

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
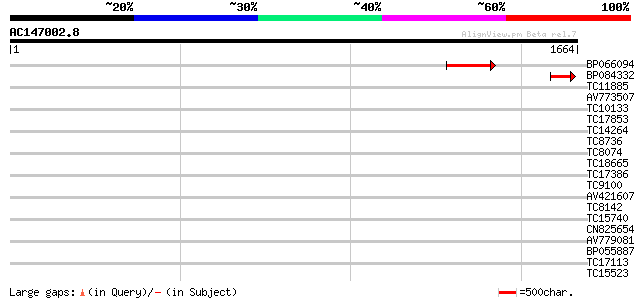
Query= AC147002.8 + phase: 0 /pseudo
(1664 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP066094 102 5e-22
BP084332 65 9e-11
TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 37 0.019
AV773507 35 0.13
TC10133 similar to UP|Q9LSS5 (Q9LSS5) Myosin heavy chain-like, p... 33 0.37
TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, ... 32 0.63
TC14264 similar to UP|Q84NF9 (Q84NF9) Histone H1 (Fragment), par... 32 0.82
TC8736 similar to PIR|T06396|T06396 isoprenylated protein - soyb... 32 0.82
TC8074 similar to UP|AAQ72789 (AAQ72789) 60S ribosomal protein L... 32 1.1
TC18665 similar to UP|O23061 (O23061) BAC IG005I10, partial (65%) 32 1.1
TC17386 similar to UP|Q9LU46 (Q9LU46) DEAD-box protein abstrakt,... 32 1.1
TC9100 similar to UP|Q8W544 (Q8W544) NADH-ubiquinone oxidoreduct... 32 1.1
AV421607 31 1.8
TC8142 UP|BAD11133 (BAD11133) Glutamate-rich protein, complete 30 2.4
TC15740 weakly similar to GB|AAM19935.1|20453393|AY097419 AT5g58... 30 2.4
CN825654 30 2.4
AV779081 30 2.4
BP055887 30 2.4
TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protei... 30 3.1
TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-bindi... 30 3.1
>BP066094
Length = 532
Score = 102 bits (254), Expect = 5e-22
Identities = 68/144 (47%), Positives = 90/144 (62%)
Frame = +1
Query: 1283 GSNQVTSILRN*SWHNIISNGCQKCLS*WCH*RRSVC*TTSWV*GS*AS*PCL*T*EITI 1342
GS Q T ++ + S HN S+ C +CLS W H R S+C +T *P EI++
Sbjct: 94 GSYQTTDLILSESQHNSSSDEC*ECLSKWLHLRGSLCSSTPRFXR*EEL*PYFQIEEISL 273
Query: 1343 WLETSSQSLV**TK*FLN*K*F*KRTS*HNTLQKDS*ERYFDCANIC**YNIWFY*CISL 1402
W E SSQS+V* T+ + K* K S +N+L ++ Y +CAN+C**Y IW * IS+
Sbjct: 274 WSEASSQSMV*ETQLIPSGK*VXKG*SRYNSLLQNLQR*YINCANLC**YYIWIC*SISM 453
Query: 1403 QRIF*VNAG*I*NEYDGRIEILSG 1426
QR+F* +AG*I*N YDGR +ILSG
Sbjct: 454 QRVF*DDAG*I*NAYDGRTQILSG 525
>BP084332
Length = 368
Score = 65.1 bits (157), Expect = 9e-11
Identities = 33/76 (43%), Positives = 47/76 (61%)
Frame = +1
Query: 1586 NYCYVYSGSRVHFSCKLLHTTTLDETSVGRLSDQC*QYSHLL**YCCYLFVKESNSTFKS 1645
N+C +Y R++ S + H+ LDETS G LS+ QY +LL**+CC ES TFK
Sbjct: 37 NHCTIYCRGRIYLSSNMQHSDALDETSAGGLSNP*EQYPNLL**HCCNFIE*ESYLTFKG 216
Query: 1646 QAY*NQTPFYQRLCSK 1661
+A+* + Y RLC++
Sbjct: 217 KAH*GKVSLY*RLCTE 264
>TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (26%)
Length = 555
Score = 37.4 bits (85), Expect = 0.019
Identities = 16/45 (35%), Positives = 29/45 (63%), Gaps = 8/45 (17%)
Frame = +2
Query: 251 KKFLSKRGSYKN--FKKEDQKG------CFNCKKPGHFIADCPDL 287
+K S R SY++ ++++ ++G C NCK+PGHF +CP++
Sbjct: 260 RKIRSDRFSYRDAPYRRDSRRGFSRDNLCKNCKRPGHFARECPNV 394
Score = 31.2 bits (69), Expect = 1.4
Identities = 9/16 (56%), Positives = 13/16 (81%)
Frame = +2
Query: 271 CFNCKKPGHFIADCPD 286
C+NCK+PGH + CP+
Sbjct: 458 CWNCKEPGHMASSCPN 505
>AV773507
Length = 496
Score = 34.7 bits (78), Expect = 0.13
Identities = 14/29 (48%), Positives = 16/29 (54%)
Frame = +2
Query: 257 RGSYKNFKKEDQKGCFNCKKPGHFIADCP 285
RGSY + Q CF C +PGH DCP
Sbjct: 308 RGSYGAGDRVGQDDCFECGRPGHRARDCP 394
>TC10133 similar to UP|Q9LSS5 (Q9LSS5) Myosin heavy chain-like, partial
(34%)
Length = 1827
Score = 33.1 bits (74), Expect = 0.37
Identities = 31/132 (23%), Positives = 56/132 (41%)
Frame = +1
Query: 311 KKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPR 370
+ L A+ + L +S + A++ V L + E+V ++S D +
Sbjct: 292 QSELEASQKHLSVDSAALASAANEIQLLKVQLELVANCESVQTHHAESAD-------VEL 450
Query: 371 QELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKASEEELKGFNLISATYE 430
L +L E LSL E+ N+L + KE TLL+L+ ++ ++ A
Sbjct: 451 LNLKQNLSETLSLVENMKNQLRNFKEFEAQAQGLVSETLLQLETAKRTVEILRGDVAKSV 630
Query: 431 DRLKSLCQKLQE 442
D KS+ +L +
Sbjct: 631 DGYKSIALELDQ 666
>TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, partial
(58%)
Length = 881
Score = 32.3 bits (72), Expect = 0.63
Identities = 13/25 (52%), Positives = 16/25 (64%)
Frame = +2
Query: 271 CFNCKKPGHFIADCPDLQKEKFKGK 295
CF C KPGHF +CP + E +GK
Sbjct: 419 CFKCGKPGHFARECPS-EGETRRGK 490
>TC14264 similar to UP|Q84NF9 (Q84NF9) Histone H1 (Fragment), partial (76%)
Length = 1277
Score = 32.0 bits (71), Expect = 0.82
Identities = 37/139 (26%), Positives = 57/139 (40%), Gaps = 17/139 (12%)
Frame = +2
Query: 507 LKSTF-VPEGTNAKTAVQSKPEA-----------SGSQAKITSKPENLKIKVMTKSDPKS 554
+K++F +P ++ A SKP A S + K +KP K K PKS
Sbjct: 419 VKNSFKLPSAKSSAAAAASKPAAAKKKPQPAAAKSKPKPKAKAKPAAAATKAKAKPAPKS 598
Query: 555 Q--KIKILKRSEP--VHQNLIKPESKIPKQKDQKNKAATASEKTIPKGVKPKVLNDQKP- 609
+ K +++P + KP +K K AA A K P K K + P
Sbjct: 599 KAAATKTTAKAKPAAAAKPKAKPAAKAKPAAKAKAVAAPAKAKASPAKPKAKAKSKTAPR 778
Query: 610 LSIHPKVQGRKSKTSKANP 628
++++P V K +KA P
Sbjct: 779 MNVNPPVPKAK-PAAKAKP 832
>TC8736 similar to PIR|T06396|T06396 isoprenylated protein - soybean
(fragment) {Glycine max;}, partial (84%)
Length = 1268
Score = 32.0 bits (71), Expect = 0.82
Identities = 26/104 (25%), Positives = 43/104 (41%), Gaps = 9/104 (8%)
Frame = +2
Query: 535 ITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLIKPESKIPKQKDQKNKAATASEKT 594
+ PE ++ K+ K + I+I++ L KP+ P++K + +K A +
Sbjct: 377 VCCSPEKIRDKLCCKGGGTIKSIEIVE--------LPKPKPPEPEKKKEADKPKAAEHEK 532
Query: 595 IPKGVKPKVL-------NDQKPLSIHP--KVQGRKSKTSKANPK 629
G KPK + +KP + P K G K K PK
Sbjct: 533 KKDGEKPKAAAEPEKKKDSEKPKAAEPEKKKDGDKEKPKSDGPK 664
>TC8074 similar to UP|AAQ72789 (AAQ72789) 60S ribosomal protein L5, partial
(96%)
Length = 1086
Score = 31.6 bits (70), Expect = 1.1
Identities = 16/52 (30%), Positives = 31/52 (58%), Gaps = 1/52 (1%)
Frame = +3
Query: 285 PDLQKEKFKGKSKKSSFNSSKFRKQIKKS-LMATWEDLDSESGSDKEEADDD 335
P ++K + K K +N K + +K+ L+A + L+S +G+D++E DD+
Sbjct: 744 PTIKKAEKKPPKKSKRYNLKKLTYEERKAKLIARLQALNSAAGADEDEEDDE 899
>TC18665 similar to UP|O23061 (O23061) BAC IG005I10, partial (65%)
Length = 451
Score = 31.6 bits (70), Expect = 1.1
Identities = 20/54 (37%), Positives = 24/54 (44%)
Frame = -2
Query: 319 EDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPRQE 372
EDLDS+ D+E DDD EA EA DE+EV + P E
Sbjct: 189 EDLDSDGFRDEEIGDDDEGE-----GEEEDEATEEASGFDGDEHEVRERAPGDE 43
>TC17386 similar to UP|Q9LU46 (Q9LU46) DEAD-box protein abstrakt, partial
(19%)
Length = 507
Score = 31.6 bits (70), Expect = 1.1
Identities = 17/46 (36%), Positives = 23/46 (49%), Gaps = 2/46 (4%)
Frame = +2
Query: 269 KGCFNCKKPGHFIADCPDLQKEKFK--GKSKKSSFNSSKFRKQIKK 312
KGC C GH I DCP L+ +K ++ F S +R +I K
Sbjct: 203 KGCAYCGGLGHRIRDCPKLEHQKSMAIANNRHDYFGSGGYRGEI*K 340
>TC9100 similar to UP|Q8W544 (Q8W544) NADH-ubiquinone oxidoreductase
(Fragment), partial (97%)
Length = 712
Score = 31.6 bits (70), Expect = 1.1
Identities = 19/41 (46%), Positives = 23/41 (55%)
Frame = -3
Query: 378 KELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKASEEE 418
+ LLSL TN L KEKY+ + KQQ S + E A E E
Sbjct: 644 ERLLSLKLLNTNNLLWRKEKYIQIEKQQSSNVAEDHAFEGE 522
>AV421607
Length = 245
Score = 30.8 bits (68), Expect = 1.8
Identities = 9/16 (56%), Positives = 12/16 (74%)
Frame = +3
Query: 270 GCFNCKKPGHFIADCP 285
GC+ C +PGH+ DCP
Sbjct: 51 GCYKCGRPGHWSRDCP 98
>TC8142 UP|BAD11133 (BAD11133) Glutamate-rich protein, complete
Length = 999
Score = 30.4 bits (67), Expect = 2.4
Identities = 18/81 (22%), Positives = 38/81 (46%), Gaps = 1/81 (1%)
Frame = +1
Query: 552 PKSQKIKILKRSEPVHQNLIKPESKIPKQKDQKNKAATASEKTIPKGVKPK-VLNDQKPL 610
P+ ++ +EP + + ++ PK+ + + +A A E P V+ K V + KP
Sbjct: 214 PEQPAAEVPAPAEPATEEPKEETTEEPKETETETEAPVAPETQDPLEVENKEVTEEAKPE 393
Query: 611 SIHPKVQGRKSKTSKANPKGP 631
+ P+ + + KT + + P
Sbjct: 394 EVKPETEKTEEKTEEPKTEEP 456
Score = 29.6 bits (65), Expect = 4.1
Identities = 19/91 (20%), Positives = 39/91 (41%)
Frame = +1
Query: 512 VPEGTNAKTAVQSKPEASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLI 571
+PE + PE ++ ++P + K T +PK + + P Q+ +
Sbjct: 172 IPEPEQPAATEVAAPEQPAAEVPAPAEPATEEPKEETTEEPKETETETEAPVAPETQDPL 351
Query: 572 KPESKIPKQKDQKNKAATASEKTIPKGVKPK 602
+ E+K ++ + + +EKT K +PK
Sbjct: 352 EVENKEVTEEAKPEEVKPETEKTEEKTEEPK 444
>TC15740 weakly similar to GB|AAM19935.1|20453393|AY097419 AT5g58640/mzn1_90
{Arabidopsis thaliana;}, partial (65%)
Length = 575
Score = 30.4 bits (67), Expect = 2.4
Identities = 17/44 (38%), Positives = 23/44 (51%), Gaps = 3/44 (6%)
Frame = -3
Query: 654 TKVMVPRQRMFKAHDWRESFVPY--PYNERW-RRSEVWWQPNWK 694
T+V +PR+R K H W+ F Y PYN + R+ Q WK
Sbjct: 534 TQVEIPRRRCNKTHHWKNLFPSYYSPYNTKLNNRNHFA*QTLWK 403
>CN825654
Length = 580
Score = 30.4 bits (67), Expect = 2.4
Identities = 39/143 (27%), Positives = 61/143 (42%), Gaps = 9/143 (6%)
Frame = +2
Query: 202 PSKGKTSKSSKAY---KASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRG 258
PSKG + K + A +++++ DG ++++ + + K K+ SK+G
Sbjct: 161 PSKGGSKKGGASLFTASAFDAIDDENDGGEEQEEEEPVVSFAGK------KKPSKGSKKG 322
Query: 259 SYKNFKK----EDQKGCF--NCKKPGHFIADCPDLQKEKFKGKSKKSSFNSSKFRKQIKK 312
F +D G N G D D F GK KKSS S K
Sbjct: 323 GGSLFNAAAFDDDNDGDVMDNSVDDGRDGNDDVDEPVIAFTGK-KKSSKGSKKGGALFSA 499
Query: 313 SLMATWEDLDSESGSDKEEADDD 335
A+++D+D E+ DKEE DD+
Sbjct: 500 ---ASFDDVD-EAKDDKEEDDDE 556
>AV779081
Length = 467
Score = 30.4 bits (67), Expect = 2.4
Identities = 21/82 (25%), Positives = 35/82 (42%), Gaps = 3/82 (3%)
Frame = -2
Query: 476 TYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGL---KSTFVPEGTNAKTAVQSKPEASGSQ 532
T +K Y+ S S D + L +S VP TA+ S PEA+G++
Sbjct: 460 TASSKTLAASYNAWTSPSQKSSSSSDLVNFNLSFSRSVLVPASGLPLTAIPSSPEANGNR 281
Query: 533 AKITSKPENLKIKVMTKSDPKS 554
+ EN++ + + P+S
Sbjct: 280 CTLLLGLENIR*NFLQPTKPRS 215
>BP055887
Length = 506
Score = 30.4 bits (67), Expect = 2.4
Identities = 18/41 (43%), Positives = 23/41 (55%)
Frame = +2
Query: 378 KELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKASEEE 418
++ LSL TN L KEKY+ + KQQ S + E A E E
Sbjct: 44 RKTLSLKLLNTNNLLWRKEKYIQIEKQQSSNVAEDHAFEGE 166
>TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, partial
(25%)
Length = 1077
Score = 30.0 bits (66), Expect = 3.1
Identities = 13/33 (39%), Positives = 16/33 (48%), Gaps = 5/33 (15%)
Frame = -1
Query: 675 PYPYNERWR-RSEVWWQPNW----KDHWYRYYW 702
P + E+WR R E WW W WY Y+W
Sbjct: 816 PARWREQWRERREKWWWRRWWWWWS*TWYYYWW 718
>TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-binding,
abscisic acid-inducible protein, complete
Length = 972
Score = 30.0 bits (66), Expect = 3.1
Identities = 10/27 (37%), Positives = 13/27 (48%), Gaps = 1/27 (3%)
Frame = +1
Query: 677 PYNERWRRSEVWWQPNWKD-HWYRYYW 702
P + WRR WW W W R++W
Sbjct: 586 PGSRPWRRRRWWWIRRWSS*RWIRWWW 666
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.357 0.156 0.559
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,830,610
Number of Sequences: 28460
Number of extensions: 367501
Number of successful extensions: 4311
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 3144
Number of HSP's successfully gapped in prelim test: 149
Number of HSP's that attempted gapping in prelim test: 1009
Number of HSP's gapped (non-prelim): 3493
length of query: 1664
length of database: 4,897,600
effective HSP length: 103
effective length of query: 1561
effective length of database: 1,966,220
effective search space: 3069269420
effective search space used: 3069269420
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC147002.8


