

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147000.4 + phase: 1 /pseudo
(187 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
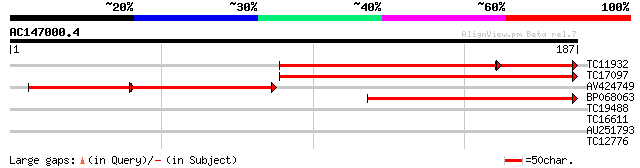
Sequences producing significant alignments: (bits) Value
TC11932 similar to UP|KAPS_CATRO (O49204) Adenylylsulfate kinase... 149 2e-45
TC17097 similar to UP|Q9SE92 (Q9SE92) Adenosine-5'-phosphosulfat... 140 1e-34
AV424749 74 9e-26
BP068063 108 6e-25
TC19488 similar to UP|MRAW_BUCAP (O85295) S-adenosyl-methyltrans... 26 4.8
TC16611 homologue to GB|AAA85390.1|1161312|ATU04876 myo-inositol... 26 4.8
AU251793 25 6.3
TC12776 25 8.2
>TC11932 similar to UP|KAPS_CATRO (O49204) Adenylylsulfate kinase,
chloroplast precursor (APS kinase)
(Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase) ,
partial (30%)
Length = 764
Score = 149 bits (376), Expect(2) = 2e-45
Identities = 67/73 (91%), Positives = 72/73 (97%)
Frame = +2
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+KDRDACRALLP+GDF+EVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 17 LISPYRKDRDACRALLPKGDFVEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 196
Query: 150 EPPCCCEIILQQK 162
EPPC CEI+LQQ+
Sbjct: 197 EPPCSCEIVLQQE 235
Score = 48.5 bits (114), Expect(2) = 2e-45
Identities = 21/27 (77%), Positives = 25/27 (91%)
Frame = +3
Query: 161 QKGSDCKSPKDMAETVISYLEKSGHLQ 187
+KGSDCKSP D+AE VISYL+KSG+LQ
Sbjct: 231 RKGSDCKSPNDLAEIVISYLQKSGYLQ 311
>TC17097 similar to UP|Q9SE92 (Q9SE92) Adenosine-5'-phosphosulfate kinase
(Fragment) , partial (39%)
Length = 576
Score = 140 bits (354), Expect = 1e-34
Identities = 65/98 (66%), Positives = 79/98 (80%)
Frame = +2
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY++DRD CRA+L + FIEVF+++PL +CEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 80 LISPYRRDRDVCRAMLSDAHFIEVFMNMPLELCEARDPKGLYKLARAGKIKGFTGIDDPY 259
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP CEI ++Q+G C +P MA V YLE+ G L+
Sbjct: 260 EPPLNCEIEIKQEGGVCPTPNVMAGQVACYLEEKGFLE 373
>AV424749
Length = 416
Score = 73.6 bits (179), Expect(2) = 9e-26
Identities = 39/48 (81%), Positives = 44/48 (91%)
Frame = +1
Query: 41 QYSCMCFESKLALQRKTDLHP*R*QYSAWSKP*S*F*SRRSFRKHTKD 88
++SCM FESKLALQRKT L P* *Q+SAWS+P*S*F*SRRSFRKHTK+
Sbjct: 196 KFSCMRFESKLALQRKTVLCP*W*QHSAWSEP*S*F*SRRSFRKHTKN 339
Score = 58.5 bits (140), Expect(2) = 9e-26
Identities = 27/35 (77%), Positives = 30/35 (85%)
Frame = +3
Query: 7 RRTFCGMIVQFKNVIDSSCFSKKDVLYG*LVSVVQ 41
R+TFCGM VQF+N ID SCFS+KDVLYG*L S VQ
Sbjct: 87 RQTFCGMTVQFRNRIDRSCFSRKDVLYG*LASAVQ 191
>BP068063
Length = 520
Score = 108 bits (270), Expect = 6e-25
Identities = 50/69 (72%), Positives = 60/69 (86%)
Frame = +1
Query: 119 LHVCEARDPKGLYKLARAGKIKGFTGIDDPYEPPCCCEIILQQKGSDCKSPKDMAETVIS 178
++V EARDPKGLYKLAR GKIKG+TGIDDPYEPP EI++QQKGS+C SP D AE VIS
Sbjct: 46 VNVYEARDPKGLYKLARTGKIKGWTGIDDPYEPPSSSEIVVQQKGSNCMSPSDTAEIVIS 225
Query: 179 YLEKSGHLQ 187
YL+K+G+L+
Sbjct: 226 YLDKNGYLR 252
Score = 34.3 bits (77), Expect = 0.014
Identities = 14/25 (56%), Positives = 18/25 (72%)
Frame = +3
Query: 105 LPEGDFIEVFIDVPLHVCEARDPKG 129
LPEGD IEV++DVP ++ PKG
Sbjct: 3 LPEGDVIEVYLDVPRKCVRSKGPKG 77
>TC19488 similar to UP|MRAW_BUCAP (O85295) S-adenosyl-methyltransferase
mraW , partial (5%)
Length = 502
Score = 25.8 bits (55), Expect = 4.8
Identities = 17/41 (41%), Positives = 20/41 (48%)
Frame = +3
Query: 23 SSCFSKKDVLYG*LVSVVQYSCMCFESKLALQRKTDLHP*R 63
S C SK +V G V + C C E L L TD+HP R
Sbjct: 24 SRCSSKLEVRRG---MVPRGVCECGERLLYLTSHTDIHPER 137
>TC16611 homologue to GB|AAA85390.1|1161312|ATU04876
myo-inositol-1-phosphate synthase {Arabidopsis
thaliana;} , partial (21%)
Length = 527
Score = 25.8 bits (55), Expect = 4.8
Identities = 10/30 (33%), Positives = 14/30 (46%)
Frame = +1
Query: 148 PYEPPCCCEIILQQKGSDCKSPKDMAETVI 177
P PPCC L +GS C + E ++
Sbjct: 157 PLIPPCCYHPQLLDQGSSCSTRHTSGECIV 246
>AU251793
Length = 349
Score = 25.4 bits (54), Expect = 6.3
Identities = 14/43 (32%), Positives = 18/43 (41%), Gaps = 9/43 (20%)
Frame = +1
Query: 152 PCCCEI---------ILQQKGSDCKSPKDMAETVISYLEKSGH 185
P CC L + S KSP++ + SYL SGH
Sbjct: 76 PLCCSSRLIRGRYMEFLNHRRSRAKSPRNEIHGIPSYLNPSGH 204
>TC12776
Length = 560
Score = 25.0 bits (53), Expect = 8.2
Identities = 15/45 (33%), Positives = 19/45 (41%)
Frame = -1
Query: 125 RDPKGLYKLARAGKIKGFTGIDDPYEPPCCCEIILQQKGSDCKSP 169
RDP+G + K G T PP LQQ+ C+SP
Sbjct: 503 RDPRGWLQWILLDKPWGSTPAHSQTPPPEVSATKLQQRTRACRSP 369
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.340 0.150 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,849,890
Number of Sequences: 28460
Number of extensions: 51517
Number of successful extensions: 322
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 320
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 322
length of query: 187
length of database: 4,897,600
effective HSP length: 85
effective length of query: 102
effective length of database: 2,478,500
effective search space: 252807000
effective search space used: 252807000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC147000.4


