

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147013.10 - phase: 0 /pseudo
(339 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
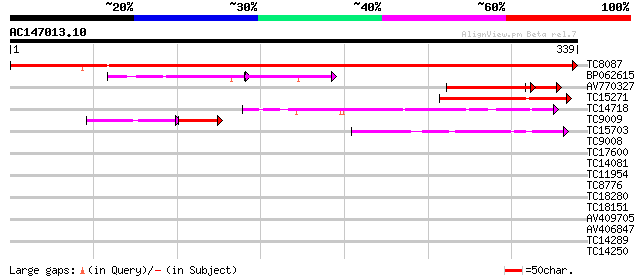
Score E
Sequences producing significant alignments: (bits) Value
TC8087 homologue to UP|FENR_PEA (P10933) Ferredoxin--NADP reduct... 597 e-172
BP062615 62 1e-19
AV770327 66 2e-15
TC15271 similar to PIR|T06773|T06773 ferredoxin-NADP reductase ... 70 7e-13
TC14718 similar to UP|NCPR_CATRO (Q05001) NADPH--cytochrome P450... 60 4e-10
TC9009 similar to PIR|T06773|T06773 ferredoxin-NADP reductase -... 38 3e-05
TC15703 weakly similar to UP|Q7VSP1 (Q7VSP1) CDP-6-deoxy-delta-3... 40 7e-04
TC9008 homologue to PIR|T06773|T06773 ferredoxin-NADP reductase ... 38 0.003
TC17600 similar to PIR|A47298|A47298 NADPH-ferrihemoprotein redu... 35 0.018
TC14081 similar to UP|RCA_PHAAU (O98997) Ribulose bisphosphate c... 32 0.12
TC11954 homologue to GB|AAP68290.1|31711868|BT008851 At3g12156 {... 28 2.9
TC8776 27 3.8
TC18280 27 3.8
TC18151 weakly similar to UP|Q9PYU9 (Q9PYU9) ORF94, partial (40%) 27 6.4
AV409705 27 6.4
AV406847 26 8.4
TC14289 similar to UP|LEU1_SOYBN (Q39891) Probable 2-isopropylma... 26 8.4
TC14250 homologue to UP|Q8GTQ9 (Q8GTQ9) Succinyl-CoA ligase alph... 26 8.4
>TC8087 homologue to UP|FENR_PEA (P10933) Ferredoxin--NADP reductase, leaf
isozyme, chloroplast precursor (FNR) , partial (94%)
Length = 1449
Score = 597 bits (1540), Expect = e-172
Identities = 294/342 (85%), Positives = 316/342 (91%), Gaps = 3/342 (0%)
Frame = +1
Query: 1 MAAAVTAAVSFPYSNSTSLPIRTSVISPDRLVFKKVSLNNVS---ISGRLTVRAEVATEA 57
MAAAVTAAVSFP + STSL RTS+++P+R+VFKKV L S +S R V + T+A
Sbjct: 136 MAAAVTAAVSFPSTKSTSLTSRTSIVAPERVVFKKVPLQYTSGRVVSIRAQVTTDTDTQA 315
Query: 58 PAPVKVEKISKKQEEGIVVNKFKPKTPYIGRCLLNTKITGDDAPGETWHMVFSTEGEVPY 117
P P KVEK+ KKQEEG+VVNKFKPK PYIGR LLN+KITGDDAPGETWHMVFSTEGEVPY
Sbjct: 316 P-PAKVEKVHKKQEEGVVVNKFKPKNPYIGRVLLNSKITGDDAPGETWHMVFSTEGEVPY 492
Query: 118 REGQSIGIVPDGIDKNGKPHKLRLYSIASSALGDFGDSKTVSLCVKRLVYTNDAGEVVKG 177
REGQSIG++P+GIDKNGKPHKLRLYSIASSALGDFGDSKTVSLCVKRLVYTND+GE+VKG
Sbjct: 493 REGQSIGVIPEGIDKNGKPHKLRLYSIASSALGDFGDSKTVSLCVKRLVYTNDSGEIVKG 672
Query: 178 VCSNFLCDLRPGSEVQITGPVGKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHE 237
VCSNFLCDL+PGSEV ITGPVGKEMLMPKDPNATVIML TGTGIAPFRSFLWKMFFEKH+
Sbjct: 673 VCSNFLCDLKPGSEVTITGPVGKEMLMPKDPNATVIMLATGTGIAPFRSFLWKMFFEKHD 852
Query: 238 DYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTR 297
DYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEK+PENFRLDFAVSREQ NDKGEKMYIQTR
Sbjct: 853 DYKFNGLAWLFLGVPTSSSLLYKEEFEKMKEKSPENFRLDFAVSREQTNDKGEKMYIQTR 1032
Query: 298 MAQYAEELWELLKKDNTFVYMCGLKGMEKGIDDIMVSLAAKD 339
MAQYAEELW+LLKKDNTFVYMCGLKGMEKGIDDIMVSLAA+D
Sbjct: 1033MAQYAEELWQLLKKDNTFVYMCGLKGMEKGIDDIMVSLAARD 1158
>BP062615
Length = 496
Score = 61.6 bits (148), Expect(2) = 1e-19
Identities = 37/88 (42%), Positives = 50/88 (56%), Gaps = 2/88 (2%)
Frame = -3
Query: 59 APVKVEKISKKQEEGIVVNKFKPKTPYIGRCLLNTKITGDDAPGETWHMVFSTEGEVPYR 118
+P+++E S+ +N KPK PY + +I G APGET H+V G VPY
Sbjct: 488 SPLELEAASEPP-----LNLHKPKQPYTTIVSVE-RIVGPKAPGETCHIVIDHGGNVPYW 327
Query: 119 EGQSIGIVPDGID--KNGKPHKLRLYSI 144
EGQS G +P G + K G P+ +RLYSI
Sbjct: 326 EGQSYGFIPPGENPKKPGSPYNVRLYSI 243
Score = 50.8 bits (120), Expect(2) = 1e-19
Identities = 30/59 (50%), Positives = 34/59 (56%), Gaps = 4/59 (6%)
Frame = -2
Query: 141 LYSIASSALGDFGDSKTVSLCVKRLVYTNDA----GEVVKGVCSNFLCDLRPGSEVQIT 195
LYSIAS+ D D K SLCV R VY + GVCSNFLC+ +PG VQIT
Sbjct: 240 LYSIASTRYRDSFDGKIASLCVCRAVYYDPET**EDPSKNGVCSNFLCNSKPGD*VQIT 64
>AV770327
Length = 329
Score = 65.9 bits (159), Expect(2) = 2e-15
Identities = 35/53 (66%), Positives = 39/53 (73%)
Frame = -3
Query: 262 EFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLKKDNT 314
E E MKE+AP +FRLD AVSRE +D GEK YI T M +YAEEL LLK DNT
Sbjct: 327 EVE*MKERAPAHFRLDLAVSREPTHDIGEKKYIPTPMGRYAEELGLLLK*DNT 169
Score = 32.7 bits (73), Expect(2) = 2e-15
Identities = 15/22 (68%), Positives = 15/22 (68%)
Frame = -1
Query: 309 LKKDNTFVYMCGLKGMEKGIDD 330
L K FVYMCGL GM KG DD
Sbjct: 185 LSKTTLFVYMCGL*GMGKGNDD 120
>TC15271 similar to PIR|T06773|T06773 ferredoxin-NADP reductase - garden
pea (fragment) {Pisum sativum;} ,
partial (27%)
Length = 729
Score = 69.7 bits (169), Expect = 7e-13
Identities = 32/79 (40%), Positives = 52/79 (65%)
Frame = +2
Query: 258 LYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLKKDNTFVY 317
LY +EF K + P +FR D A+SREQ N G +MY+Q ++ +Y++E+++LL + +Y
Sbjct: 2 LYHDEFTKYLKDYPTHFRYDIALSREQKNQNGGQMYVQDKIEEYSDEIFKLL-DNGAHIY 178
Query: 318 MCGLKGMEKGIDDIMVSLA 336
CGL+GM GI + + +A
Sbjct: 179 FCGLRGMMPGIQETLKRVA 235
>TC14718 similar to UP|NCPR_CATRO (Q05001) NADPH--cytochrome P450 reductase
(CPR) (P450R) , partial (36%)
Length = 1163
Score = 60.5 bits (145), Expect = 4e-10
Identities = 58/199 (29%), Positives = 90/199 (45%), Gaps = 10/199 (5%)
Frame = +1
Query: 140 RLYSIASSALGDFGDSKTVSLCVKRLVYTND---AGEVVKGVCSNFLCDLRPGSEVQITG 196
R YSI+SS S+ C ND G + +GVCS ++ + P + Q
Sbjct: 109 RFYSISSSPR--MAPSRIHVTCA----LVNDKMPTGRIHRGVCSTWMKNSVPLEKSQDCS 270
Query: 197 --PV---GKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGV 251
P+ +P D +IM+G GTG+APFR FL + K ED G + LF G
Sbjct: 271 WAPIFVRQSNFKLPADNKVPIIMIGPGTGLAPFRGFLQERLALK-EDGAELGPSVLFFGC 447
Query: 252 PT-SSSLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLK 310
+Y++E + L A SRE K Y+Q +M + A ++W ++
Sbjct: 448 RNRQMDYIYEDELNHFVNSGALS-ELIVAFSREGPT----KEYVQHKMMEKASDIWNMIS 612
Query: 311 KDNTFVYMCG-LKGMEKGI 328
+ ++Y+CG KGM + +
Sbjct: 613 Q-GAYIYVCGDAKGMARDV 666
>TC9009 similar to PIR|T06773|T06773 ferredoxin-NADP reductase - garden
pea (fragment) {Pisum sativum;} ,
partial (33%)
Length = 585
Score = 37.7 bits (86), Expect(2) = 3e-05
Identities = 15/26 (57%), Positives = 19/26 (72%)
Frame = +1
Query: 102 GETWHMVFSTEGEVPYREGQSIGIVP 127
GET H+V +G VPY EGQS G++P
Sbjct: 505 GETCHIVIDHDGNVPYWEGQSYGVIP 582
Score = 25.8 bits (55), Expect(2) = 3e-05
Identities = 17/56 (30%), Positives = 25/56 (44%)
Frame = +3
Query: 47 LTVRAEVATEAPAPVKVEKISKKQEEGIVVNKFKPKTPYIGRCLLNTKITGDDAPG 102
L+V + AT V ++ + E + N KPK PY + +I G APG
Sbjct: 345 LSVPVQQATGPKVTVSPLELEEAPEPPL--NLHKPKQPYTATIVSVERIVGPKAPG 506
>TC15703 weakly similar to UP|Q7VSP1 (Q7VSP1)
CDP-6-deoxy-delta-3,4-glucoseen reductase , partial
(6%)
Length = 560
Score = 39.7 bits (91), Expect = 7e-04
Identities = 33/131 (25%), Positives = 63/131 (47%), Gaps = 1/131 (0%)
Frame = +3
Query: 205 PKDPNATVIMLGTGTGIAPFRSFLWKMF-FEKHEDYKFNGLAWLFLGVPTSSSLLYKEEF 263
P + TVI+ TG+GI+P RS + F +K D + ++ G + Y+++F
Sbjct: 9 PPEKFGTVIVFATGSGISPIRSLIESGFSADKRSDVR------VYYGARNLQRMAYQDKF 170
Query: 264 EKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLKKDNTFVYMCGLKG 323
K+ ++ +S+ + + GE Y+Q + A+E+ L +T +CG K
Sbjct: 171 ---KQWESSGVKIVPVLSQPEDSWTGESGYVQAAFTR-AKEISNPL---STGAVLCGQKQ 329
Query: 324 MEKGIDDIMVS 334
M + + I+V+
Sbjct: 330 MTEEVTSILVA 362
>TC9008 homologue to PIR|T06773|T06773 ferredoxin-NADP reductase - garden
pea (fragment) {Pisum sativum;} ,
partial (18%)
Length = 737
Score = 37.7 bits (86), Expect = 0.003
Identities = 16/27 (59%), Positives = 18/27 (66%)
Frame = +2
Query: 101 PGETWHMVFSTEGEVPYREGQSIGIVP 127
PGET H+V G VPY EGQS G +P
Sbjct: 182 PGETCHIVIDHGGNVPYWEGQSYGFIP 262
>TC17600 similar to PIR|A47298|A47298 NADPH-ferrihemoprotein reductase -
mung bean {Vigna radiata;} , partial (19%)
Length = 717
Score = 35.0 bits (79), Expect = 0.018
Identities = 31/98 (31%), Positives = 45/98 (45%), Gaps = 2/98 (2%)
Frame = +3
Query: 233 FEKHEDYKFNGLAWLFLGVPTSS-SLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEK 291
F ED G A LF G +Y++E E+ + L A SRE EK
Sbjct: 12 FALKEDGIELGPAILFFGCRNRRMDFIYEDELNNFLEQGSLS-ELIVAFSREG----SEK 176
Query: 292 MYIQTRMAQYAEELWELLKKDNTFVYMCG-LKGMEKGI 328
Y+Q +M A LW L+ + ++Y+CG KGM + +
Sbjct: 177 EYVQHKMMDKAAYLWSLISQ-GAYLYVCGDAKGMARDV 287
>TC14081 similar to UP|RCA_PHAAU (O98997) Ribulose bisphosphate
carboxylase/oxygenase activase, chloroplast precursor
(RuBisCO activase) (RA), partial (98%)
Length = 1625
Score = 32.3 bits (72), Expect = 0.12
Identities = 25/97 (25%), Positives = 42/97 (42%), Gaps = 10/97 (10%)
Frame = -3
Query: 1 MAAAVTAAVSFPYSNSTSLPIRTSVISPDRLVFKKVSLNNVSISGRLTVRAEVATEAPAP 60
++ +V S + LP+ T V+ + F K+ L +AE+ TEAPAP
Sbjct: 270 LSPSVCLLSSISAATILKLPLETLVLGSLEITFFKL----------LPKKAELGTEAPAP 121
Query: 61 ----------VKVEKISKKQEEGIVVNKFKPKTPYIG 87
V + + +EG+VV + + K Y+G
Sbjct: 120 EPFKLNGVLTVPIVETEAAIDEGVVVQRTERKREYLG 10
>TC11954 homologue to GB|AAP68290.1|31711868|BT008851 At3g12156 {Arabidopsis
thaliana;}, partial (26%)
Length = 512
Score = 27.7 bits (60), Expect = 2.9
Identities = 18/62 (29%), Positives = 30/62 (48%)
Frame = +1
Query: 204 MPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPTSSSLLYKEEF 263
+PK+PNA + + T G P S + E + + + + W+ G SS LL+ EF
Sbjct: 82 IPKNPNAVIFVAATDDGYIPKHSVM-----ELQKAWPGSEVRWV-TGGHVSSFLLHNGEF 243
Query: 264 EK 265
+
Sbjct: 244 RR 249
>TC8776
Length = 611
Score = 27.3 bits (59), Expect = 3.8
Identities = 19/63 (30%), Positives = 30/63 (47%)
Frame = -3
Query: 32 VFKKVSLNNVSISGRLTVRAEVATEAPAPVKVEKISKKQEEGIVVNKFKPKTPYIGRCLL 91
VF SL+ VSI L +R P + +KK + ++ F + P++ C+L
Sbjct: 249 VFGSTSLSLVSI---LKLRRYSQKSLPNFKSYQTCTKKFGKSLMTLSFHVQLPHLVTCML 79
Query: 92 NTK 94
NTK
Sbjct: 78 NTK 70
>TC18280
Length = 572
Score = 27.3 bits (59), Expect = 3.8
Identities = 19/50 (38%), Positives = 28/50 (56%)
Frame = -1
Query: 29 DRLVFKKVSLNNVSISGRLTVRAEVATEAPAPVKVEKISKKQEEGIVVNK 78
DR + S N + + + V+ E TE A VKVEKI+K+++ VV K
Sbjct: 182 DRNGTPRSSSENTNKARNVDVKIE-KTERAASVKVEKIAKEEKPVSVVKK 36
>TC18151 weakly similar to UP|Q9PYU9 (Q9PYU9) ORF94, partial (40%)
Length = 540
Score = 26.6 bits (57), Expect = 6.4
Identities = 11/21 (52%), Positives = 14/21 (66%)
Frame = +2
Query: 144 IASSALGDFGDSKTVSLCVKR 164
+ SS GDFGD++ L VKR
Sbjct: 65 VLSSVAGDFGDNRNPKLLVKR 127
>AV409705
Length = 263
Score = 26.6 bits (57), Expect = 6.4
Identities = 15/47 (31%), Positives = 25/47 (52%)
Frame = +2
Query: 12 PYSNSTSLPIRTSVISPDRLVFKKVSLNNVSISGRLTVRAEVATEAP 58
P+S +SLP S++ P RL F++ +N G +T + + AP
Sbjct: 53 PFSPLSSLPSNPSLLPPPRLTFRRRIASNF---GPMTTTSRLRHLAP 184
>AV406847
Length = 415
Score = 26.2 bits (56), Expect = 8.4
Identities = 12/28 (42%), Positives = 14/28 (49%)
Frame = -3
Query: 181 NFLCDLRPGSEVQITGPVGKEMLMPKDP 208
+FLC RP S T P + L P DP
Sbjct: 203 SFLCPTRPKSSPSSTVPPPRSHLRPSDP 120
>TC14289 similar to UP|LEU1_SOYBN (Q39891) Probable 2-isopropylmalate
synthase (Alpha-isopropylmalate synthase) (Alpha-IPM
synthetase) (Late nodulin 56) (N-56) , partial (85%)
Length = 1822
Score = 26.2 bits (56), Expect = 8.4
Identities = 9/23 (39%), Positives = 16/23 (69%)
Frame = -1
Query: 4 AVTAAVSFPYSNSTSLPIRTSVI 26
A +A+SFPYSN ++P + ++
Sbjct: 709 AFISAISFPYSNGIAMPTVSGIV 641
>TC14250 homologue to UP|Q8GTQ9 (Q8GTQ9) Succinyl-CoA ligase alpha subunit
, partial (89%)
Length = 1395
Score = 26.2 bits (56), Expect = 8.4
Identities = 13/30 (43%), Positives = 17/30 (56%), Gaps = 1/30 (3%)
Frame = -2
Query: 79 FKPKTPY-IGRCLLNTKITGDDAPGETWHM 107
F P P+ I R LL+T GD+A + W M
Sbjct: 203 FFPVIPWQITRVLLSTNTAGDEAVADRWRM 114
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.135 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,342,089
Number of Sequences: 28460
Number of extensions: 67739
Number of successful extensions: 313
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 307
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 307
length of query: 339
length of database: 4,897,600
effective HSP length: 91
effective length of query: 248
effective length of database: 2,307,740
effective search space: 572319520
effective search space used: 572319520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Medicago: description of AC147013.10


