

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147179.5 - phase: 0
(706 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
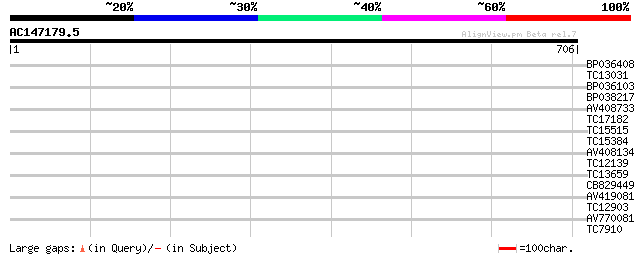
Score E
Sequences producing significant alignments: (bits) Value
BP036408 34 0.070
TC13031 31 0.59
BP036103 30 1.0
BP038217 29 2.2
AV408733 29 2.2
TC17182 similar to UP|Q8L748 (Q8L748) AT3g14120/MAG2_7, partial ... 29 2.9
TC15515 28 3.8
TC15384 weakly similar to UP|Q9LVB8 (Q9LVB8) HSR203J protein-lik... 28 5.0
AV408134 28 6.5
TC12139 28 6.5
TC13659 homologue to UP|Q9M060 (Q9M060) Eukaryotic translation i... 28 6.5
CB829449 28 6.5
AV419081 28 6.5
TC12903 weakly similar to UP|ALA9_ARATH (Q9SX33) Potential phosp... 28 6.5
AV770081 27 8.5
TC7910 homologue to UP|P93166 (P93166) SCOF-1, partial (52%) 27 8.5
>BP036408
Length = 516
Score = 34.3 bits (77), Expect = 0.070
Identities = 12/14 (85%), Positives = 13/14 (92%)
Frame = -1
Query: 690 LCVVCLAICWKHIM 703
LCV+CLAICWKH M
Sbjct: 516 LCVLCLAICWKHFM 475
>TC13031
Length = 462
Score = 31.2 bits (69), Expect = 0.59
Identities = 17/41 (41%), Positives = 25/41 (60%), Gaps = 2/41 (4%)
Frame = +1
Query: 243 FSAPSSSGRTLKEVF--LKHSISAYLCGHLHSRFGKNLKRH 281
F+AP+ SG+ EVF L H+IS L + H +F + +RH
Sbjct: 106 FTAPAKSGKFPHEVFALLNHAISKKLETYTHYQFKRQPRRH 228
>BP036103
Length = 423
Score = 30.4 bits (67), Expect = 1.0
Identities = 13/33 (39%), Positives = 17/33 (51%)
Frame = +2
Query: 481 HSFRLSWSWKEFFVMGCQWASLYYPLLWSALGF 513
HS RLSW E +G W+ PL W + G+
Sbjct: 308 HSHRLSWRDPELVFLGRNWSIRNLPLCWVSRGW 406
>BP038217
Length = 514
Score = 29.3 bits (64), Expect = 2.2
Identities = 13/35 (37%), Positives = 18/35 (51%)
Frame = -2
Query: 465 MGRSTLTELRPFSINGHSFRLSWSWKEFFVMGCQW 499
+G T +RPF++ H F L W KE F Q+
Sbjct: 390 LGPITFQRIRPFNLTIHLFFLLWQLKELFPHSLQF 286
>AV408733
Length = 430
Score = 29.3 bits (64), Expect = 2.2
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = -2
Query: 367 SRFMQTSSWHHNYECQSVASSSYETI 392
SRF +SWHHN+ +SS T+
Sbjct: 78 SRFAAAASWHHNFTVLGAMASSSSTV 1
>TC17182 similar to UP|Q8L748 (Q8L748) AT3g14120/MAG2_7, partial (10%)
Length = 562
Score = 28.9 bits (63), Expect = 2.9
Identities = 26/87 (29%), Positives = 41/87 (46%), Gaps = 7/87 (8%)
Frame = +2
Query: 470 LTELRPFSINGHSFRLSW-----SWKEFFVMGCQWASLYYPLLWS--ALGFMFSFLLVPK 522
LT++ P+S H+ + S K FFV C++ L + L + A+ ++L V K
Sbjct: 62 LTKVVPWSARLHTLLKDFVGGAVSLKSFFV-ACRFLFLSWDLEFYLIAMTLWLNWLAVLK 238
Query: 523 ALLFFQMNLYTYRNFIANKGVVNGALW 549
LF + +YRNF +G A W
Sbjct: 239 LSLFTCLVNSSYRNFYCLRGNTQSAKW 319
>TC15515
Length = 1203
Score = 28.5 bits (62), Expect = 3.8
Identities = 12/30 (40%), Positives = 19/30 (63%)
Frame = +3
Query: 290 SLQNFFQFNVHQNSFESTVNCSIGAPPQEF 319
S+Q+FF+ + Q + E T ++ APP EF
Sbjct: 141 SIQSFFRRGISQPAAERTKEATLPAPPTEF 230
>TC15384 weakly similar to UP|Q9LVB8 (Q9LVB8) HSR203J protein-like protein,
partial (21%)
Length = 1193
Score = 28.1 bits (61), Expect = 5.0
Identities = 13/38 (34%), Positives = 23/38 (60%)
Frame = +1
Query: 64 VHHPNRALDFTNLVGHALSFINPSLVLITGDLTDGKSK 101
+H+PN ++ N + +NP+L +++ DLT KSK
Sbjct: 154 IHNPNGSVTRPNKPPESPPSLNPTLPVLSKDLTINKSK 267
>AV408134
Length = 428
Score = 27.7 bits (60), Expect = 6.5
Identities = 13/32 (40%), Positives = 20/32 (61%)
Frame = -1
Query: 218 LELSHWDSQSEKPVTKISFGHFPLSFSAPSSS 249
+ELS + + + K++ H PLSF APSS+
Sbjct: 419 MELSTFGLEGMQLPKKLNLNHHPLSFFAPSST 324
>TC12139
Length = 492
Score = 27.7 bits (60), Expect = 6.5
Identities = 11/30 (36%), Positives = 20/30 (66%)
Frame = +1
Query: 596 WAVETSNGKGKFEWVGFPDIMVLVLPHILF 625
W++ + KGKF ++ FP+ ++ PH+LF
Sbjct: 298 WSLLFAI*KGKFRFM*FPEFTLIHPPHLLF 387
>TC13659 homologue to UP|Q9M060 (Q9M060) Eukaryotic translation initiation
factor 6 (EIF-6)-like protein (At3g55620), partial (43%)
Length = 411
Score = 27.7 bits (60), Expect = 6.5
Identities = 11/33 (33%), Positives = 17/33 (51%)
Frame = +3
Query: 281 HHQLSNRFLSLQNFFQFNVHQNSFESTVNCSIG 313
HHQ+ N S++ F F++ + NC IG
Sbjct: 39 HHQIFNLITSVKEFCPFSIMATRLQFENNCEIG 137
>CB829449
Length = 561
Score = 27.7 bits (60), Expect = 6.5
Identities = 16/43 (37%), Positives = 19/43 (43%), Gaps = 4/43 (9%)
Frame = +1
Query: 662 NSRRPLLNGNHNSTISTLHLGKRW----IRKLLCVVCLAICWK 700
N R LL +ST L RW IRK+ C C CW+
Sbjct: 61 NETRSLLRPP*HSTNHRPPLTARWRTAGIRKMCCATC*RCCWR 189
>AV419081
Length = 399
Score = 27.7 bits (60), Expect = 6.5
Identities = 13/29 (44%), Positives = 14/29 (47%)
Frame = +3
Query: 489 WKEFFVMGCQWASLYYPLLWSALGFMFSF 517
W FFV C W S PL+W F F F
Sbjct: 237 WSVFFVFWCAWFS-SAPLIWGLSFFPFFF 320
>TC12903 weakly similar to UP|ALA9_ARATH (Q9SX33) Potential
phospholipid-transporting ATPase 9 (Aminophospholipid
flippase 9) , partial (14%)
Length = 746
Score = 27.7 bits (60), Expect = 6.5
Identities = 14/38 (36%), Positives = 20/38 (51%), Gaps = 3/38 (7%)
Frame = -1
Query: 36 CNGETVIEPKGTPDSAVW---VVQLSDLHLSVHHPNRA 70
C E + E G+PD+ + V + HL VH+PN A
Sbjct: 335 CQEEEMPESNGSPDANMLNQGEVPYGECHLQVHNPNHA 222
>AV770081
Length = 244
Score = 27.3 bits (59), Expect = 8.5
Identities = 9/15 (60%), Positives = 11/15 (73%)
Frame = -1
Query: 257 FLKHSISAYLCGHLH 271
F +H+I YLC HLH
Sbjct: 130 FFRHTIPCYLCDHLH 86
>TC7910 homologue to UP|P93166 (P93166) SCOF-1, partial (52%)
Length = 1035
Score = 27.3 bits (59), Expect = 8.5
Identities = 11/21 (52%), Positives = 12/21 (56%)
Frame = +1
Query: 470 LTELRPFSINGHSFRLSWSWK 490
LT+ P SI SFR W WK
Sbjct: 43 LTQFSPLSITQLSFRFRWLWK 105
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.138 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,600,811
Number of Sequences: 28460
Number of extensions: 227626
Number of successful extensions: 1798
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 1786
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1797
length of query: 706
length of database: 4,897,600
effective HSP length: 97
effective length of query: 609
effective length of database: 2,136,980
effective search space: 1301420820
effective search space used: 1301420820
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC147179.5


