLIYQDRDFKKLHLDRSPKNNNQVILTLQK 269
Query: 80 ECSMDVIVSPTSLSHLLEN--PSKPTTLTNDPYYSLNDKD---CRSVAGSCNGLLCLLGL 134
P + LL++ PS + + + P+ ++ + GSCNGL+CL G+
Sbjct: 270 PLFSGRTFFPFPVRRLLQDQEPSSSSIIDDIPFEEEDEDEEDNYHDTIGSCNGLVCLYGI 449
Query: 135 SEDREMWLRFWNPATRAISDKLGHYPADFTGGFEVAFGYDNS 176
+ D +W R WNPATR K A AFGYD+S
Sbjct: 450 N-DHGLWFRLWNPATRFRFHKSPPLDAVIGSTLHFAFGYDHS 572
>TC14802 weakly similar to UP|CAD56661 (CAD56661) S locus F-box (SLF)-S4
protein, partial (10%)
Length = 551
Score = 95.9 bits (237), Expect = 1e-20
Identities = 73/212 (34%), Positives = 98/212 (45%), Gaps = 6/212 (2%)
Frame = +3
Query: 20 IILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHL--QRSIQNPRLAVTQE 77
+ LP +LI E+LS+LPVKSLLQFR V +WK ISD FVKLHL + S +N T
Sbjct: 15 LCLPDELIIEILSWLPVKSLLQFRVVSKTWKSFISDPQFVKLHLLHRLSFRNADFEHTSL 194
Query: 78 SRECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGLSED 137
+C D PY S
Sbjct: 195 LIKCHTDDF--------------------GRPYISSRT---------------------- 248
Query: 138 REMWLRFWNPATRAISDKLGHYPADFTGGFEVAFGYDNSTDTYKVVYLQKGMT--GVFSL 195
+RFWNPATR +S P+ + FGYD S+DTYKVV + G+T V+++
Sbjct: 249 ----VRFWNPATRTMSQDSP--PSWSPRNLHLGFGYDCSSDTYKVVGMIPGLTMVNVYNM 410
Query: 196 GDNVWRNIESFPLG--YYLDNRVHLRDSLNWL 225
GDN WR I+ P + + V++ ++LNWL
Sbjct: 411 GDNCWRTIQISPHAPMHLQGSAVYVSNTLNWL 506
>BG662160
Length = 445
Score = 84.7 bits (208), Expect = 2e-17
Identities = 57/160 (35%), Positives = 78/160 (48%), Gaps = 4/160 (2%)
Frame = +2
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAVTQESREC 81
LP +L E+LS+LPVKSL++FRCV SWK +ISDS F+KLHL RS R
Sbjct: 23 LPEELRLEILSWLPVKSLVRFRCVSKSWKSIISDSQFIKLHLHRSSSTTR---------- 172
Query: 82 SMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGLSEDRE-- 139
+ D + ++ ++ S T D Y G+CNGL+CL DR+
Sbjct: 173 NTDFAYLQSLITSPRKSRSASTIAIPDDY---------DFCGTCNGLVCLHSSKYDRKDK 325
Query: 140 --MWLRFWNPATRAISDKLGHYPADFTGGFEVAFGYDNST 177
+R WNPA R++S + F YD+ST
Sbjct: 326 YTSHVRLWNPAMRSMSQRSPPLYLPIVYDSYFGFDYDSST 445
>TC9055
Length = 577
Score = 84.0 bits (206), Expect = 4e-17
Identities = 51/155 (32%), Positives = 84/155 (53%), Gaps = 2/155 (1%)
Frame = +2
Query: 234 CDDYDCEYITSIEQFMIVALDLRTETCKELLLPRGFDEVPCYEPSLCVLMDCICFSHLVK 293
C ++ EQ +IV+LD+R E + L LP+G E+P E + VL +C+C + K
Sbjct: 5 CVNWLANKFEPFEQLVIVSLDMRQEAYRLLSLPQGTSELPSAE--IQVLGNCLCLFYDFK 178
Query: 294 KTHLVIWKMMDYGDDDSWTQLLEINLQILKKIDEW--SAWVPLHLSKNYDTLILGSTLEH 351
+TH V WKM +YG +SWT LL I+ + L+ + + W+ + + + + +
Sbjct: 179 RTHFVAWKMSEYGLPESWTPLLTISYEHLRWDHRFYLNQWLICLCGDGH--IFMLAKKQK 352
Query: 352 EFAVYNLRDGSIEKTRITNGESSWIYINDYVESLV 386
+YNL D S++ + + + W+ NDYVESLV
Sbjct: 353 LVIIYNLSDNSVKYVELASNK-LWLDANDYVESLV 454
>BP035281
Length = 498
Score = 79.7 bits (195), Expect = 8e-16
Identities = 53/132 (40%), Positives = 72/132 (54%), Gaps = 2/132 (1%)
Frame = +2
Query: 21 ILPLDLITELLSFLPVKSLLQFRCVCMSWKILIS-DSFFVKLHLQR-SIQNPRLAVTQES 78
+ P +L+ E+LS+LPV SL++F+CVC SWK LIS + F KL L+R S++ + +T
Sbjct: 119 VFPSELVIEILSWLPVLSLIRFKCVCKSWKSLISHNKAFAKLQLERTSLKINHVILT--- 289
Query: 79 RECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGLSEDR 138
S D P SL LLE+PS D L+ + V GSCNGL+CL
Sbjct: 290 ---SSDSSFIPCSLPRLLEDPSSIMDQDQDRCLRLDKFEYNEVIGSCNGLICLHADYCAP 460
Query: 139 EMWLRFWNPATR 150
R NP+TR
Sbjct: 461 GFCFRPCNPSTR 496
>CB826914
Length = 575
Score = 70.9 bits (172), Expect = 4e-13
Identities = 49/155 (31%), Positives = 77/155 (49%), Gaps = 9/155 (5%)
Frame = +1
Query: 4 SPPKSQQPPS---NVASSPIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVK 60
SPP++ + + + I + DLI E+L LPVKS+++F+ VC W+ LISD F
Sbjct: 103 SPPEAASVSTIEIQIQTLHIDMDRDLIIEILLMLPVKSIVRFKAVCKLWRSLISDPLFAS 282
Query: 61 LHLQRSIQNPRLAVTQESRECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLN-DKDCR 119
LH QR+ + A + R ++ + SH + P L+N Y+ L+ ++C
Sbjct: 283 LHFQRAAPSLLFADSHAIRTIDLEGPLQ----SHRVSQPINCHFLSN--YHDLSLPRNCI 444
Query: 120 SVAGSCNGLLCL-----LGLSEDREMWLRFWNPAT 149
S+ GSC G L + +G L WNP+T
Sbjct: 445 SIHGSCRGFLLITWISNIGYGHPWNDSLYLWNPST 549
>TC10709
Length = 578
Score = 68.2 bits (165), Expect = 2e-12
Identities = 45/112 (40%), Positives = 61/112 (54%)
Frame = +3
Query: 275 YEPSLCVLMDCICFSHLVKKTHLVIWKMMDYGDDDSWTQLLEINLQILKKIDEWSAWVPL 334
+EPSL VL + + H HL +WKM+ YG +SWTQLL I+ + L I VP+
Sbjct: 6 FEPSLKVLGNHMYLFHDHNSIHLFVWKMVKYGALESWTQLLSISYERLHCIGFPYIPVPM 185
Query: 335 HLSKNYDTLILGSTLEHEFAVYNLRDGSIEKTRITNGESSWIYINDYVESLV 386
L ++ D L++ T EF YN+RD S+E N E W+ YV SLV
Sbjct: 186 FLLEDGDILMIAITENLEFIKYNIRDYSMEYIPNPN-EVFWMDAYGYVPSLV 338
>BP050476
Length = 520
Score = 67.8 bits (164), Expect = 3e-12
Identities = 59/172 (34%), Positives = 78/172 (45%), Gaps = 19/172 (11%)
Frame = +2
Query: 55 DSFFVKLHLQRSIQNPRLAVTQESRECSM---DVIVSPTSLSHLLENPSKPTTLTNDPYY 111
DS FVKLHL RS +N + + + D V P+S+ L+E+ S + + Y
Sbjct: 17 DSSFVKLHLNRSPKNTHILLNIANDPYDFENDDTWVVPSSVCCLIEDLS--SMIDAKGCY 190
Query: 112 SLNDKDCRSVAGSCNGLLCL---LGLSEDREMWLRFWNPATRAISDKLGHYPADFT---- 164
L KD V GS NGL+C + E W++ WNPAT S K +
Sbjct: 191 LL--KDGHLVIGSSNGLICFGNFYDVGPIEEFWVQLWNPATHLKSKKSPTFNLSMRTSVD 364
Query: 165 ---GGFEVAFGYDNSTDTYKVVYL------QKGMTGVFSLGDNVWRNIESFP 207
G + FGYDN DTYKVV + QK T V+ + D R I S P
Sbjct: 365 APPGKVNLGFGYDNLHDTYKVVLVHWNCTKQKMETMVYCVNDIFCRKILSDP 520
>AV414925
Length = 414
Score = 66.6 bits (161), Expect = 7e-12
Identities = 40/127 (31%), Positives = 69/127 (53%), Gaps = 1/127 (0%)
Frame = +1
Query: 11 PPSNVASSPIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNP 70
PP N +P+++I ++LS LPV SLL+FR + SW+ LI F+KLHL S+ NP
Sbjct: 64 PPQNPTLMAEEIPVEVIADILSRLPVTSLLRFRSISKSWRSLIDSKHFMKLHLNNSLTNP 243
Query: 71 RLAVTQESRECSMDVIVSPTSLSHLLENPSKPTTLT-NDPYYSLNDKDCRSVAGSCNGLL 129
+T ++I+ + + + PS + N P +++ ++ GSC+GLL
Sbjct: 244 NPNLT--------NLILRHNTDLYRADFPSIGAAVNLNHPLMCYSNR--INILGSCHGLL 393
Query: 130 CLLGLSE 136
C+ +++
Sbjct: 394 CICNVAD 414
>TC19187
Length = 632
Score = 65.1 bits (157), Expect = 2e-11
Identities = 31/72 (43%), Positives = 48/72 (66%)
Frame = +1
Query: 248 FMIVALDLRTETCKELLLPRGFDEVPCYEPSLCVLMDCICFSHLVKKTHLVIWKMMDYGD 307
++IV+LDL+ ET KE+L P +++ C P+L VL C+C ++ K+T V+W M DYG
Sbjct: 70 WIIVSLDLQKETYKEIL-PPDYEKEECSTPTLSVLKGCLCMNYDHKRTDFVVWMMKDYGV 246
Query: 308 DDSWTQLLEINL 319
+SW +L+ I L
Sbjct: 247 RESWVKLVTIPL 282
>AV768207
Length = 374
Score = 62.8 bits (151), Expect = 1e-10
Identities = 34/61 (55%), Positives = 38/61 (61%)
Frame = +1
Query: 6 PKSQQPPSNVASSPIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQR 65
P S PP + P LP +L E+L LPVKSLLQ RCVC SWK LISD F K HL+
Sbjct: 28 PSSSNPPGDSPPFPT-LPFELGVEILCRLPVKSLLQLRCVCKSWKSLISDPKFAKNHLRC 204
Query: 66 S 66
S
Sbjct: 205S 207
>AV427033
Length = 420
Score = 62.4 bits (150), Expect = 1e-10
Identities = 34/64 (53%), Positives = 39/64 (60%)
Frame = +1
Query: 3 LSPPKSQQPPSNVASSPIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLH 62
L+PP PP+ + LP DL+ E+L LPV SLLQFRCVC SW LISD F K H
Sbjct: 73 LTPPSL--PPAGDSLHAPPLPFDLVVEILCRLPVNSLLQFRCVCKSWNSLISDPKFAKNH 246
Query: 63 LQRS 66
L S
Sbjct: 247 LHCS 258
>TC14801
Length = 595
Score = 62.0 bits (149), Expect = 2e-10
Identities = 38/109 (34%), Positives = 60/109 (54%), Gaps = 1/109 (0%)
Frame = +2
Query: 279 LCVLMDCICFSHLVKKTH-LVIWKMMDYGDDDSWTQLLEINLQILKKIDEWSAWVPLHLS 337
L VL C+C S K+TH V+ +M +YG ++SWTQL I I+ + S + +S
Sbjct: 101 LGVLRGCLCISQYHKETHSFVVSQMKEYGVNESWTQLFNI---IVHERILHSPCFAMCMS 271
Query: 338 KNYDTLILGSTLEHEFAVYNLRDGSIEKTRITNGESSWIYINDYVESLV 386
+N D L+ + + +YN RD +E T+I N Y+ +Y+ESL+
Sbjct: 272 ENGDVLLFAESGVSQAVLYNQRDNKLEDTKIANNIVG-SYVRNYIESLI 415
>BI419994
Length = 549
Score = 59.7 bits (143), Expect = 8e-10
Identities = 53/174 (30%), Positives = 79/174 (44%), Gaps = 9/174 (5%)
Frame = +2
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAVTQESREC 81
LP +LI ++L LPV+SLL+F+ VC SW+ LISD F K H + +
Sbjct: 47 LPDELILQILLRLPVRSLLRFKTVCKSWRYLISDPQFTKSHFNLAASPTHRIFLNNTNGF 226
Query: 82 ---SMDV---IVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGLS 135
S+D + +SL H L+ P P N + S V GSC G + L
Sbjct: 227 *IESLDTDSSLHDESSLVH-LKFPLAPPHPDNHRFGS--SVTGLKVLGSCRGFVLL---- 385
Query: 136 EDREMWLRFWNPAT---RAISDKLGHYPADFTGGFEVAFGYDNSTDTYKVVYLQ 186
+ + + WNP+T R + + + + F GYD S D Y VV+++
Sbjct: 386 ANHQGNVVVWNPSTGVRRRVPEWCSYLASQCE--FLYGIGYDESNDDYLVVFIE 541
>TC12990
Length = 518
Score = 58.9 bits (141), Expect = 1e-09
Identities = 36/107 (33%), Positives = 60/107 (55%), Gaps = 1/107 (0%)
Frame = +1
Query: 281 VLMDCICFSHLVKKTH-LVIWKMMDYGDDDSWTQLLEINLQILKKIDEWSAWVPLHLSKN 339
VL C+C S K+TH V+W+M ++G +SWTQL N+ + ++I + + +S+N
Sbjct: 4 VLGGCLCISQNNKETHSFVVWQMKEFGVHESWTQL--FNIIVHERILHTPCFA-MCMSEN 174
Query: 340 YDTLILGSTLEHEFAVYNLRDGSIEKTRITNGESSWIYINDYVESLV 386
D L+ + + +YN R+ E T+I N Y+ +Y+ESLV
Sbjct: 175 GDALLFAKSGVSQAVLYNQREKKFEDTKIGNNIVG-SYVRNYIESLV 312
>BP059894
Length = 474
Score = 56.6 bits (135), Expect(2) = 3e-09
Identities = 35/114 (30%), Positives = 59/114 (51%), Gaps = 2/114 (1%)
Frame = -3
Query: 265 LPRGFDEVPCYEPSLCVLMDCICFSHLVKKTHLVIWKMMDYGDDDSWTQLLEINLQILKK 324
LP+G E+P E + VL +C+C H +TH V WKM +YG +SWT++L I+ Q +
Sbjct: 469 LPKGTSELPYVE--IQVLGNCLCLFHDDNRTHFVGWKMTEYGVPESWTRMLSISYQHFQC 296
Query: 325 IDE--WSAWVPLHLSKNYDTLILGSTLEHEFAVYNLRDGSIEKTRITNGESSWI 376
D W+ + L + + L+L ++NL D S++ + + W+
Sbjct: 295 NDVILRHRWL-VCLCGDGNILMLAKDEGGLVIIFNLSDNSVKYIKFPPKDFGWV 137
Score = 20.8 bits (42), Expect(2) = 3e-09
Identities = 7/12 (58%), Positives = 9/12 (74%)
Frame = -2
Query: 375 WIYINDYVESLV 386
W+ DYVESL+
Sbjct: 146 WLGAEDYVESLI 111
>AV423520
Length = 468
Score = 55.5 bits (132), Expect = 2e-08
Identities = 32/65 (49%), Positives = 38/65 (58%), Gaps = 4/65 (6%)
Frame = +3
Query: 6 PKSQQPPSNVASSPI---ILPLDLITELLSFLPVKSLLQFRCVCMSWKILIS-DSFFVKL 61
P S P + P LP +L+ E+L LPVKSLLQFRCVC SW LIS D F +
Sbjct: 108 PSSSNPHGDSLHPPPPLPTLPFELVVEILCRLPVKSLLQFRCVCKSWNSLISGDPKFARK 287
Query: 62 HLQRS 66
HL+ S
Sbjct: 288 HLRCS 302
>BP051299
Length = 539
Score = 54.3 bits (129), Expect = 3e-08
Identities = 43/164 (26%), Positives = 75/164 (45%), Gaps = 2/164 (1%)
Frame = +2
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAVTQESREC 81
LP ++ E+L LP K+L++ VC SW+ LI+ + F+ LH S + L + E
Sbjct: 5 LPQEIWAEILHRLPPKTLVKCTSVCKSWRSLITSTSFISLHRNHSPSSLLLQLCDER--- 175
Query: 82 SMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGLSEDREMW 141
P + + L + ++ + SV G C+GL+C+ R++
Sbjct: 176 -----AIPNFIHYSLRRDDPFLSESSSLRLPSSFNREFSVVGICHGLVCITSTENCRDLI 340
Query: 142 LRFWNPATRA--ISDKLGHYPADFTGGFEVAFGYDNSTDTYKVV 183
+ NP+ R + YP+ + VAFG+D+ + YKV+
Sbjct: 341 I--CNPSLRLHITLPEPSDYPSLHSAS--VAFGFDSRNNDYKVI 460
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.139 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,794,658
Number of Sequences: 28460
Number of extensions: 144526
Number of successful extensions: 960
Number of sequences better than 10.0: 84
Number of HSP's better than 10.0 without gapping: 920
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 935
length of query: 389
length of database: 4,897,600
effective HSP length: 92
effective length of query: 297
effective length of database: 2,279,280
effective search space: 676946160
effective search space used: 676946160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC147202.5


