

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147001.13 + phase: 0
(136 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
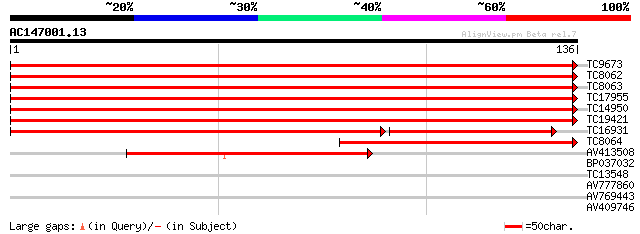
Sequences producing significant alignments: (bits) Value
TC9673 UP|BAA31218 (BAA31218) Histone H3, complete 249 9e-68
TC8062 UP|BAA31218 (BAA31218) Histone H3, complete 249 9e-68
TC8063 UP|BAA31218 (BAA31218) Histone H3, complete 249 9e-68
TC17955 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {M... 240 5e-65
TC14950 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {M... 240 5e-65
TC19421 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {M... 238 3e-64
TC16931 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {M... 161 4e-57
TC8064 homologue to GB|AAB36496.1|488573|MSU09463 histone H3.2 p... 104 6e-24
AV413508 89 3e-19
BP037032 29 0.32
TC13548 similar to GB|AAO63329.1|28950811|BT005265 At5g21920 {Ar... 26 2.7
AV777860 25 3.6
AV769443 24 7.9
AV409746 24 7.9
>TC9673 UP|BAA31218 (BAA31218) Histone H3, complete
Length = 690
Score = 249 bits (637), Expect = 9e-68
Identities = 126/136 (92%), Positives = 131/136 (95%)
Frame = +1
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR+ST GKAPRKQLATKAARKS PTTGG+KKPHRYRPGTVALREIRKYQK TE
Sbjct: 76 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGGVKKPHRYRPGTVALREIRKYQKSTE 255
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
LLIRKLPFQRLVREIAQ+FKTDLRFQSHAVLALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 256 LLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAKRVTI 435
Query: 121 MVKDIQLARRIRGERA 136
M KDIQLARRIRGERA
Sbjct: 436 MPKDIQLARRIRGERA 483
>TC8062 UP|BAA31218 (BAA31218) Histone H3, complete
Length = 768
Score = 249 bits (637), Expect = 9e-68
Identities = 126/136 (92%), Positives = 131/136 (95%)
Frame = +3
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR+ST GKAPRKQLATKAARKS PTTGG+KKPHRYRPGTVALREIRKYQK TE
Sbjct: 51 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGGVKKPHRYRPGTVALREIRKYQKSTE 230
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
LLIRKLPFQRLVREIAQ+FKTDLRFQSHAVLALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 231 LLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAKRVTI 410
Query: 121 MVKDIQLARRIRGERA 136
M KDIQLARRIRGERA
Sbjct: 411 MPKDIQLARRIRGERA 458
>TC8063 UP|BAA31218 (BAA31218) Histone H3, complete
Length = 723
Score = 249 bits (637), Expect = 9e-68
Identities = 126/136 (92%), Positives = 131/136 (95%)
Frame = +2
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR+ST GKAPRKQLATKAARKS PTTGG+KKPHRYRPGTVALREIRKYQK TE
Sbjct: 71 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGGVKKPHRYRPGTVALREIRKYQKSTE 250
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
LLIRKLPFQRLVREIAQ+FKTDLRFQSHAVLALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 251 LLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAKRVTI 430
Query: 121 MVKDIQLARRIRGERA 136
M KDIQLARRIRGERA
Sbjct: 431 MPKDIQLARRIRGERA 478
>TC17955 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {Medicago
sativa;}, complete
Length = 639
Score = 240 bits (613), Expect = 5e-65
Identities = 122/136 (89%), Positives = 128/136 (93%)
Frame = +1
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR+ST GKAPRKQLATKAARKS P TGG+KKPHR+RPGTVALREIRKYQK TE
Sbjct: 58 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 237
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
LLIRKLPFQRLVREIAQ+FKTDLRFQS AV ALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 238 LLIRKLPFQRLVREIAQDFKTDLRFQSSAVSALQEAAEAYLVGLFEDTNLCAIHAKRVTI 417
Query: 121 MVKDIQLARRIRGERA 136
M KDIQLARRIRGERA
Sbjct: 418 MPKDIQLARRIRGERA 465
>TC14950 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {Medicago
sativa;}, complete
Length = 641
Score = 240 bits (613), Expect = 5e-65
Identities = 122/136 (89%), Positives = 128/136 (93%)
Frame = +2
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR+ST GKAPRKQLATKAARKS P TGG+KKPHR+RPGTVALREIRKYQK TE
Sbjct: 65 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 244
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
LLIRKLPFQRLVREIAQ+FKTDLRFQS AV ALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 245 LLIRKLPFQRLVREIAQDFKTDLRFQSSAVSALQEAAEAYLVGLFEDTNLCAIHAKRVTI 424
Query: 121 MVKDIQLARRIRGERA 136
M KDIQLARRIRGERA
Sbjct: 425 MPKDIQLARRIRGERA 472
>TC19421 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {Medicago
sativa;}, complete
Length = 672
Score = 238 bits (607), Expect = 3e-64
Identities = 121/136 (88%), Positives = 127/136 (92%)
Frame = +3
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR+ST GKAPRKQLATKAARKS P TGG+KKPHR+RPGTVALREIRKYQK TE
Sbjct: 93 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 272
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITV 120
LLIRKLPFQRLVREIAQ+FKTDLRFQS AV ALQEA EAYLVGLFEDTNLCAIHAKR+T+
Sbjct: 273 LLIRKLPFQRLVREIAQDFKTDLRFQSSAVSALQEAAEAYLVGLFEDTNLCAIHAKRVTI 452
Query: 121 MVKDIQLARRIRGERA 136
M KDIQLAR IRGERA
Sbjct: 453 MPKDIQLARXIRGERA 500
>TC16931 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {Medicago
sativa;}, partial (96%)
Length = 440
Score = 161 bits (407), Expect(2) = 4e-57
Identities = 81/90 (90%), Positives = 85/90 (94%)
Frame = +1
Query: 1 MARTKQTARRSTLGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGTVALREIRKYQKGTE 60
MARTKQTAR+ST GKAPRKQLATKAARKS P TGG+KKPHR+RPGTVALREIRKYQK TE
Sbjct: 46 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 225
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAV 90
LLIRKLPFQRLVREIAQ+FKTDLRFQS AV
Sbjct: 226 LLIRKLPFQRLVREIAQDFKTDLRFQSSAV 315
Score = 75.1 bits (183), Expect(2) = 4e-57
Identities = 36/40 (90%), Positives = 38/40 (95%)
Frame = +2
Query: 92 ALQEAVEAYLVGLFEDTNLCAIHAKRITVMVKDIQLARRI 131
ALQEA EAYLVGLFEDTNLCAIHAKR+T+M KDIQLARRI
Sbjct: 320 ALQEAAEAYLVGLFEDTNLCAIHAKRVTIMPKDIQLARRI 439
>TC8064 homologue to GB|AAB36496.1|488573|MSU09463 histone H3.2 precursor
{Medicago sativa;} , partial (45%)
Length = 486
Score = 104 bits (259), Expect = 6e-24
Identities = 51/57 (89%), Positives = 55/57 (96%)
Frame = +1
Query: 80 KTDLRFQSHAVLALQEAVEAYLVGLFEDTNLCAIHAKRITVMVKDIQLARRIRGERA 136
+TDLRFQSHAVLALQEA EAYLVGLFEDT+LCAIHAKR+T+M KDIQLARRIRGERA
Sbjct: 79 QTDLRFQSHAVLALQEAAEAYLVGLFEDTHLCAIHAKRVTIMPKDIQLARRIRGERA 249
>AV413508
Length = 265
Score = 88.6 bits (218), Expect = 3e-19
Identities = 50/87 (57%), Positives = 54/87 (61%), Gaps = 28/87 (32%)
Frame = +3
Query: 29 SVPTTGGIKKPHRYRPGTVALR----------------------------EIRKYQKGTE 60
S PTTGG+KKPHRYRPGTVALR EIRKYQK TE
Sbjct: 3 SAPTTGGVKKPHRYRPGTVALR*FTVFPQILSRFLSVAED*IFLFVYNFSEIRKYQKSTE 182
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQS 87
LLIRKLPFQRLVREIAQ+FK L F++
Sbjct: 183LLIRKLPFQRLVREIAQDFKVILVFKT 263
>BP037032
Length = 577
Score = 28.9 bits (63), Expect = 0.32
Identities = 21/69 (30%), Positives = 32/69 (45%), Gaps = 1/69 (1%)
Frame = +1
Query: 23 TKAARKSVPTTGGIKK-PHRYRPGTVALREIRKYQKGTELLIRKLPFQRLVREIAQNFKT 81
T+ R++ P G + PHR RP V + + + LIR LP R++R+ + K
Sbjct: 142 TQRPRRTNPPPGAERHHPHRLRPRNVRVHTVSDQKPLGSELIRILPHVRILRQ-PRRVKL 318
Query: 82 DLRFQSHAV 90
R H V
Sbjct: 319 HARVFRHGV 345
>TC13548 similar to GB|AAO63329.1|28950811|BT005265 At5g21920 {Arabidopsis
thaliana;}, partial (22%)
Length = 515
Score = 25.8 bits (55), Expect = 2.7
Identities = 14/34 (41%), Positives = 19/34 (55%)
Frame = -3
Query: 13 LGKAPRKQLATKAARKSVPTTGGIKKPHRYRPGT 46
L KA R + A R SVP GGI +++R G+
Sbjct: 135 LVKAFRTRNARMGERSSVPPRGGINPRNKFRYGS 34
>AV777860
Length = 558
Score = 25.4 bits (54), Expect = 3.6
Identities = 13/42 (30%), Positives = 24/42 (56%)
Frame = -3
Query: 61 LLIRKLPFQRLVREIAQNFKTDLRFQSHAVLALQEAVEAYLV 102
LL+ K P +V +++ F++DL+ Q H + ++ YLV
Sbjct: 403 LLLVKKP*YTVVPMLSKLFQSDLQLQKHTEELILMVLDIYLV 278
>AV769443
Length = 546
Score = 24.3 bits (51), Expect = 7.9
Identities = 9/23 (39%), Positives = 14/23 (60%)
Frame = -3
Query: 90 VLALQEAVEAYLVGLFEDTNLCA 112
++ +Q +E L+GLF N CA
Sbjct: 187 IVEIQTHIELLLIGLFSKINRCA 119
>AV409746
Length = 411
Score = 24.3 bits (51), Expect = 7.9
Identities = 12/41 (29%), Positives = 22/41 (53%), Gaps = 6/41 (14%)
Frame = -2
Query: 27 RKSVPTTGGIKKPHRYRPGTVALREI------RKYQKGTEL 61
R+ +P GG+ +PHR LR++ R++++G L
Sbjct: 230 RRGIPVGGGLLRPHRSSRPRRRLRKLECNWRSRRWRRGRGL 108
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.135 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,501,038
Number of Sequences: 28460
Number of extensions: 13791
Number of successful extensions: 85
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 84
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 85
length of query: 136
length of database: 4,897,600
effective HSP length: 81
effective length of query: 55
effective length of database: 2,592,340
effective search space: 142578700
effective search space used: 142578700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 50 (23.9 bits)
Medicago: description of AC147001.13


