

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147010.4 - phase: 0 /pseudo
(618 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
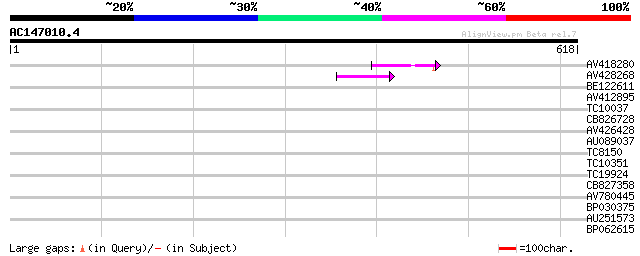
Score E
Sequences producing significant alignments: (bits) Value
AV418280 42 3e-04
AV428268 41 5e-04
BE122611 30 0.002
AV412895 34 0.079
TC10037 34 0.079
CB826728 34 0.079
AV426428 33 0.10
AU089037 33 0.18
TC8150 similar to UP|AAR88248 (AAR88248) Mitochondrial citrate s... 33 0.18
TC10351 similar to UP|Q84TG3 (Q84TG3) At2g35930, partial (20%) 31 0.51
TC19924 28 5.7
CB827358 28 5.7
AV780445 28 5.7
BP030375 27 7.4
AU251573 27 7.4
BP062615 27 9.7
>AV418280
Length = 398
Score = 42.0 bits (97), Expect = 3e-04
Identities = 28/81 (34%), Positives = 35/81 (42%), Gaps = 6/81 (7%)
Frame = -2
Query: 395 CVSCPSNHEDMSHIFFECPFAIHVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELN 454
C C + E H+FF C F++ VW W I AL ++T A F + N
Sbjct: 388 CPFCTTVEECSGHLFFTCVFSMGVWQALHRWLGISVALPASTLA---NFAQFSITARNKN 218
Query: 455 QRLTSL------FWSLWKHRN 469
QRL L WSLW RN
Sbjct: 217 QRLGELAIWIATVWSLWIQRN 155
>AV428268
Length = 429
Score = 41.2 bits (95), Expect = 5e-04
Identities = 18/63 (28%), Positives = 27/63 (42%)
Frame = +3
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
+W +KVP L W + T+V L +G+ C C + E H+ F CP
Sbjct: 78 LWSVKVPSNAIALSWKVLINRVQTKVNLNRRGVVISNVCPLCSLDEESTDHLLFSCPIV* 257
Query: 417 HVW 419
+W
Sbjct: 258 RIW 266
>BE122611
Length = 357
Score = 30.4 bits (67), Expect(2) = 0.002
Identities = 11/31 (35%), Positives = 16/31 (51%)
Frame = -3
Query: 353 FWSGIWKLKVPPKFKNLVWCMCRGFQPTRVR 383
FWS +WK P K + +W + R P+ R
Sbjct: 238 FWSLVWKANAPEKVRFFLWLVGRNALPSNAR 146
Score = 28.1 bits (61), Expect(2) = 0.002
Identities = 9/25 (36%), Positives = 15/25 (60%)
Frame = -2
Query: 395 CVSCPSNHEDMSHIFFECPFAIHVW 419
C+ C + ED +H+F CP + +W
Sbjct: 113 CIRCGAPSEDANHVFRWCPGSAWMW 39
>AV412895
Length = 405
Score = 33.9 bits (76), Expect = 0.079
Identities = 20/72 (27%), Positives = 28/72 (38%), Gaps = 3/72 (4%)
Frame = +1
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGI---QCPTNCVSCPSNHEDMSHIFFECP 413
IW + P K VW + G TR LL + + C C E SH+ F C
Sbjct: 82 IWMVPAPLNIKAFVWRVLLGRIQTRDNLLKRQVIHNALDAICPLCGLAEESGSHLLFSCA 261
Query: 414 FAIHVW*MTSLW 425
++ +W W
Sbjct: 262 ESMLIWYECFAW 297
>TC10037
Length = 560
Score = 33.9 bits (76), Expect = 0.079
Identities = 18/41 (43%), Positives = 21/41 (50%), Gaps = 1/41 (2%)
Frame = -3
Query: 380 TRVRLLDKGIQCPTN-CVSCPSNHEDMSHIFFECPFAIHVW 419
TR RL +KG PT C+ C E M H +CP A VW
Sbjct: 435 TRKRLWNKGAVVPTIFCLRCDRLEETMDHALRDCP*ARRVW 313
>CB826728
Length = 509
Score = 33.9 bits (76), Expect = 0.079
Identities = 14/59 (23%), Positives = 23/59 (38%)
Frame = +1
Query: 354 WSGIWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFEC 412
W +W P+ K ++W C G+ P R L +G+ + E H+ C
Sbjct: 331 WKTVWSASSLPRCKEVMWRACAGYLPVRSALKRRGLDVDPSSPWSGLEDEAEVHVLLTC 507
>AV426428
Length = 413
Score = 33.5 bits (75), Expect = 0.10
Identities = 17/68 (25%), Positives = 27/68 (39%)
Frame = +3
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
+WK + P+ W C G P + L +G+ C C + + H CP
Sbjct: 54 LWKAEALPRCCETGWRACLGVLPVKASLHARGLAEDPICPRCLAAPKTADHAVLFCP*VH 233
Query: 417 HVW*MTSL 424
+W +SL
Sbjct: 234 PIWFASSL 257
>AU089037
Length = 245
Score = 32.7 bits (73), Expect = 0.18
Identities = 18/70 (25%), Positives = 27/70 (37%), Gaps = 3/70 (4%)
Frame = -1
Query: 349 PYVGFWSGIWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTN---CVSCPSNHEDM 405
P + +WK+ P + W + TR L + + + C C E
Sbjct: 215 PSSDIFQKLWKVPAPSNAVSFAWRLILDRVQTRGNLRRRQVIQQSEEALCPMCSQCEESS 36
Query: 406 SHIFFECPFA 415
SH+FF CP A
Sbjct: 35 SHLFFSCPLA 6
>TC8150 similar to UP|AAR88248 (AAR88248) Mitochondrial citrate synthase,
partial (98%)
Length = 1956
Score = 32.7 bits (73), Expect = 0.18
Identities = 10/14 (71%), Positives = 11/14 (78%)
Frame = +3
Query: 240 IYCSRWCSLEYWVW 253
+YCS WCS EYW W
Sbjct: 1335 LYCSLWCSEEYWSW 1376
>TC10351 similar to UP|Q84TG3 (Q84TG3) At2g35930, partial (20%)
Length = 702
Score = 31.2 bits (69), Expect = 0.51
Identities = 10/18 (55%), Positives = 13/18 (71%), Gaps = 3/18 (16%)
Frame = +3
Query: 240 IYCSRWCSLEYW---VWC 254
++C RWCSL +W VWC
Sbjct: 99 VFCKRWCSLGWWPNCVWC 152
>TC19924
Length = 525
Score = 27.7 bits (60), Expect = 5.7
Identities = 23/83 (27%), Positives = 33/83 (39%), Gaps = 3/83 (3%)
Frame = -2
Query: 395 CVSCPSNHEDMSHIFFECPFAIHVW*MTSL-WGTIQHAL--SSTTSAINVIFMLLETFSA 451
C C + E + H F C V + W T + S S I + + E F
Sbjct: 308 CPICFAAPETVEHAIFHCNGTRLVGLGSQFQWNTSDASRFDSGLLSKIECLRLREENFVQ 129
Query: 452 ELNQRLTSLFWSLWKHRNLRAWE 474
E + L L W +WK RN+ +E
Sbjct: 128 ECSS-LGCLLWEIWKTRNITVFE 63
>CB827358
Length = 453
Score = 27.7 bits (60), Expect = 5.7
Identities = 22/94 (23%), Positives = 39/94 (41%), Gaps = 1/94 (1%)
Frame = +3
Query: 361 KVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAIHVW* 420
K P+ K+L WC+C R L KG+ +C C + + + F + W
Sbjct: 6 KALPRCKDLAWCVC-----CR*SLWRKGVNSDPSCPLCGNEF*TIFYCSFWWKLIVMSW- 167
Query: 421 MTSLWGTIQHALSSTTS-AINVIFMLLETFSAEL 453
+ W + + T+S + F++ FS +L
Sbjct: 168 RAAKWLLVLYGKRITSSFSRRFRFVVRPQFSGQL 269
>AV780445
Length = 504
Score = 27.7 bits (60), Expect = 5.7
Identities = 9/28 (32%), Positives = 14/28 (49%)
Frame = +2
Query: 398 CPSNHEDMSHIFFECPFAIHVW*MTSLW 425
CP+ ED+ H +CP + +W W
Sbjct: 68 CPAPEEDI*HCLRDCPHSQEIWLRLGAW 151
>BP030375
Length = 419
Score = 27.3 bits (59), Expect = 7.4
Identities = 17/42 (40%), Positives = 24/42 (56%)
Frame = -3
Query: 423 SLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLFWSL 464
+LWGT+Q+ S IF+ LE F N+R+ SL +SL
Sbjct: 132 TLWGTLQYIALS-----EAIFLCLEYF----NRRIQSLLFSL 34
>AU251573
Length = 332
Score = 27.3 bits (59), Expect = 7.4
Identities = 15/59 (25%), Positives = 20/59 (33%), Gaps = 2/59 (3%)
Frame = -1
Query: 412 CPFAIHVW*MTSLWGT--IQHALSSTTSAINVIFMLLETFSAELNQRLTSLFWSLWKHR 468
CPF W + +G I L T V+ + S + FW LW R
Sbjct: 311 CPFGQRCWLLMPCYGVWGIHWWLEKTHQGFGVLVRKIYALSHVMGSNSKPCFWGLWGER 135
>BP062615
Length = 496
Score = 26.9 bits (58), Expect = 9.7
Identities = 10/21 (47%), Positives = 13/21 (61%)
Frame = +3
Query: 450 SAELNQRLTSLFWSLWKHRNL 470
S +N +TS WSLW H +L
Sbjct: 342 STMINHNMTSFPWSLWSHYSL 404
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.364 0.161 0.683
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,097,405
Number of Sequences: 28460
Number of extensions: 243433
Number of successful extensions: 4006
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 2200
Number of HSP's successfully gapped in prelim test: 161
Number of HSP's that attempted gapping in prelim test: 1715
Number of HSP's gapped (non-prelim): 2508
length of query: 618
length of database: 4,897,600
effective HSP length: 96
effective length of query: 522
effective length of database: 2,165,440
effective search space: 1130359680
effective search space used: 1130359680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 58 (26.9 bits)
Medicago: description of AC147010.4


