

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147179.9 + phase: 1 /partial
(375 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
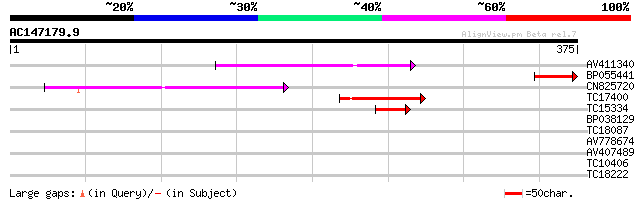
Score E
Sequences producing significant alignments: (bits) Value
AV411340 97 3e-21
BP055441 57 5e-09
CN825720 53 9e-08
TC17400 similar to UP|Q9C5B2 (Q9C5B2) Translation releasing fact... 49 1e-06
TC15334 43 7e-05
BP038129 37 0.007
TC18087 similar to UP|Q9LF46 (Q9LF46) 2-hydroxyphytanoyl-CoA lya... 29 1.5
AV778674 28 2.5
AV407489 27 4.2
TC10406 similar to UP|Q94A21 (Q94A21) AT5g15330/F8M21_220, parti... 27 5.5
TC18222 similar to UP|SY32_ARATH (Q9LK09) Syntaxin 32 (AtSYP32),... 27 7.2
>AV411340
Length = 391
Score = 97.4 bits (241), Expect = 3e-21
Identities = 51/132 (38%), Positives = 77/132 (57%)
Frame = +2
Query: 137 GTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKG 196
G +A W L+RMY R+ E+ +K + S E+ G K+ +E++G Y L E
Sbjct: 2 GDEAGIWVGDLVRMYERYSERNSWKHSTISSSEAEKGGFKTFVMEIKGNCVYSKLKYESS 181
Query: 197 THRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNK 256
HR+ R ++G TS + V +MP + E + VEI +D+E++ +R+GG GGQNVNK
Sbjct: 182 VHRVQRVPLTETQGRVHTSTATVAIMPEVDE--VEVEIDPKDIELTTARSGGAGGQNVNK 355
Query: 257 VETAVRITHIPT 268
VETA+ + H PT
Sbjct: 356 VETAIDLFHKPT 391
>BP055441
Length = 394
Score = 57.0 bits (136), Expect = 5e-09
Identities = 26/28 (92%), Positives = 28/28 (99%)
Frame = -1
Query: 348 SVIDGELDPFIKSYLKHKYSMTMSTSGV 375
SV+DGELDPFIKSYLKHK+SMTMSTSGV
Sbjct: 394 SVMDGELDPFIKSYLKHKFSMTMSTSGV 311
>CN825720
Length = 699
Score = 52.8 bits (125), Expect = 9e-08
Identities = 40/170 (23%), Positives = 80/170 (46%), Gaps = 9/170 (5%)
Frame = +1
Query: 24 LQLLEQELAKLEEEASCSSFW--------DDRAKAQQTLSTLADVKEKIKLLNDYKTQVE 75
L ++EQ L+ +E ++C S + A+A + L L+ E I L + +++
Sbjct: 181 LNIMEQRLSAIEHRSACLSNLLNQPEATPSEYARANKELRKLSGSVELINELRAKQKEID 360
Query: 76 DAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLL-SGPYDKEGAVISITAG 134
+T++ + E + + EE KE + + L LL D+ +++ + AG
Sbjct: 361 GLKTLMAESSEDKDMLSMATEEFGQA-KEEERRLQNMLLKSLLPKDDADERDSILEVRAG 537
Query: 135 AGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEG 184
+GG +A +A + +MY ++ K +K VV+ + + G K A+ + G
Sbjct: 538 SGGEEASLFAMDIFKMYEKYAHKNGWKFEVVDIAQSDMKGYKEASAAIAG 687
>TC17400 similar to UP|Q9C5B2 (Q9C5B2) Translation releasing factor2,
partial (6%)
Length = 572
Score = 49.3 bits (116), Expect = 1e-06
Identities = 31/57 (54%), Positives = 38/57 (66%)
Frame = +3
Query: 219 VEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCT 275
+EVMPLL E+S++ E SR GGKGGQ V+KVE AV+ITH TGVT+ T
Sbjct: 309 LEVMPLL-EKSMDTERLGPGN*FFNSREGGKGGQIVDKVEIAVQITHNQTGVTIH*T 476
>TC15334
Length = 537
Score = 43.1 bits (100), Expect = 7e-05
Identities = 19/23 (82%), Positives = 21/23 (90%)
Frame = +1
Query: 243 FSRAGGKGGQNVNKVETAVRITH 265
FSR GGKGGQ VN+VETA+RITH
Sbjct: 178 FSREGGKGGQTVNQVETALRITH 246
>BP038129
Length = 558
Score = 36.6 bits (83), Expect = 0.007
Identities = 16/29 (55%), Positives = 22/29 (75%)
Frame = +2
Query: 233 EIPEEDLEISFSRAGGKGGQNVNKVETAV 261
+I + + +SF+R+GG GGQNVNKV T V
Sbjct: 263 KITLDHVTVSFARSGGPGGQNVNKVNTKV 349
>TC18087 similar to UP|Q9LF46 (Q9LF46) 2-hydroxyphytanoyl-CoA lyase-like
protein, partial (30%)
Length = 571
Score = 28.9 bits (63), Expect = 1.5
Identities = 19/49 (38%), Positives = 27/49 (54%)
Frame = -2
Query: 93 GLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQ 141
G + S L EL + + RF+L QLL + A IS+ AGAG D++
Sbjct: 456 GDLRDISGLHGELREWLHRFDLVQLL----EVTAASISLIAGAGDHDSR 322
>AV778674
Length = 606
Score = 28.1 bits (61), Expect = 2.5
Identities = 25/123 (20%), Positives = 55/123 (44%)
Frame = +3
Query: 11 ASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDY 70
A Q +KE ++ +Q LE+ +A+L +E + + + + +K + + + Y
Sbjct: 138 AEQAMKEEMDTR-IQTLEKTVARLRDEVK-------KEREDKNMERNRRLKTEKAIKDSY 293
Query: 71 KTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVIS 130
++ + + E + K L +E L + + ++ QLLSGP + + I
Sbjct: 294 NNVEQEKKQFINELERHKEALKRLSDEVEKLKIVVGNLPEGSDVAQLLSGPNVDDFSAIY 473
Query: 131 ITA 133
++A
Sbjct: 474 MSA 482
>AV407489
Length = 415
Score = 27.3 bits (59), Expect = 4.2
Identities = 17/63 (26%), Positives = 33/63 (51%)
Frame = -1
Query: 214 TSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITHIPTGVTLR 273
++FS + +LP++S N+E P+ L+ ++ + E AV+ IP+G+ +R
Sbjct: 322 SAFSTSLIGKILPKKSPNIEFPQSHLKFLLVQSYTI**PSWQHFEQAVQTPSIPSGMQVR 143
Query: 274 CTE 276
E
Sbjct: 142 QEE 134
>TC10406 similar to UP|Q94A21 (Q94A21) AT5g15330/F8M21_220, partial (57%)
Length = 986
Score = 26.9 bits (58), Expect = 5.5
Identities = 27/101 (26%), Positives = 43/101 (41%)
Frame = +3
Query: 24 LQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVEDAETIVML 83
L++L QEL K + F+ D K ++ + ++KE+I+ L D Q E
Sbjct: 9 LRILNQELEKFND------FYVD--KEEEFVIRFQELKERIERLKDKSYQREMYTFDYEF 164
Query: 84 TEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDK 124
+EEM + K L ++ N S F + YDK
Sbjct: 165 SEEMMEIRKDLVTIHGEMVLLKNYSSLNFAGLIKILKKYDK 287
>TC18222 similar to UP|SY32_ARATH (Q9LK09) Syntaxin 32 (AtSYP32), partial
(37%)
Length = 493
Score = 26.6 bits (57), Expect = 7.2
Identities = 21/83 (25%), Positives = 40/83 (47%)
Frame = +2
Query: 23 GLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVEDAETIVM 82
G+ Q+LAKL + A +S +DD Q L+++ +K+ I LN + V D + +
Sbjct: 182 GIHQTSQKLAKLAKLAKRTSVFDDPTMEIQELTSV--IKQDITALN---SAVVDLQLLSN 346
Query: 83 LTEEMESVDKGLYEEASSLIKEL 105
E + +++L+ +L
Sbjct: 347 SRNESGNASADTTSHSTTLVDDL 415
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.131 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,666,365
Number of Sequences: 28460
Number of extensions: 47663
Number of successful extensions: 182
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 181
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 181
length of query: 375
length of database: 4,897,600
effective HSP length: 92
effective length of query: 283
effective length of database: 2,279,280
effective search space: 645036240
effective search space used: 645036240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC147179.9


