

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
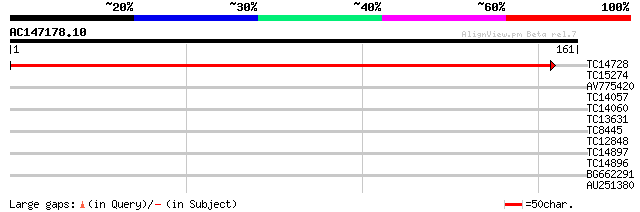
Query= AC147178.10 + phase: 0
(161 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC14728 similar to UP|Q9LSL0 (Q9LSL0) Emb|CAB16809.1, partial (86%) 287 5e-79
TC15274 homologue to UP|Q8GVC8 (Q8GVC8) Enoyl ACP reductase, par... 27 1.7
AV775420 27 2.2
TC14057 similar to GB|AAM62519.1|21553426|AY085287 50S ribosomal... 26 2.9
TC14060 GB|AAA34124.1|170354|TOBUBI4A pentameric polyubiquitin {... 26 2.9
TC13631 similar to UP|Q9SGN7 (Q9SGN7) F3M18.17 (At1g28390/F3M18_... 26 2.9
TC8445 similar to UP|O80910 (O80910) At2g38410 protein, partial ... 26 3.8
TC12848 GB|AAA37921.1|553942|MUSIGF2RA insulin-like growth facto... 26 3.8
TC14897 homologue to UP|RS18_ARATH (P34788) 40S ribosomal protei... 25 6.4
TC14896 homologue to UP|RS18_ARATH (P34788) 40S ribosomal protei... 25 6.4
BG662291 25 6.4
AU251380 25 8.4
>TC14728 similar to UP|Q9LSL0 (Q9LSL0) Emb|CAB16809.1, partial (86%)
Length = 1142
Score = 287 bits (735), Expect = 5e-79
Identities = 139/155 (89%), Positives = 147/155 (94%)
Frame = +3
Query: 1 MKDDDGLPTTTAAINKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLRWSGNLN 60
MKDDDGLPTTTA KK+N+DSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLRWSGNLN
Sbjct: 294 MKDDDGLPTTTAPTAKKDNLDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLRWSGNLN 473
Query: 61 TLSSDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQEAFEAAYEELTSDV 120
+LS+DLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQEAFEAAYEELTSD
Sbjct: 474 SLSNDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQEAFEAAYEELTSDA 653
Query: 121 PGVRDDAITEIAKMSVRSVNYDPPPIQSTVSFPFS 155
PGVRD AITEIAKMSVRS+++DPPPIQST + FS
Sbjct: 654 PGVRDAAITEIAKMSVRSIDFDPPPIQSTRAQEFS 758
>TC15274 homologue to UP|Q8GVC8 (Q8GVC8) Enoyl ACP reductase, partial (25%)
Length = 629
Score = 26.9 bits (58), Expect = 1.7
Identities = 16/41 (39%), Positives = 22/41 (53%)
Frame = +1
Query: 36 AAILLLAFWSMFTGTVSLRWSGNLNTLSSDLDTPIHDDLDV 76
AA L S TGTV L LN + +D+PI ++LD+
Sbjct: 172 AAFLASPLASAITGTV-LYVDNGLNAMGVGVDSPIFENLDI 291
>AV775420
Length = 457
Score = 26.6 bits (57), Expect = 2.2
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = -1
Query: 25 FGKGRYKFWALAAILLLAFWS 45
F KG+Y+FW L ++L WS
Sbjct: 289 FLKGQYRFWRLRVLVLGFVWS 227
>TC14057 similar to GB|AAM62519.1|21553426|AY085287 50S ribosomal protein L3
{Arabidopsis thaliana;} , partial (83%)
Length = 1408
Score = 26.2 bits (56), Expect = 2.9
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -3
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F +SL WSG + +LS
Sbjct: 158 FWML*SARVLPSSSCFPAKMSLCWSGGMPSLS 63
>TC14060 GB|AAA34124.1|170354|TOBUBI4A pentameric polyubiquitin {Nicotiana
sylvestris;} , partial (49%)
Length = 567
Score = 26.2 bits (56), Expect = 2.9
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -2
Query: 32 FWALAAILLLAFWSMFTGTVSLRWSGNLNTLS 63
FW L + +L S F +SL WSG + +LS
Sbjct: 338 FWML*SARVLPSSSCFPAKMSLCWSGGMPSLS 243
>TC13631 similar to UP|Q9SGN7 (Q9SGN7) F3M18.17 (At1g28390/F3M18_17),
partial (8%)
Length = 656
Score = 26.2 bits (56), Expect = 2.9
Identities = 11/30 (36%), Positives = 16/30 (52%)
Frame = +2
Query: 1 MKDDDGLPTTTAAINKKENMDSGLFGKGRY 30
++D DG+ T + E + S FG GRY
Sbjct: 518 LRDADGVEHTQQSSKPAEKVSSSRFGSGRY 607
>TC8445 similar to UP|O80910 (O80910) At2g38410 protein, partial (7%)
Length = 841
Score = 25.8 bits (55), Expect = 3.8
Identities = 14/33 (42%), Positives = 16/33 (48%)
Frame = +3
Query: 129 TEIAKMSVRSVNYDPPPIQSTVSFPFSTHGAVS 161
T A S S PP +TVS P TH A+S
Sbjct: 54 TFTASASTSSAFSASPPTTATVSAPTPTHSAIS 152
>TC12848 GB|AAA37921.1|553942|MUSIGF2RA insulin-like growth factor 2
receptor {Mus musculus;}, partial (23%)
Length = 664
Score = 25.8 bits (55), Expect = 3.8
Identities = 8/18 (44%), Positives = 16/18 (88%)
Frame = +3
Query: 43 FWSMFTGTVSLRWSGNLN 60
F S+F+G++S+RWS +++
Sbjct: 153 FQSLFSGSISVRWSSSVS 206
>TC14897 homologue to UP|RS18_ARATH (P34788) 40S ribosomal protein S18,
complete
Length = 669
Score = 25.0 bits (53), Expect = 6.4
Identities = 10/29 (34%), Positives = 17/29 (58%)
Frame = +2
Query: 62 LSSDLDTPIHDDLDVLEMEEREKVVRHMW 90
+S+ LD + DDL+ L+ + +RH W
Sbjct: 377 VSNALDMKLRDDLERLKKIRNHRGLRHYW 463
>TC14896 homologue to UP|RS18_ARATH (P34788) 40S ribosomal protein S18,
complete
Length = 906
Score = 25.0 bits (53), Expect = 6.4
Identities = 10/29 (34%), Positives = 17/29 (58%)
Frame = +3
Query: 62 LSSDLDTPIHDDLDVLEMEEREKVVRHMW 90
+S+ LD + DDL+ L+ + +RH W
Sbjct: 354 VSNALDMKLRDDLERLKKIRNHRGLRHYW 440
>BG662291
Length = 394
Score = 25.0 bits (53), Expect = 6.4
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = +3
Query: 7 LPTTTAAINKKENMDSGL 24
LPT +N+ EN+DSGL
Sbjct: 207 LPTYNEELNQNENIDSGL 260
>AU251380
Length = 372
Score = 24.6 bits (52), Expect = 8.4
Identities = 11/33 (33%), Positives = 16/33 (48%), Gaps = 5/33 (15%)
Frame = +1
Query: 26 GKGRYKFWALAAI-----LLLAFWSMFTGTVSL 53
G G + WA A L+ FW F+G ++L
Sbjct: 100 GNGGFVIWANEAFGPFWGSLMGFWKFFSGVINL 198
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.133 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,820,968
Number of Sequences: 28460
Number of extensions: 32689
Number of successful extensions: 165
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 165
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 165
length of query: 161
length of database: 4,897,600
effective HSP length: 83
effective length of query: 78
effective length of database: 2,535,420
effective search space: 197762760
effective search space used: 197762760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 51 (24.3 bits)
Medicago: description of AC147178.10


