

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147011.7 + phase: 0 /pseudo
(384 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
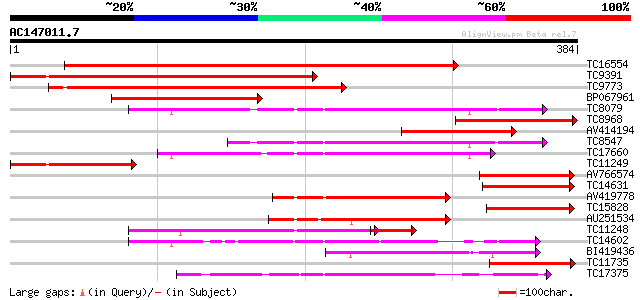
Score E
Sequences producing significant alignments: (bits) Value
TC16554 homologue to UP|AAN71719 (AAN71719) Glycogen synthase ki... 455 e-129
TC9391 homologue to UP|MSK1_MEDSA (P51137) Glycogen synthase kin... 377 e-105
TC9773 homologue to UP|KSGI_ARATH (Q39012) Shaggy-related protei... 324 1e-89
BP067961 198 1e-51
TC8079 homologue to UP|Q9M6S1 (Q9M6S1) MAP kinase 3, complete 178 1e-45
TC8968 homologue to UP|Q41619 (Q41619) Protein kinase (Fragment)... 166 5e-42
AV414194 154 3e-38
TC8547 homologue to UP|MMK1_MEDSA (Q07176) Mitogen-activated pro... 153 5e-38
TC17660 homologue to UP|Q9M6R8 (Q9M6R8) MAP kinase PsMAPK2, part... 135 1e-32
TC11249 similar to GB|CAA48472.1|313148|AMMSK3A protein kinase {... 119 6e-28
AV766574 115 1e-26
TC14631 homologue to GB|CAA48472.1|313148|AMMSK3A protein kinase... 110 4e-25
AV419778 105 2e-23
TC15828 similar to UP|P93774 (P93774) Shaggy-like kinase (Fragme... 101 2e-22
AU251534 100 7e-22
TC11248 homologue to UP|P93323 (P93323) Cdc2MsF protein, partial... 79 1e-21
TC14602 UP|Q8GS56 (Q8GS56) Phosphoenolpyruvate carboxylase kinas... 95 2e-20
BI419436 95 2e-20
TC11735 similar to GB|CAA68872.1|1504063|ATASKKAP shaggy-like ki... 94 4e-20
TC17375 mitogen-activated kinase kinase kinase alpha [Lotus corn... 92 1e-19
>TC16554 homologue to UP|AAN71719 (AAN71719) Glycogen synthase kinase 3 beta
protein kinase DWARF12 , partial (71%)
Length = 927
Score = 455 bits (1171), Expect = e-129
Identities = 214/267 (80%), Positives = 245/267 (91%)
Frame = +1
Query: 38 DDREMEATVVDGNGTETGHIIVTTIGGRNGQPKQTISYMAERVVGHGSFGVVFQAKCLET 97
+D+EM +V++GN + TGHII TTIGG+NG+PKQTISYMAERVVG GSFG+VFQAKCLE+
Sbjct: 127 EDKEMSTSVINGNDSLTGHIISTTIGGKNGEPKQTISYMAERVVGTGSFGIVFQAKCLES 306
Query: 98 GETVAIKKVLQDKRYKNRELQTMRLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPETV 157
GE VAIKKVLQD+RYKNRELQ MR++DHPNVVSLKHCFFSTT DEL+LNLV+EYVPE++
Sbjct: 307 GEAVAIKKVLQDRRYKNRELQLMRVMDHPNVVSLKHCFFSTTSTDELFLNLVMEYVPESM 486
Query: 158 HRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHRCIGVCHRDIKPQNLLVNPHTHQVKL 217
+RVIKHYS NQRMP+IYVKLY YQIFR L+YIH GVCHRD+KPQN+LV+P THQVKL
Sbjct: 487 YRVIKHYSNANQRMPIIYVKLYMYQIFRGLAYIHTVPGVCHRDLKPQNILVDPLTHQVKL 666
Query: 218 CDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTTAIDVWSVGCVLAELLLGQPLFP 277
CDFGSAK+LVKGE NISYICSR+YRAPELIFGATEY+T+ID+WS GCVLAELLLGQPLFP
Sbjct: 667 CDFGSAKMLVKGEANISYICSRFYRAPELIFGATEYSTSIDIWSAGCVLAELLLGQPLFP 846
Query: 278 GESGVDQLVEIIKVLGTPTREEIKCMN 304
GE+ VDQLV IIKVLGTPTREE++CMN
Sbjct: 847 GENAVDQLVHIIKVLGTPTREEVRCMN 927
>TC9391 homologue to UP|MSK1_MEDSA (P51137) Glycogen synthase kinase-3
homolog MsK-1 , partial (48%)
Length = 841
Score = 377 bits (967), Expect = e-105
Identities = 191/210 (90%), Positives = 199/210 (93%), Gaps = 2/210 (0%)
Frame = +1
Query: 1 MASVGVAPTSGFKESLG-DGEIGVDDILPEEMSDMKIRDDREMEATVVDGNG-TETGHII 58
MASVGVAPTSG +E+ G G GVD LPEEM+DMKIRDDREMEATVVDGN TETGHII
Sbjct: 214 MASVGVAPTSGVREASGCHGAAGVDR-LPEEMNDMKIRDDREMEATVVDGNDVTETGHII 390
Query: 59 VTTIGGRNGQPKQTISYMAERVVGHGSFGVVFQAKCLETGETVAIKKVLQDKRYKNRELQ 118
VTTIGGRNGQPKQTISYMAER+VGHGSFGVVFQAKCLETGETVAIKKVLQDKRYKNRELQ
Sbjct: 391 VTTIGGRNGQPKQTISYMAERIVGHGSFGVVFQAKCLETGETVAIKKVLQDKRYKNRELQ 570
Query: 119 TMRLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKL 178
TMRLLDHPNVV+LKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHY+KLNQR+P+IYVKL
Sbjct: 571 TMRLLDHPNVVALKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHYNKLNQRLPLIYVKL 750
Query: 179 YTYQIFRALSYIHRCIGVCHRDIKPQNLLV 208
YTYQIFRALSYIHRCI VCHRDIKPQNLLV
Sbjct: 751 YTYQIFRALSYIHRCIRVCHRDIKPQNLLV 840
>TC9773 homologue to UP|KSGI_ARATH (Q39012) Shaggy-related protein kinase
iota (ASK-iota) , partial (47%)
Length = 774
Score = 324 bits (831), Expect = 1e-89
Identities = 155/202 (76%), Positives = 178/202 (87%)
Frame = +1
Query: 27 LPEEMSDMKIRDDREMEATVVDGNGTETGHIIVTTIGGRNGQPKQTISYMAERVVGHGSF 86
+P SDM+ D+ M TV++GN TGHII TTIGG+NG+PKQTISYMAERVVG GSF
Sbjct: 175 VPRRSSDMET--DKAMSTTVIEGNDAVTGHIISTTIGGKNGEPKQTISYMAERVVGTGSF 348
Query: 87 GVVFQAKCLETGETVAIKKVLQDKRYKNRELQTMRLLDHPNVVSLKHCFFSTTEKDELYL 146
GVVFQAKCLETGE VAIKKVLQD+RYKNRELQ MRL+DHPNV+SLKHCFFSTT +DEL+L
Sbjct: 349 GVVFQAKCLETGEAVAIKKVLQDRRYKNRELQLMRLMDHPNVISLKHCFFSTTSRDELFL 528
Query: 147 NLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHRCIGVCHRDIKPQNL 206
NLV++YVPET++RV+KHY+ +NQRMP+IYVKLYTYQIFR L+YIH GV HRD+KPQNL
Sbjct: 529 NLVMDYVPETLYRVLKHYNSMNQRMPLIYVKLYTYQIFRGLAYIHTVPGVSHRDVKPQNL 708
Query: 207 LVNPHTHQVKLCDFGSAKVLVK 228
LV+P THQVKLCDFGSAKVLVK
Sbjct: 709 LVHPLTHQVKLCDFGSAKVLVK 774
>BP067961
Length = 368
Score = 198 bits (503), Expect = 1e-51
Identities = 95/102 (93%), Positives = 101/102 (98%)
Frame = +3
Query: 70 KQTISYMAERVVGHGSFGVVFQAKCLETGETVAIKKVLQDKRYKNRELQTMRLLDHPNVV 129
++TISYMAER+VGHGS GVVFQAKCLETGETVAIKKVLQDKRYKNRELQTMRLLDHPNVV
Sbjct: 63 RRTISYMAERMVGHGSLGVVFQAKCLETGETVAIKKVLQDKRYKNRELQTMRLLDHPNVV 242
Query: 130 SLKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRM 171
+LKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHY+KLNQR+
Sbjct: 243 ALKHCFFSTTEKDELYLNLVLEYVPETVHRVIKHYNKLNQRL 368
>TC8079 homologue to UP|Q9M6S1 (Q9M6S1) MAP kinase 3, complete
Length = 1550
Score = 178 bits (452), Expect = 1e-45
Identities = 101/293 (34%), Positives = 165/293 (55%), Gaps = 9/293 (3%)
Frame = +3
Query: 81 VGHGSFGVVFQAKCLETGETVAIKKVLQ------DKRYKNRELQTMRLLDHPNVVSLKHC 134
+G G++G+V ET E VA+KK+ D + RE++ +R LDH NV++L+
Sbjct: 309 IGRGAYGIVCSLLNTETNELVAVKKIANAFDNHMDAKRTLREIKLLRHLDHENVIALRDV 488
Query: 135 FFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHRCI 194
++ + + E + +H++I+ NQ + + + + YQ+ R L YIH
Sbjct: 489 IPPPLRREFTDVYITTELMDTDLHQIIRS----NQGLSEEHCQYFLYQVLRGLKYIHSA- 653
Query: 195 GVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYT 254
+ HRD+KP NLL+N + +K+ DFG A+ V+ + Y+ +R+YRAPEL+ +++YT
Sbjct: 654 NIIHRDLKPSNLLLNSNC-DLKIIDFGLARPTVENDFMTEYVVTRWYRAPELLLNSSDYT 830
Query: 255 TAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCM-NPNYTEF--K 311
+AIDVWSVGC+ EL+ +PLFPG+ V QL + ++LGTPT ++ + N + + +
Sbjct: 831 SAIDVWSVGCIFMELMNKKPLFPGKDHVHQLRLLTELLGTPTEADLGLVKNDDVRRYIRQ 1010
Query: 312 FPQIKAHPWHKIFHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFDELRD 364
PQ P K+F P A+DLV ++L P R L HP+ ++L D
Sbjct: 1011LPQYPRQPLTKVFPHVHPM-AMDLVDKMLTIDPTKRITVEQALAHPYLEKLHD 1166
>TC8968 homologue to UP|Q41619 (Q41619) Protein kinase (Fragment), partial
(32%)
Length = 612
Score = 166 bits (421), Expect = 5e-42
Identities = 75/82 (91%), Positives = 77/82 (93%)
Frame = +1
Query: 303 MNPNYTEFKFPQIKAHPWHKIFHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFDEL 362
MNPNYTEFKFPQIKAHPWHKIFHKRMP EAVDLVSRLLQYSPNLRC ALD LTHPFFDEL
Sbjct: 1 MNPNYTEFKFPQIKAHPWHKIFHKRMPPEAVDLVSRLLQYSPNLRCTALDALTHPFFDEL 180
Query: 363 RDPNARLPTGRFLPPLFNFKPH 384
R+P+ RLPTGRFLPPLFNFK H
Sbjct: 181 REPSTRLPTGRFLPPLFNFKSH 246
>AV414194
Length = 236
Score = 154 bits (388), Expect = 3e-38
Identities = 69/78 (88%), Positives = 77/78 (98%)
Frame = +3
Query: 266 LAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFH 325
LAELLLGQP+FPG+SGVDQLVEIIK+LGTPTREEIKCMNPNYTEFKFPQIKAHPWHK+FH
Sbjct: 3 LAELLLGQPMFPGDSGVDQLVEIIKILGTPTREEIKCMNPNYTEFKFPQIKAHPWHKVFH 182
Query: 326 KRMPAEAVDLVSRLLQYS 343
K++P EA+DLVSR+LQYS
Sbjct: 183 KKVPLEALDLVSRMLQYS 236
>TC8547 homologue to UP|MMK1_MEDSA (Q07176) Mitogen-activated protein
kinase homolog MMK1 (MAP kinase MSK7) (MAP kinase ERK1)
, partial (71%)
Length = 1098
Score = 153 bits (386), Expect = 5e-38
Identities = 80/219 (36%), Positives = 130/219 (58%), Gaps = 2/219 (0%)
Frame = +1
Query: 148 LVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHRCIGVCHRDIKPQNLL 207
+ E + +H++I+ NQ + + + + YQI R L YIH V HRD+KP NLL
Sbjct: 70 IAYELMDTDLHQIIRS----NQGLSEEHCQYFLYQILRGLKYIHSA-NVLHRDLKPSNLL 234
Query: 208 VNPHTHQVKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTTAIDVWSVGCVLA 267
+N + +K+CDFG A+V + + Y+ +R+YRAPEL+ +++YT AIDVWSVGC+
Sbjct: 235 LNANC-DLKICDFGLARVTSETDFMTEYVVTRWYRAPELLLNSSDYTAAIDVWSVGCIFM 411
Query: 268 ELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEF--KFPQIKAHPWHKIFH 325
EL+ +PLFPG V QL +++++GTP+ +++ +N N + + P + + + F
Sbjct: 412 ELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFLNENAKRYIRQLPPYRRQSFQEKFP 591
Query: 326 KRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFDELRD 364
+ P EA+DLV ++L + P R + L HP+ L D
Sbjct: 592 QVHP-EAIDLVEKMLTFDPRKRITVEEALAHPYLTSLHD 705
>TC17660 homologue to UP|Q9M6R8 (Q9M6R8) MAP kinase PsMAPK2, partial (64%)
Length = 717
Score = 135 bits (339), Expect = 1e-32
Identities = 83/239 (34%), Positives = 130/239 (53%), Gaps = 10/239 (4%)
Frame = +2
Query: 101 VAIKKVLQ------DKRYKNRELQTMRLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVP 154
VAIKK+ D REL+ +R L H NV++LK + LV E +
Sbjct: 8 VAIKKIQNAFENRVDALRTLRELKLLRHLHHDNVIALKDIMMPVHRSSFKDVYLVYELMD 187
Query: 155 ETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHRCIGVCHRDIKPQNLLVNPHTHQ 214
+H++IK L+ + + + +Q+ R L Y+H + HRD+KP NLL+N +
Sbjct: 188 TDLHQIIKSSQSLSND----HCQYFLFQLLRGLKYLHSA-NILHRDLKPGNLLINANC-D 349
Query: 215 VKLCDFGSAKV-LVKGEPNISYICSRYYRAPELIFGATEYTTAIDVWSVGCVLAELLLGQ 273
+K+CDFG A++ K + Y+ +R+YRAPEL+ Y T+IDVWSVGC+ AELL +
Sbjct: 350 LKICDFGLARINCSKNQFMTEYVVTRWYRAPELLLCCDNYGTSIDVWSVGCIFAELLGRK 529
Query: 274 PLFPGESGVDQLVEIIKVLGTPTREEIKCM-NPNYTEF--KFPQIKAHPWHKIFHKRMP 329
P+FPG ++QL II +LG+ E+I+ + NP ++ P P+ +++ P
Sbjct: 530 PIFPGSECLNQLKLIINILGSQREEDIEFIDNPKAKKYIKSLPYSIGAPFSRLYPNAHP 706
>TC11249 similar to GB|CAA48472.1|313148|AMMSK3A protein kinase {Medicago
sativa;} , partial (20%)
Length = 461
Score = 119 bits (299), Expect = 6e-28
Identities = 62/86 (72%), Positives = 70/86 (81%)
Frame = +2
Query: 1 MASVGVAPTSGFKESLGDGEIGVDDILPEEMSDMKIRDDREMEATVVDGNGTETGHIIVT 60
MAS V P SGF + +GV+ LP E++ MKIRDD+EMEATVVDG GTETGHIIVT
Sbjct: 206 MASGVVPPASGFTDMNTSSVVGVER-LPHELNVMKIRDDKEMEATVVDGTGTETGHIIVT 382
Query: 61 TIGGRNGQPKQTISYMAERVVGHGSF 86
TIGG+NGQPKQTISYMAERVVG+GSF
Sbjct: 383 TIGGKNGQPKQTISYMAERVVGNGSF 460
>AV766574
Length = 258
Score = 115 bits (288), Expect = 1e-26
Identities = 51/64 (79%), Positives = 57/64 (88%)
Frame = -3
Query: 319 PWHKIFHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFDELRDPNARLPTGRFLPPL 378
PWHK+FHKRMP EA+DL SRLLQYSP+LRC AL+ LTHPFF ELR+PNARLP GR LPPL
Sbjct: 256 PWHKVFHKRMPPEAIDLASRLLQYSPSLRCTALEALTHPFFVELREPNARLPNGRPLPPL 77
Query: 379 FNFK 382
F+FK
Sbjct: 76 FDFK 65
>TC14631 homologue to GB|CAA48472.1|313148|AMMSK3A protein kinase {Medicago
sativa;} , partial (22%)
Length = 572
Score = 110 bits (275), Expect = 4e-25
Identities = 51/62 (82%), Positives = 54/62 (86%)
Frame = +2
Query: 321 HKIFHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFDELRDPNARLPTGRFLPPLFN 380
HKIFHKRMP EAVDLVSRLLQYSPNLR AL+ L HPF+DELR+ N RLP GRFLPPLFN
Sbjct: 2 HKIFHKRMPPEAVDLVSRLLQYSPNLRSTALEALVHPFYDELREANTRLPNGRFLPPLFN 181
Query: 381 FK 382
FK
Sbjct: 182 FK 187
>AV419778
Length = 411
Score = 105 bits (261), Expect = 2e-23
Identities = 48/120 (40%), Positives = 81/120 (67%)
Frame = +1
Query: 179 YTYQIFRALSYIHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICS 238
+ +Q+F+ L+Y+H+ G HRD+KP+NLLV +K+ DFG A+ + P Y+ +
Sbjct: 7 WCFQVFQGLAYMHQR-GYFHRDLKPENLLVTKDI--IKVSDFGLAREISSHPPYTEYVST 177
Query: 239 RYYRAPELIFGATEYTTAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTRE 298
R+YRAPE++ + Y++ +D+W++G ++AEL +PLFPG S D++ +I V+G+PT E
Sbjct: 178 RWYRAPEVLLQSYLYSSKVDMWAMGAIMAELFTLRPLFPGTSEADEIYKICSVIGSPTTE 357
>TC15828 similar to UP|P93774 (P93774) Shaggy-like kinase (Fragment),
partial (31%)
Length = 510
Score = 101 bits (252), Expect = 2e-22
Identities = 45/59 (76%), Positives = 51/59 (86%)
Frame = +3
Query: 324 FHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFDELRDPNARLPTGRFLPPLFNFK 382
FHKRMP EA+DL SRLL+YSP+LRC AL+ HPFFDELR+PNARLP GR LPPLFNF+
Sbjct: 3 FHKRMPPEAIDLASRLLRYSPSLRCSALEACAHPFFDELREPNARLPNGRPLPPLFNFR 179
>AU251534
Length = 391
Score = 99.8 bits (247), Expect = 7e-22
Identities = 53/125 (42%), Positives = 76/125 (60%), Gaps = 2/125 (1%)
Frame = +2
Query: 176 VKLYTYQIFRALSYIHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGE--PNI 233
VK Y Q+ L + H V HRDIK NLL++ + +K+ DFG A V P
Sbjct: 20 VKCYMRQLLSGLEHCHHR-RVLHRDIKGSNLLID-NEGVLKIADFGLASVFDPNHKHPMT 193
Query: 234 SYICSRYYRAPELIFGATEYTTAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLG 293
S + + +YR PEL+ GAT+Y +D+WS GC+L ELL G+P+ PG + V+QL +I K+ G
Sbjct: 194 SRVVTLWYRPPELLLGATDYDVGVDLWSAGCILGELLAGKPIMPGRTEVEQLHKIYKLCG 373
Query: 294 TPTRE 298
+P+ E
Sbjct: 374 SPSDE 388
>TC11248 homologue to UP|P93323 (P93323) Cdc2MsF protein, partial (72%)
Length = 747
Score = 79.0 bits (193), Expect(2) = 1e-21
Identities = 53/178 (29%), Positives = 95/178 (52%), Gaps = 8/178 (4%)
Frame = +1
Query: 81 VGHGSFGVVFQAKCLETGETVAIKKVLQDKRYKN------RELQTMRLLDH-PNVVSLKH 133
VG G++G V++A+ TG+ VA+KK + + RE+ +R+L P+VV L
Sbjct: 67 VGEGTYGKVYRAREKATGKIVALKKTRLHEDDEGVPPTTLREVSILRMLSRDPHVVRLMD 246
Query: 134 CFFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHRC 193
+++ + L LV EY+ + + I+ + + Q +P VK YQ+ + +++ H
Sbjct: 247 VKQGQSKEGKTVLYLVFEYMDTDLKKFIRTFRQTGQNVPPKTVKSLMYQLRKGVAFCHGH 426
Query: 194 IGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISY-ICSRYYRAPELIFGA 250
G+ HRD+KP NLL++ T+ +K+ D G A+ ++ I + +YRA FG+
Sbjct: 427 -GILHRDLKPHNLLMDRETNMLKIADLGLARAFTVPIKKYTHEILTLWYRAR*SPFGS 597
Score = 40.8 bits (94), Expect(2) = 1e-21
Identities = 16/31 (51%), Positives = 23/31 (73%)
Frame = +2
Query: 245 ELIFGATEYTTAIDVWSVGCVLAELLLGQPL 275
E++ GAT Y+ A+D+WSV C+ AEL+ Q L
Sbjct: 581 EVLLGATHYSMAVDMWSVACIFAELVTKQAL 673
>TC14602 UP|Q8GS56 (Q8GS56) Phosphoenolpyruvate carboxylase kinase, complete
Length = 1256
Score = 95.1 bits (235), Expect = 2e-20
Identities = 81/287 (28%), Positives = 134/287 (46%), Gaps = 8/287 (2%)
Frame = +2
Query: 81 VGHGSFGVVFQAKCLETGETVAIKKVLQ-------DKRYKNRELQTMRLLD-HPNVVSLK 132
+G G FG +F+ + +T A+K + + D+ E + M LL HPN++ +
Sbjct: 86 IGRGRFGTIFRCFHPISTDTFAVKLIDKSLLADSTDRHCLENEPKYMSLLSPHPNILQI- 262
Query: 133 HCFFSTTEKDELYLNLVLEYVPETVHRVIKHYSKLNQRMPMIYVKLYTYQIFRALSYIHR 192
F E D++ L++V+E + + + + + +P + Q+ A+++ HR
Sbjct: 263 ---FDVFEDDDV-LSMVIELC-QPLTLLDRIVAANGTSIPEVEAAGLMKQLLEAVAHCHR 427
Query: 193 CIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATE 252
+GV HRD+KP N+L +KL DFGSA+ G + + YY APE++ G E
Sbjct: 428 -LGVAHRDVKPDNVLFGGGG-DLKLADFGSAEWFGDGRRMSGVVGTPYYVAPEVLMGR-E 598
Query: 253 YTTAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKF 312
Y +DVWS G +L +L G P F G+S + +I+ +F
Sbjct: 599 YGEKVDVWSCGVILYIMLSGTPPFYGDSAAEIFEAVIR-----------------GNLRF 727
Query: 313 PQIKAHPWHKIFHKRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFF 359
P +IF PA A DL+ +++ P+ R A L HP+F
Sbjct: 728 PS-------RIFRNVSPA-AKDLLRKMICRDPSNRISAEQALRHPWF 844
>BI419436
Length = 601
Score = 95.1 bits (235), Expect = 2e-20
Identities = 58/151 (38%), Positives = 86/151 (56%), Gaps = 6/151 (3%)
Frame = +1
Query: 215 VKLCDFGSAKVLVKG--EPNISYICSRYYRAPELIFGATEYTTAIDVWSVGCVLAELLLG 272
+K+ DFG A G +P S + + +YR PEL+ GAT+Y ++D+WSVGCV AELL+G
Sbjct: 22 LKVADFGLANFTSSGHKQPLTSRVVTLWYRPPELLLGATDYGPSVDLWSVGCVFAELLVG 201
Query: 273 QPLFPGESGVDQLVEIIKVLGTPTREEIKCMN-PNYTEFKFPQIKAHPWHKIFH---KRM 328
+P+ G + V+QL +I K+ G+P + K P+ T FK PQ HP+ K +
Sbjct: 202 KPVLQGRTEVEQLHKIFKLCGSPPEDYWKKTRLPHATLFK-PQ---HPYDSCLRESFKDL 369
Query: 329 PAEAVDLVSRLLQYSPNLRCQALDCLTHPFF 359
P +V+L+ LL P R A L+ +F
Sbjct: 370 PPASVNLLQTLLSVEPYKRGTATSALSSEYF 462
>TC11735 similar to GB|CAA68872.1|1504063|ATASKKAP shaggy-like kinase kappa
{Arabidopsis thaliana;} , partial (23%)
Length = 564
Score = 94.0 bits (232), Expect = 4e-20
Identities = 43/58 (74%), Positives = 45/58 (77%)
Frame = +3
Query: 326 KRMPAEAVDLVSRLLQYSPNLRCQALDCLTHPFFDELRDPNARLPTGRFLPPLFNFKP 383
KR+P EAVDLV R QYSPNLRC AL+ HPFFDELRDP RLP GR LPPLFNF P
Sbjct: 6 KRLPPEAVDLVCRFFQYSPNLRCTALEACIHPFFDELRDPTPRLPNGRPLPPLFNFNP 179
>TC17375 mitogen-activated kinase kinase kinase alpha [Lotus corniculatus
var. japonicus]
Length = 1422
Score = 92.0 bits (227), Expect = 1e-19
Identities = 70/255 (27%), Positives = 119/255 (46%), Gaps = 1/255 (0%)
Frame = +3
Query: 114 NRELQTMRLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPE-TVHRVIKHYSKLNQRMP 172
N+E+ + HPN+V + +E E L++ LEYV ++H++++ Y + +
Sbjct: 75 NQEINLLNQFSHPNIVQ-----YYGSELGEESLSVYLEYVSGGSIHKLLQEYGAFKEPV- 236
Query: 173 MIYVKLYTYQIFRALSYIHRCIGVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPN 232
++ YT QI L+Y+H V HRDIK N+LV+P+ ++KL DFG +K +
Sbjct: 237 ---IQNYTRQIVSGLAYLHSRNTV-HRDIKGANILVDPNG-EIKLADFGMSKHINSAASM 401
Query: 233 ISYICSRYYRAPELIFGATEYTTAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVL 292
+S+ S Y+ APE++ Y +D+ S+GC + E+ +P + GV + +I
Sbjct: 402 LSFKGSPYWMAPEVVMNTNGYGLPVDISSLGCTILEMATSKPPWSQFEGVAAIFKI---- 569
Query: 293 GTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFHKRMPAEAVDLVSRLLQYSPNLRCQALD 352
P+I H + +A + + + LQ P R A
Sbjct: 570 --------------GNSKDMPEIPEH---------LSDDAKNFIKQCLQRDPLARPTAQS 680
Query: 353 CLTHPFFDELRDPNA 367
L HPF +RD +A
Sbjct: 681 LLNHPF---IRDQSA 716
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.140 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,892,369
Number of Sequences: 28460
Number of extensions: 103757
Number of successful extensions: 645
Number of sequences better than 10.0: 151
Number of HSP's better than 10.0 without gapping: 583
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 585
length of query: 384
length of database: 4,897,600
effective HSP length: 92
effective length of query: 292
effective length of database: 2,279,280
effective search space: 665549760
effective search space used: 665549760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC147011.7


