

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147014.15 - phase: 0
(138 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
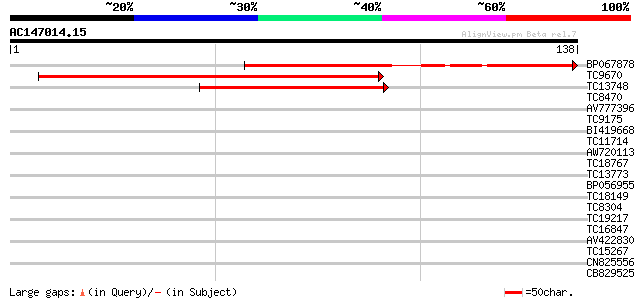
Score E
Sequences producing significant alignments: (bits) Value
BP067878 86 2e-18
TC9670 homologue to UP|O64958 (O64958) CUM1, partial (65%) 53 2e-08
TC13748 similar to UP|Q39401 (Q39401) MADS5 protein, partial (28%) 42 3e-05
TC8470 homologue to UP|Q8LK24 (Q8LK24) SOS2-like protein kinase,... 31 0.088
AV777396 31 0.088
TC9175 weakly similar to UP|INAD_DROME (Q24008) Inactivation-no-... 31 0.088
BI419668 30 0.15
TC11714 similar to UP|Q16861 (Q16861) Super cysteine rich protei... 30 0.15
AW720113 30 0.20
TC18767 30 0.20
TC13773 similar to UP|AAR20741 (AAR20741) At4g22560, partial (11%) 29 0.33
BP056955 28 0.44
TC18149 weakly similar to UP|Q91BL3 (Q91BL3) Essential structura... 28 0.44
TC8304 homologue to UP|LB41_ARATH (Q9M886) LOB domain protein 41... 28 0.74
TC19217 27 0.97
TC16847 weakly similar to PIR|T51733|T51733 cytidine deaminase ... 27 0.97
AV422830 27 1.3
TC15267 similar to UP|EXTN_TOBAC (P13983) Extensin precursor (Ce... 27 1.3
CN825556 27 1.3
CB829525 27 1.7
>BP067878
Length = 395
Score = 86.3 bits (212), Expect = 2e-18
Identities = 51/81 (62%), Positives = 61/81 (74%)
Frame = -3
Query: 58 VRARKNQAYKHQIDQLKEKEKNLVAENARLSKQVKRNYCPPQPQPQPTTKDHQREDQQPY 117
VR++K+Q Y +I+QLKEKEK L+AENARL++Q QPQP T + QRE+Q Y
Sbjct: 390 VRSKKSQVYHERIEQLKEKEKVLLAENARLTEQYGS-------QPQPAT-NAQRENQS-Y 238
Query: 118 AESSPSSDVVTELFIGLHRSS 138
AESSPSSDV TELFIGLHRSS
Sbjct: 237 AESSPSSDVETELFIGLHRSS 175
>TC9670 homologue to UP|O64958 (O64958) CUM1, partial (65%)
Length = 1060
Score = 52.8 bits (125), Expect = 2e-08
Identities = 27/84 (32%), Positives = 51/84 (60%)
Frame = +3
Query: 8 QNLKHETASLMKKIELLEASKRKLMGEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYK 67
Q + E L +I L+ + R++MGE LGS + EL+ +E +LEK +S +R++KN+
Sbjct: 240 QFYQQEADKLRVQISNLQNNNRQMMGESLGSMNAKELKNLETKLEKGISRIRSKKNELLF 419
Query: 68 HQIDQLKEKEKNLVAENARLSKQV 91
+I+ ++++E +L N L ++
Sbjct: 420 AEIEYMQKREIDLHNNNQLLRAKI 491
>TC13748 similar to UP|Q39401 (Q39401) MADS5 protein, partial (28%)
Length = 624
Score = 42.4 bits (98), Expect = 3e-05
Identities = 19/46 (41%), Positives = 33/46 (71%)
Frame = +1
Query: 47 IEQQLEKSVSVVRARKNQAYKHQIDQLKEKEKNLVAENARLSKQVK 92
+EQQL+ ++ +R+RKNQA I +L++K+K L +N L+K++K
Sbjct: 4 LEQQLDSALKHIRSRKNQALFESISELQKKDKALQEQNNLLTKKIK 141
>TC8470 homologue to UP|Q8LK24 (Q8LK24) SOS2-like protein kinase, partial
(29%)
Length = 807
Score = 30.8 bits (68), Expect = 0.088
Identities = 20/46 (43%), Positives = 23/46 (49%), Gaps = 1/46 (2%)
Frame = -3
Query: 94 NYCPPQPQPQPTT-KDHQREDQQPYAESSPSSDVVTELFIGLHRSS 138
++CP PQP TT H Q P + SSPS D T LH SS
Sbjct: 349 HHCPSSPQPPETTASPHTCPPQ*PASPSSPSPDPATSPC--LHSSS 218
>AV777396
Length = 577
Score = 30.8 bits (68), Expect = 0.088
Identities = 18/72 (25%), Positives = 40/72 (55%), Gaps = 4/72 (5%)
Frame = -2
Query: 50 QLEKSVSVVRARKNQAYKHQIDQLKEKEKNLVAEN----ARLSKQVKRNYCPPQPQPQPT 105
+LEK +S +R++KN+ +I+ ++++E +L N A++++ +R P T
Sbjct: 576 KLEKGISRIRSKKNEMLFAEIEYMQKREIDLHNSNQLLRAKIAESDERKNHNFNMLPGTT 397
Query: 106 TKDHQREDQQPY 117
+ ++ QQP+
Sbjct: 396 NFESLQQSQQPF 361
>TC9175 weakly similar to UP|INAD_DROME (Q24008)
Inactivation-no-after-potential D protein, partial (5%)
Length = 692
Score = 30.8 bits (68), Expect = 0.088
Identities = 14/34 (41%), Positives = 21/34 (61%)
Frame = +2
Query: 86 RLSKQVKRNYCPPQPQPQPTTKDHQREDQQPYAE 119
R KQ K PQP+P+P K+ +E+++P AE
Sbjct: 26 RTKKQAK---IVPQPEPEPEKKEEGKEEEKPAAE 118
>BI419668
Length = 647
Score = 30.0 bits (66), Expect = 0.15
Identities = 15/54 (27%), Positives = 24/54 (43%), Gaps = 2/54 (3%)
Frame = +3
Query: 75 EKEKNLVAENARLSKQVKRNYCPPQPQPQPTTKD--HQREDQQPYAESSPSSDV 126
+++K E L Q + + PP P P P K+ + RE Q P + D+
Sbjct: 240 KEDKPTKVEGGGLKNQTESEFLPPPPPPPPIKKNPLNSREKQGPAVARTDDDDI 401
>TC11714 similar to UP|Q16861 (Q16861) Super cysteine rich protein
(Fragment), partial (48%)
Length = 519
Score = 30.0 bits (66), Expect = 0.15
Identities = 13/34 (38%), Positives = 22/34 (64%)
Frame = -1
Query: 61 RKNQAYKHQIDQLKEKEKNLVAENARLSKQVKRN 94
+K Q + Q+ QLK++++ L + R SKQV R+
Sbjct: 138 KKQQQQQQQLQQLKQQQQQLQQQKTRNSKQVPRH 37
>AW720113
Length = 512
Score = 29.6 bits (65), Expect = 0.20
Identities = 15/36 (41%), Positives = 18/36 (49%), Gaps = 5/36 (13%)
Frame = +1
Query: 97 PPQPQPQ-----PTTKDHQREDQQPYAESSPSSDVV 127
PPQPQPQ P + H+ E P+ PS VV
Sbjct: 133 PPQPQPQPPPPPPPPQFHENEG*NPFIHDQPSVGVV 240
>TC18767
Length = 1004
Score = 29.6 bits (65), Expect = 0.20
Identities = 25/103 (24%), Positives = 39/103 (37%), Gaps = 1/103 (0%)
Frame = +2
Query: 33 GEGLGSCSLDELQQIEQQLEKSVSVVRARKNQAYKHQIDQLKEKEKNLVAENARLSKQVK 92
G LG S D ++ LE R +Y H + + N+ +AR ++ +
Sbjct: 86 GYALGLTSADGSSNLDGGLEIIEDASRCFNCGSYNHSLRECSRPRDNVAVNSARKQRKSR 265
Query: 93 RNYCPPQPQPQPTTKDHQREDQQPYAESSPSS-DVVTELFIGL 134
RN P T+ +Q Y P + D VT +GL
Sbjct: 266 RNQNVSSRNP---TRYYQNSPAGKYDGLRPGALDNVTRRLLGL 385
>TC13773 similar to UP|AAR20741 (AAR20741) At4g22560, partial (11%)
Length = 441
Score = 28.9 bits (63), Expect = 0.33
Identities = 10/20 (50%), Positives = 12/20 (60%)
Frame = +2
Query: 97 PPQPQPQPTTKDHQREDQQP 116
PP P P P+ K+H R Q P
Sbjct: 278 PPSPSPNPSPKNHSRNTQVP 337
>BP056955
Length = 581
Score = 28.5 bits (62), Expect = 0.44
Identities = 27/99 (27%), Positives = 45/99 (45%), Gaps = 14/99 (14%)
Frame = +2
Query: 54 SVSVVRARKNQAYKHQIDQLKEKEK----NLVAENARL--------SKQVKRNYCPPQPQ 101
S+S R +K + K + +++E+ NL++ N L S R+ PQPQ
Sbjct: 23 SLSRERGKKKKKKKKKKKKMEEQALIGALNLISRNLPLPPDLFTTVSSIYHRSQPQPQPQ 202
Query: 102 PQPTTKDHQREDQQPYAESSPSSDVVTELFIGL--HRSS 138
PQP + + QP + D++ +L L HR+S
Sbjct: 203 PQPQPQPQPQSQPQPQQQ-----DLLADLQDALSNHRAS 304
>TC18149 weakly similar to UP|Q91BL3 (Q91BL3) Essential structural protein
pp78/81, partial (5%)
Length = 710
Score = 28.5 bits (62), Expect = 0.44
Identities = 11/24 (45%), Positives = 14/24 (57%)
Frame = +3
Query: 98 PQPQPQPTTKDHQREDQQPYAESS 121
PQP PQP HQ+ Q P+ + S
Sbjct: 294 PQPHPQPGAPPHQQYQQTPHPQYS 365
>TC8304 homologue to UP|LB41_ARATH (Q9M886) LOB domain protein 41, partial
(8%)
Length = 617
Score = 27.7 bits (60), Expect = 0.74
Identities = 19/56 (33%), Positives = 26/56 (45%), Gaps = 6/56 (10%)
Frame = -1
Query: 74 KEKEKN-----LVAENARLSKQVKRNYCPPQPQPQPTTKDH-QREDQQPYAESSPS 123
K K++N L +R ++ RN P P P+ K H Q Q P A+SS S
Sbjct: 359 KRKQQNGF*SLLHITESRFKSKLNRNIKPVIEAPAPSCKSHSQPSHQNPSAQSSSS 192
>TC19217
Length = 637
Score = 27.3 bits (59), Expect = 0.97
Identities = 25/101 (24%), Positives = 45/101 (43%), Gaps = 3/101 (2%)
Frame = +2
Query: 41 LDELQQIEQQLEKSVSVVRARKNQAYKHQIDQLKEKEKNLVAENARLSKQVKRNYCPPQP 100
+DE+++++ + V +ARK KHQ D + + ++ A L+++ K+
Sbjct: 347 VDEIKKLDSSKD---GVPKARK---VKHQKDHIGMRSLEMLTVRASLAEKWKKKVPSTDT 508
Query: 101 QPQPTT-KDHQREDQQPYAESSPSSDVV--TELFIGLHRSS 138
P P+T R D + S D V L GL +S+
Sbjct: 509 TPVPSTGLKRSRPDNSQHGTSVFDDDAVGPARLSQGLRKSN 631
>TC16847 weakly similar to PIR|T51733|T51733 cytidine deaminase [validated]
- Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (74%)
Length = 1098
Score = 27.3 bits (59), Expect = 0.97
Identities = 16/53 (30%), Positives = 21/53 (39%), Gaps = 2/53 (3%)
Frame = -1
Query: 65 AYKHQIDQLKEKEKNLVAENARLSKQVKRNYCP--PQPQPQPTTKDHQREDQQ 115
++K Q +QLK K K R + P P P P P T+ Q Q
Sbjct: 948 SHKFQFNQLKYKGKP*TGTEGRFLAAATLQWSPV*PPPHPSPPTRRRQHRHNQ 790
>AV422830
Length = 440
Score = 26.9 bits (58), Expect = 1.3
Identities = 12/26 (46%), Positives = 13/26 (49%)
Frame = +3
Query: 97 PPQPQPQPTTKDHQREDQQPYAESSP 122
PP P P PTT H+ P SSP
Sbjct: 159 PPPPLPPPTTTTHRLLPHIPSLSSSP 236
>TC15267 similar to UP|EXTN_TOBAC (P13983) Extensin precursor (Cell wall
hydroxyproline-rich glycoprotein), partial (6%)
Length = 1003
Score = 26.9 bits (58), Expect = 1.3
Identities = 13/29 (44%), Positives = 16/29 (54%)
Frame = -2
Query: 97 PPQPQPQPTTKDHQREDQQPYAESSPSSD 125
PPQ QP PT HQ + P SP+S+
Sbjct: 654 PPQTQPPPT---HQGKSADPSPPRSPTSE 577
>CN825556
Length = 655
Score = 26.9 bits (58), Expect = 1.3
Identities = 11/30 (36%), Positives = 14/30 (46%)
Frame = +1
Query: 98 PQPQPQPTTKDHQREDQQPYAESSPSSDVV 127
PQPQPQP + + QP + P V
Sbjct: 469 PQPQPQPQPQPQPQPQPQPQPQPQPQPQPV 558
Score = 26.6 bits (57), Expect = 1.7
Identities = 10/25 (40%), Positives = 13/25 (52%)
Frame = +1
Query: 98 PQPQPQPTTKDHQREDQQPYAESSP 122
PQPQPQP + + QP + P
Sbjct: 463 PQPQPQPQPQPQPQPQPQPQPQPQP 537
Score = 26.6 bits (57), Expect = 1.7
Identities = 11/27 (40%), Positives = 14/27 (51%)
Frame = +1
Query: 98 PQPQPQPTTKDHQREDQQPYAESSPSS 124
PQPQPQP + + QP P+S
Sbjct: 493 PQPQPQPQPQPQPQPQPQPQPVPPPAS 573
Score = 25.8 bits (55), Expect = 2.8
Identities = 10/25 (40%), Positives = 13/25 (52%)
Frame = +1
Query: 98 PQPQPQPTTKDHQREDQQPYAESSP 122
PQPQPQP + + QP + P
Sbjct: 487 PQPQPQPQPQPQPQPQPQPQPQPVP 561
>CB829525
Length = 432
Score = 26.6 bits (57), Expect = 1.7
Identities = 17/50 (34%), Positives = 23/50 (46%), Gaps = 2/50 (4%)
Frame = +2
Query: 89 KQVKRNYCPPQPQPQPTTKDHQREDQQPYAESSPSSDVVTELF--IGLHR 136
K NYC QP P P+ E P A +S + T L+ +G+HR
Sbjct: 140 KNSSANYCSTQPNPSPSAS----EPSSPSA-TSKVPPLATPLYSLLGIHR 274
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.310 0.126 0.343
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,331,362
Number of Sequences: 28460
Number of extensions: 33365
Number of successful extensions: 366
Number of sequences better than 10.0: 69
Number of HSP's better than 10.0 without gapping: 329
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 354
length of query: 138
length of database: 4,897,600
effective HSP length: 81
effective length of query: 57
effective length of database: 2,592,340
effective search space: 147763380
effective search space used: 147763380
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 50 (23.9 bits)
Medicago: description of AC147014.15


