

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147001.15 + phase: 0
(146 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
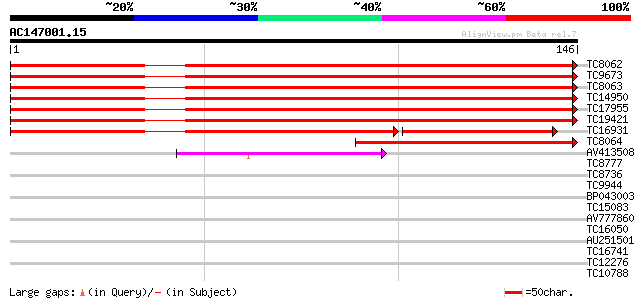
Score E
Sequences producing significant alignments: (bits) Value
TC8062 UP|BAA31218 (BAA31218) Histone H3, complete 235 2e-63
TC9673 UP|BAA31218 (BAA31218) Histone H3, complete 235 2e-63
TC8063 UP|BAA31218 (BAA31218) Histone H3, complete 235 2e-63
TC14950 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {M... 231 5e-62
TC17955 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {M... 231 5e-62
TC19421 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {M... 228 2e-61
TC16931 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {M... 145 3e-54
TC8064 homologue to GB|AAB36496.1|488573|MSU09463 histone H3.2 p... 110 1e-25
AV413508 76 3e-15
TC8777 similar to UP|Q9FFD8 (Q9FFD8) Gb|AAF26981.1 (At5g16470), ... 32 0.057
TC8736 similar to PIR|T06396|T06396 isoprenylated protein - soyb... 27 1.1
TC9944 26 3.1
BP043003 25 4.1
TC15083 similar to UP|GAR1_YEAST (P28007) snoRNP protein GAR1, p... 25 5.3
AV777860 24 9.1
TC16050 similar to UP|O82529 (O82529) Ribosomal protein L27a, pa... 24 9.1
AU251501 24 9.1
TC16741 weakly similar to UP|Q9FNS9 (Q9FNS9) Fructan 1-exohydrol... 24 9.1
TC12276 homologue to UP|R25A_ARATH (Q9SIK2) 40S ribosomal protei... 24 9.1
TC10788 weakly similar to UP|Q8YG29 (Q8YG29) Cytochrome C-type b... 24 9.1
>TC8062 UP|BAA31218 (BAA31218) Histone H3, complete
Length = 768
Score = 235 bits (600), Expect = 2e-63
Identities = 126/146 (86%), Positives = 128/146 (87%)
Frame = +3
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKSTGGKAPRK+LATK AR GG VKKPHR+R GTVALR
Sbjct: 51 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGG----------VKKPHRYRPGTVALR 200
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL
Sbjct: 201 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 380
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPKDIQLARRIRGERA
Sbjct: 381 CAIHAKRVTIMPKDIQLARRIRGERA 458
>TC9673 UP|BAA31218 (BAA31218) Histone H3, complete
Length = 690
Score = 235 bits (600), Expect = 2e-63
Identities = 126/146 (86%), Positives = 128/146 (87%)
Frame = +1
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKSTGGKAPRK+LATK AR GG VKKPHR+R GTVALR
Sbjct: 76 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGG----------VKKPHRYRPGTVALR 225
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL
Sbjct: 226 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 405
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPKDIQLARRIRGERA
Sbjct: 406 CAIHAKRVTIMPKDIQLARRIRGERA 483
>TC8063 UP|BAA31218 (BAA31218) Histone H3, complete
Length = 723
Score = 235 bits (600), Expect = 2e-63
Identities = 126/146 (86%), Positives = 128/146 (87%)
Frame = +2
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKSTGGKAPRK+LATK AR GG VKKPHR+R GTVALR
Sbjct: 71 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGG----------VKKPHRYRPGTVALR 220
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL
Sbjct: 221 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 400
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPKDIQLARRIRGERA
Sbjct: 401 CAIHAKRVTIMPKDIQLARRIRGERA 478
>TC14950 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {Medicago
sativa;}, complete
Length = 641
Score = 231 bits (588), Expect = 5e-62
Identities = 125/146 (85%), Positives = 126/146 (85%)
Frame = +2
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKSTGGKAPRK+LATK AR GG VKKPHRFR GTVALR
Sbjct: 65 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGG----------VKKPHRFRPGTVALR 214
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQS AV ALQEAAEAYLVGLFEDTNL
Sbjct: 215 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSSAVSALQEAAEAYLVGLFEDTNL 394
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPKDIQLARRIRGERA
Sbjct: 395 CAIHAKRVTIMPKDIQLARRIRGERA 472
>TC17955 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {Medicago
sativa;}, complete
Length = 639
Score = 231 bits (588), Expect = 5e-62
Identities = 125/146 (85%), Positives = 126/146 (85%)
Frame = +1
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKSTGGKAPRK+LATK AR GG VKKPHRFR GTVALR
Sbjct: 58 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGG----------VKKPHRFRPGTVALR 207
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQS AV ALQEAAEAYLVGLFEDTNL
Sbjct: 208 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSSAVSALQEAAEAYLVGLFEDTNL 387
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPKDIQLARRIRGERA
Sbjct: 388 CAIHAKRVTIMPKDIQLARRIRGERA 465
>TC19421 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {Medicago
sativa;}, complete
Length = 672
Score = 228 bits (582), Expect = 2e-61
Identities = 124/146 (84%), Positives = 125/146 (84%)
Frame = +3
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKSTGGKAPRK+LATK AR GG VKKPHRFR GTVALR
Sbjct: 93 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGG----------VKKPHRFRPGTVALR 242
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNL 120
EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQS AV ALQEAAEAYLVGLFEDTNL
Sbjct: 243 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSSAVSALQEAAEAYLVGLFEDTNL 422
Query: 121 CAIHAKRVTIMPKDIQLARRIRGERA 146
CAIHAKRVTIMPKDIQLAR IRGERA
Sbjct: 423 CAIHAKRVTIMPKDIQLARXIRGERA 500
>TC16931 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {Medicago
sativa;}, partial (96%)
Length = 440
Score = 145 bits (367), Expect(2) = 3e-54
Identities = 80/100 (80%), Positives = 81/100 (81%)
Frame = +1
Query: 1 MARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHRFRCGTVALR 60
MARTKQTARKSTGGKAPRK+LATK AR GG VKKPHRFR GTVALR
Sbjct: 46 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGG----------VKKPHRFRPGTVALR 195
Query: 61 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAV 100
EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQS AV
Sbjct: 196 EIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSSAV 315
Score = 80.9 bits (198), Expect(2) = 3e-54
Identities = 40/40 (100%), Positives = 40/40 (100%)
Frame = +2
Query: 102 ALQEAAEAYLVGLFEDTNLCAIHAKRVTIMPKDIQLARRI 141
ALQEAAEAYLVGLFEDTNLCAIHAKRVTIMPKDIQLARRI
Sbjct: 320 ALQEAAEAYLVGLFEDTNLCAIHAKRVTIMPKDIQLARRI 439
>TC8064 homologue to GB|AAB36496.1|488573|MSU09463 histone H3.2 precursor
{Medicago sativa;} , partial (45%)
Length = 486
Score = 110 bits (274), Expect = 1e-25
Identities = 55/57 (96%), Positives = 57/57 (99%)
Frame = +1
Query: 90 KTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAKRVTIMPKDIQLARRIRGERA 146
+TDLRFQSHAVLALQEAAEAYLVGLFEDT+LCAIHAKRVTIMPKDIQLARRIRGERA
Sbjct: 79 QTDLRFQSHAVLALQEAAEAYLVGLFEDTHLCAIHAKRVTIMPKDIQLARRIRGERA 249
>AV413508
Length = 265
Score = 75.9 bits (185), Expect = 3e-15
Identities = 46/82 (56%), Positives = 49/82 (59%), Gaps = 28/82 (34%)
Frame = +3
Query: 44 GHVKKPHRFRCGTVALR----------------------------EIRKYQKSTELLIRK 75
G VKKPHR+R GTVALR EIRKYQKSTELLIRK
Sbjct: 18 GGVKKPHRYRPGTVALR*FTVFPQILSRFLSVAED*IFLFVYNFSEIRKYQKSTELLIRK 197
Query: 76 LPFQRLVREIAQDFKTDLRFQS 97
LPFQRLVREIAQDFK L F++
Sbjct: 198LPFQRLVREIAQDFKVILVFKT 263
>TC8777 similar to UP|Q9FFD8 (Q9FFD8) Gb|AAF26981.1 (At5g16470), partial
(74%)
Length = 796
Score = 31.6 bits (70), Expect = 0.057
Identities = 24/73 (32%), Positives = 35/73 (47%), Gaps = 3/73 (4%)
Frame = +3
Query: 10 KSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPH-RFRCG--TVALREIRKYQ 66
K+ K KELA K+ A +GGGK+ K R G K H ++ C V +++ Q
Sbjct: 201 KAKPKKHTAKELAAKLDAATTNRGGGKAGMKDRTGLEKGGHAKWECPHCKVTAPDVKSMQ 380
Query: 67 KSTELLIRKLPFQ 79
E K+PF+
Sbjct: 381 IHHEARHPKIPFE 419
>TC8736 similar to PIR|T06396|T06396 isoprenylated protein - soybean
(fragment) {Glycine max;}, partial (84%)
Length = 1268
Score = 27.3 bits (59), Expect = 1.1
Identities = 16/48 (33%), Positives = 27/48 (55%)
Frame = +3
Query: 2 ARTKQTARKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKP 49
+R + + R+ GK+PR L+ + R PR+ +S+RK + G K P
Sbjct: 507 SRRRLSMRRRKTGKSPRLLLSLRRRRIPRSP-RRRSLRKRKMGTRKSP 647
>TC9944
Length = 605
Score = 25.8 bits (55), Expect = 3.1
Identities = 15/42 (35%), Positives = 23/42 (54%)
Frame = -1
Query: 32 KGGGKSMRKMREGHVKKPHRFRCGTVALREIRKYQKSTELLI 73
+G GKS+RKMRE V K ++++ + TE+LI
Sbjct: 497 QGVGKSLRKMREKSVYK*KLISLLNLSMKNHKNISFITEVLI 372
>BP043003
Length = 493
Score = 25.4 bits (54), Expect = 4.1
Identities = 11/29 (37%), Positives = 17/29 (57%)
Frame = -2
Query: 23 TKVARAPRAKGGGKSMRKMREGHVKKPHR 51
T+ R K GG +M + + HVK+PH+
Sbjct: 186 TRKTRIAGQKEGGATMDQKKLVHVKRPHQ 100
>TC15083 similar to UP|GAR1_YEAST (P28007) snoRNP protein GAR1, partial (7%)
Length = 1367
Score = 25.0 bits (53), Expect = 5.3
Identities = 14/43 (32%), Positives = 19/43 (43%)
Frame = -2
Query: 9 RKSTGGKAPRKELATKVARAPRAKGGGKSMRKMREGHVKKPHR 51
R S GG A R ++ V R +GG R GH + H+
Sbjct: 517 RISGGGGACRGPISGFVRRTSHGRGGASRRRGCVGGHARLRHK 389
>AV777860
Length = 558
Score = 24.3 bits (51), Expect = 9.1
Identities = 13/42 (30%), Positives = 23/42 (53%)
Frame = -3
Query: 71 LLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLV 112
LL+ K P +V +++ F++DL+ Q H + + YLV
Sbjct: 403 LLLVKKP*YTVVPMLSKLFQSDLQLQKHTEELILMVLDIYLV 278
>TC16050 similar to UP|O82529 (O82529) Ribosomal protein L27a, partial (97%)
Length = 634
Score = 24.3 bits (51), Expect = 9.1
Identities = 19/74 (25%), Positives = 32/74 (42%), Gaps = 3/74 (4%)
Frame = -3
Query: 53 RCGTVALREI---RKYQKSTELLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEA 109
RCG + R + Q+ ELL LV E+ + TDL + VL ++A
Sbjct: 344 RCGALGFRGLVLRLLRQERPELLDVDDRAMELVPELVEVAHTDLAEVTRMVLVEEDAVVV 165
Query: 110 YLVGLFEDTNLCAI 123
+ G+ + + A+
Sbjct: 164 HATGVSPSSGMLAV 123
>AU251501
Length = 405
Score = 24.3 bits (51), Expect = 9.1
Identities = 10/20 (50%), Positives = 14/20 (70%)
Frame = +1
Query: 28 APRAKGGGKSMRKMREGHVK 47
APR K S+ K+REG+V+
Sbjct: 199 APRLKDAALSLDKLREGYVE 258
>TC16741 weakly similar to UP|Q9FNS9 (Q9FNS9) Fructan 1-exohydrolase I
precursor, partial (25%)
Length = 680
Score = 24.3 bits (51), Expect = 9.1
Identities = 10/30 (33%), Positives = 17/30 (56%)
Frame = -2
Query: 99 AVLALQEAAEAYLVGLFEDTNLCAIHAKRV 128
++L+L A Y+ +F NLC +HA +
Sbjct: 517 SLLSLSRLATFYIAYIFLIANLCFLHAPSI 428
>TC12276 homologue to UP|R25A_ARATH (Q9SIK2) 40S ribosomal protein S25-1,
complete
Length = 551
Score = 24.3 bits (51), Expect = 9.1
Identities = 10/25 (40%), Positives = 14/25 (56%)
Frame = +2
Query: 16 APRKELATKVARAPRAKGGGKSMRK 40
AP+K+ A + P GGGK +K
Sbjct: 44 APKKDKAPPPSSKPAKSGGGKQKKK 118
>TC10788 weakly similar to UP|Q8YG29 (Q8YG29) Cytochrome C-type biogenesis
protein, partial (25%)
Length = 656
Score = 24.3 bits (51), Expect = 9.1
Identities = 12/32 (37%), Positives = 16/32 (49%)
Frame = +3
Query: 67 KSTELLIRKLPFQRLVREIAQDFKTDLRFQSH 98
K+ L+RK P +I K LRF+SH
Sbjct: 63 KTLNPLLRKFPLLNFPEKIQMASKLALRFRSH 158
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.135 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,717,159
Number of Sequences: 28460
Number of extensions: 16337
Number of successful extensions: 113
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 105
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 106
length of query: 146
length of database: 4,897,600
effective HSP length: 82
effective length of query: 64
effective length of database: 2,563,880
effective search space: 164088320
effective search space used: 164088320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 51 (24.3 bits)
Medicago: description of AC147001.15


