

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147011.10 - phase: 0
(165 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
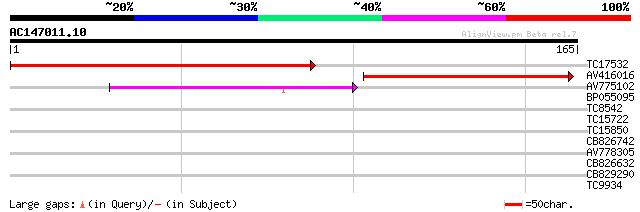
Score E
Sequences producing significant alignments: (bits) Value
TC17532 similar to UP|Q9LZ98 (Q9LZ98) ABC transporter-like prote... 163 1e-41
AV416016 107 8e-25
AV775102 65 6e-12
BP055095 28 0.80
TC8542 similar to UP|RK1_PEA (P49208) 50S ribosomal protein L1, ... 27 1.4
TC15722 similar to UP|Q9FMU0 (Q9FMU0) Similarity to ABC transpor... 26 3.0
TC15850 similar to UP|Q8LAP4 (Q8LAP4) Contains similarity to MYB... 26 3.0
CB826742 26 3.0
AV778305 26 4.0
CB826632 26 4.0
CB829290 25 5.2
TC9934 similar to UP|BAC98494 (BAC98494) AG-motif binding protei... 25 5.2
>TC17532 similar to UP|Q9LZ98 (Q9LZ98) ABC transporter-like protein
(At5g02270), partial (48%)
Length = 476
Score = 163 bits (413), Expect = 1e-41
Identities = 79/89 (88%), Positives = 87/89 (96%)
Frame = +2
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLLKPF+VLLLDEITVDLDVLARADLL FL+KECDERGATIIYATHIFDGLEDWPT++V
Sbjct: 206 MGLLKPFKVLLLDEITVDLDVLARADLLSFLKKECDERGATIIYATHIFDGLEDWPTHMV 385
Query: 61 YVAHGRLELAMPIEKVKETSKLSLMRTVE 89
YVAHG+L+LAMP++KVKE SKLSLMRTVE
Sbjct: 386 YVAHGKLQLAMPMDKVKENSKLSLMRTVE 472
>AV416016
Length = 320
Score = 107 bits (268), Expect = 8e-25
Identities = 52/61 (85%), Positives = 54/61 (88%)
Frame = -2
Query: 104 KERKAAGLPEFDKQVDGSRVVGGDPARAPVRVTNNGWAAGRLHSTIAGEENFLLSSNRVL 163
KERK AGLPEF+KQV G+RV GGDP RA VR NNGWAAGRLHSTIAGEENFLLSSNRVL
Sbjct: 319 KERKDAGLPEFEKQVKGNRVAGGDPGRAAVRAVNNGWAAGRLHSTIAGEENFLLSSNRVL 140
Query: 164 R 164
R
Sbjct: 139 R 137
>AV775102
Length = 494
Score = 65.1 bits (157), Expect = 6e-12
Identities = 34/73 (46%), Positives = 42/73 (56%), Gaps = 1/73 (1%)
Frame = -3
Query: 30 FLRKECDERGATIIYATHIFDGLEDWPTNIVYVAHGRLELAMPIEKVKE-TSKLSLMRTV 88
F ++EC +R ATI YATHIFDGLE W T + Y+ G L+ I V E S +L+ V
Sbjct: 489 FFKEECVQREATIAYATHIFDGLETWATPLAYIPEGELKRTDTISDVNELKSSANLLSVV 310
Query: 89 EVWLRKERDEDRR 101
E WLR RR
Sbjct: 309 ESWLRAAPSLKRR 271
>BP055095
Length = 538
Score = 28.1 bits (61), Expect = 0.80
Identities = 21/81 (25%), Positives = 37/81 (44%), Gaps = 5/81 (6%)
Frame = +3
Query: 3 LLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIY-----ATHIFDGLEDWPT 57
L+ P + L +DEI+ LD + KF+R+ T++ A F+ +D
Sbjct: 144 LVGPAKALFMDEISTGLDSSTTFQICKFMRQMVHIMDVTMVISLLQPAPETFELFDD--- 314
Query: 58 NIVYVAHGRLELAMPIEKVKE 78
I+ ++ G++ P E V E
Sbjct: 315 -IILLSEGQIVYQGPRENVLE 374
>TC8542 similar to UP|RK1_PEA (P49208) 50S ribosomal protein L1,
chloroplast precursor (Fragment), partial (53%)
Length = 672
Score = 27.3 bits (59), Expect = 1.4
Identities = 15/46 (32%), Positives = 21/46 (45%)
Frame = +3
Query: 68 ELAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRKERKAAGLPE 113
+L I VKET+K + TVE R D ++ R LP+
Sbjct: 399 DLKTAISLVKETAKTKFVETVEAHFRLNIDPKYNDQQLRATVSLPK 536
>TC15722 similar to UP|Q9FMU0 (Q9FMU0) Similarity to ABC transporter,
partial (37%)
Length = 624
Score = 26.2 bits (56), Expect = 3.0
Identities = 18/42 (42%), Positives = 24/42 (56%), Gaps = 3/42 (7%)
Frame = +3
Query: 9 VLLLDEITVDLDVLARAD---LLKFLRKECDERGATIIYATH 47
+L+LDE LD ARAD LLK L+KE T++ +H
Sbjct: 129 LLILDEPLAGLDWKARADVVKLLKHLKKE-----LTVLVVSH 239
>TC15850 similar to UP|Q8LAP4 (Q8LAP4) Contains similarity to MYB-related
DNA-binding protein, partial (7%)
Length = 578
Score = 26.2 bits (56), Expect = 3.0
Identities = 18/49 (36%), Positives = 23/49 (46%), Gaps = 2/49 (4%)
Frame = -3
Query: 42 IIYATHIFDGLEDWPTNIVYVAHGRLELAMPIEKVKE--TSKLSLMRTV 88
II A HI+ L NI+ H RL K+ E S+LSL + V
Sbjct: 462 IIGAVHIYTNLRTIHFNIIIFFHSRLIYIYTTSKMNEEKQSRLSLAKIV 316
>CB826742
Length = 465
Score = 26.2 bits (56), Expect = 3.0
Identities = 19/53 (35%), Positives = 23/53 (42%)
Frame = +1
Query: 91 WLRKERDEDRRQRKERKAAGLPEFDKQVDGSRVVGGDPARAPVRVTNNGWAAG 143
W R+E D KER L E + G R+VG + R V N G AG
Sbjct: 178 WQREELDSHVLSHKERFVGSLFEAQNEKRGYRLVGFNRCRF---VCNYGGQAG 327
>AV778305
Length = 361
Score = 25.8 bits (55), Expect = 4.0
Identities = 12/30 (40%), Positives = 20/30 (66%), Gaps = 4/30 (13%)
Frame = +2
Query: 77 KETSKLSLMRTV----EVWLRKERDEDRRQ 102
K+T LS+ + V E+WLRK+ + +RR+
Sbjct: 11 KQTQTLSINQGVVALQEIWLRKKEENNRRK 100
>CB826632
Length = 536
Score = 25.8 bits (55), Expect = 4.0
Identities = 14/56 (25%), Positives = 27/56 (48%)
Frame = +3
Query: 69 LAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRKERKAAGLPEFDKQVDGSRVV 124
L +P+ + L++T L +R+ D+ K ++ A P F +DGS ++
Sbjct: 99 LGLPVVQPYRQHGRHLIKTSLQVLTLQRETDKVMTKRQRTAFAPNFVHSLDGSHMM 266
>CB829290
Length = 546
Score = 25.4 bits (54), Expect = 5.2
Identities = 16/45 (35%), Positives = 24/45 (52%)
Frame = -3
Query: 3 LLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATH 47
LLK +VL+LDE T +D + + LR+ E +T+I H
Sbjct: 457 LLKKSKVLVLDEATASVDTATDNLIQQTLRQHFSE--STVITIAH 329
>TC9934 similar to UP|BAC98494 (BAC98494) AG-motif binding protein-4,
partial (37%)
Length = 1050
Score = 25.4 bits (54), Expect = 5.2
Identities = 15/40 (37%), Positives = 19/40 (47%)
Frame = +2
Query: 113 EFDKQVDGSRVVGGDPARAPVRVTNNGWAAGRLHSTIAGE 152
E +++ GS V GG R +N AAG S AGE
Sbjct: 104 EEEEEEKGSTVSGGSQDRIEEDSNSNSTAAGDYDSIFAGE 223
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.139 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,335,022
Number of Sequences: 28460
Number of extensions: 24364
Number of successful extensions: 124
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 124
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 124
length of query: 165
length of database: 4,897,600
effective HSP length: 83
effective length of query: 82
effective length of database: 2,535,420
effective search space: 207904440
effective search space used: 207904440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC147011.10


