

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
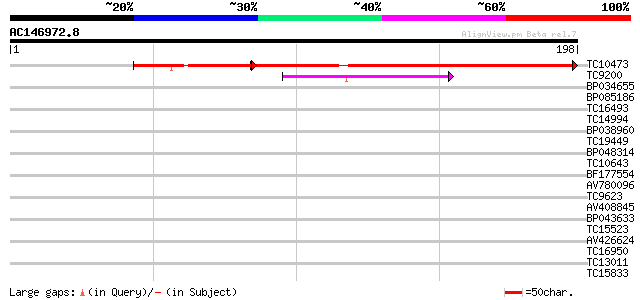
Query= AC146972.8 + phase: 0
(198 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC10473 similar to UP|Q7XBE3 (Q7XBE3) Humj1, partial (69%) 168 4e-55
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 39 5e-04
BP034655 38 0.001
BP085186 37 0.002
TC16493 similar to UP|Q7XTG2 (Q7XTG2) OJ991214_12.5 protein, par... 36 0.004
TC14994 similar to UP|O82469 (O82469) Protein phosphatase-2C , p... 35 0.007
BP038960 35 0.009
TC19449 similar to PIR|E86300|E86300 protein F3O9.30 [imported] ... 34 0.015
BP048314 34 0.019
TC10643 similar to UP|BAC84847 (BAC84847) Acidic nuclear phospho... 34 0.019
BF177554 33 0.025
AV780096 33 0.025
TC9623 homologue to UP|Q8H1K8 (Q8H1K8) Stromal ascorbate peroxid... 33 0.025
AV408845 33 0.033
BP043633 33 0.043
TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-bindi... 33 0.043
AV426624 33 0.043
TC16950 similar to UP|RGP1_HUMAN (P46060) Ran GTPase-activating ... 33 0.043
TC13011 similar to UP|Q9JM93 (Q9JM93) SRp25 nuclear protein, par... 32 0.073
TC15833 similar to UP|Q9FJ12 (Q9FJ12) Genomic DNA, chromosome 5,... 32 0.073
>TC10473 similar to UP|Q7XBE3 (Q7XBE3) Humj1, partial (69%)
Length = 786
Score = 168 bits (425), Expect(2) = 4e-55
Identities = 85/114 (74%), Positives = 89/114 (77%)
Frame = +3
Query: 85 LKTGEIYYINWRNGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISK 144
L+TGEIYYINW+NGMKAKEDPRR +E E EEES YDSEESSSES T SK
Sbjct: 213 LQTGEIYYINWKNGMKAKEDPRRTSEYNVNN---RYYESEEESRYDSEESSSESCTSSSK 383
Query: 145 EQYDQREVIEKQNVLVVAGCKICLMYFMVPKQVEDCPKCSGQLLHFDRSENCSP 198
E Y +EKQNVLVVAGCK CLMYFMVPKQVEDCPKCSGQLLHFDRSEN SP
Sbjct: 384 EHYAPLSCVEKQNVLVVAGCKSCLMYFMVPKQVEDCPKCSGQLLHFDRSENGSP 545
Score = 62.4 bits (150), Expect(2) = 4e-55
Identities = 33/46 (71%), Positives = 36/46 (77%), Gaps = 3/46 (6%)
Frame = +1
Query: 44 GIGISSSSSSSD---ENHISNNSDATLELNSNISLPYHWEQCLDLK 86
GIGISSSS +NH SN SD TLELNS+ISLP+HWEQCLDLK
Sbjct: 40 GIGISSSSDDVPVPLDNHFSN-SDTTLELNSHISLPHHWEQCLDLK 174
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 39.3 bits (90), Expect = 5e-04
Identities = 22/62 (35%), Positives = 33/62 (52%), Gaps = 2/62 (3%)
Frame = +1
Query: 96 RNGMKAKEDPRRAAERECEES--EEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVI 153
+N ++A+ E E EE EEEEEEEEEE + EE E + ++ + + E I
Sbjct: 166 QNKLEAEVQEYEEVEEEVEEEVEEEEEEEEEEEEGEEEEEEEEEEEEVEAEAEEEDDEPI 345
Query: 154 EK 155
+K
Sbjct: 346 QK 351
Score = 37.0 bits (84), Expect = 0.002
Identities = 21/68 (30%), Positives = 33/68 (47%)
Frame = +1
Query: 70 NSNISLPYHWEQCLDLKTGEIYYINWRNGMKAKEDPRRAAERECEESEEEEEEEEEESWY 129
+ + LP+ + L+ + E + + +E+ E E E EEEEEEEEEE
Sbjct: 136 SQSAELPFKLQNKLEAEVQEYEEVEEEVEEEVEEEEEEEEEEEEGEEEEEEEEEEEEVEA 315
Query: 130 DSEESSSE 137
++EE E
Sbjct: 316 EAEEEDDE 339
Score = 32.7 bits (73), Expect = 0.043
Identities = 17/47 (36%), Positives = 25/47 (53%)
Frame = +1
Query: 110 ERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
E E +E EE EEE EEE + EE E +E+ ++ E +E +
Sbjct: 178 EAEVQEYEEVEEEVEEEVEEEEEEEEEEEEGEEEEEEEEEEEEVEAE 318
>BP034655
Length = 517
Score = 38.1 bits (87), Expect = 0.001
Identities = 19/36 (52%), Positives = 21/36 (57%)
Frame = +1
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSE 137
+E+ E E EE EEEEEEEEEE D EE E
Sbjct: 145 EEEEEEEEEEEEEEEEEEEEEEEEEDEEDDEEDEDE 252
Score = 36.6 bits (83), Expect = 0.003
Identities = 18/35 (51%), Positives = 20/35 (56%)
Frame = +1
Query: 103 EDPRRAAERECEESEEEEEEEEEESWYDSEESSSE 137
+D E E EE EEEEEEEEEE + EE E
Sbjct: 136 DDDEEEEEEEEEEEEEEEEEEEEEEEEEDEEDDEE 240
Score = 36.2 bits (82), Expect = 0.004
Identities = 19/38 (50%), Positives = 20/38 (52%)
Frame = +1
Query: 110 ERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQY 147
E E EE EEEEEEEEEE + EE E E Y
Sbjct: 145 EEEEEEEEEEEEEEEEEEEEEEEEEDEEDDEEDEDEDY 258
Score = 35.8 bits (81), Expect = 0.005
Identities = 17/35 (48%), Positives = 20/35 (56%)
Frame = +1
Query: 103 EDPRRAAERECEESEEEEEEEEEESWYDSEESSSE 137
+D E E EE EEEEEEEEEE D E+ +
Sbjct: 139 DDEEEEEEEEEEEEEEEEEEEEEEEEEDEEDDEED 243
Score = 35.0 bits (79), Expect = 0.009
Identities = 17/45 (37%), Positives = 23/45 (50%)
Frame = +1
Query: 98 GMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTII 142
G+ ++ E E EE EEEEEEEEEE + ++ E I
Sbjct: 127 GLSDDDEEEEEEEEEEEEEEEEEEEEEEEEEDEEDDEEDEDEDYI 261
Score = 34.3 bits (77), Expect = 0.015
Identities = 17/36 (47%), Positives = 20/36 (55%)
Frame = +1
Query: 114 EESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQ 149
EE EEEEEEEEEE + EE E +E D+
Sbjct: 145 EEEEEEEEEEEEEEEEEEEEEEEEEDEEDDEEDEDE 252
Score = 32.3 bits (72), Expect = 0.056
Identities = 19/47 (40%), Positives = 21/47 (44%)
Frame = +1
Query: 112 ECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQNV 158
E EE EEEEEEEEEE + EE + E D V V
Sbjct: 145 EEEEEEEEEEEEEEEEEEEEEEEEEDEEDDEEDEDEDYIPVAPSHRV 285
Score = 32.3 bits (72), Expect = 0.056
Identities = 16/40 (40%), Positives = 21/40 (52%)
Frame = +1
Query: 114 EESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVI 153
+E EEEEEEEEEE + EE E E+ + + I
Sbjct: 142 DEEEEEEEEEEEEEEEEEEEEEEEEEDEEDDEEDEDEDYI 261
Score = 32.0 bits (71), Expect = 0.073
Identities = 15/38 (39%), Positives = 21/38 (54%)
Frame = +1
Query: 114 EESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQRE 151
++ EEEEEEEEEE + EE E ++ D+ E
Sbjct: 139 DDEEEEEEEEEEEEEEEEEEEEEEEEEDEEDDEEDEDE 252
Score = 31.2 bits (69), Expect = 0.12
Identities = 19/55 (34%), Positives = 28/55 (50%)
Frame = +1
Query: 108 AAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQNVLVVA 162
AA+ + +EEEEEEEEE + EE E +E + E E ++ + VA
Sbjct: 115 AADFGLSDDDEEEEEEEEEEEEEEEEEEEEEE---EEEDEEDDEEDEDEDYIPVA 270
>BP085186
Length = 388
Score = 37.4 bits (85), Expect = 0.002
Identities = 18/36 (50%), Positives = 23/36 (63%)
Frame = -1
Query: 110 ERECEESEEEEEEEEEESWYDSEESSSESSTIISKE 145
E E EE EEEEEEEEEE + EE +E + ++E
Sbjct: 256 EEEEEEEEEEEEEEEEEEEAEPEEEKNEEENVNTQE 149
Score = 32.3 bits (72), Expect = 0.056
Identities = 17/43 (39%), Positives = 23/43 (52%)
Frame = -1
Query: 109 AERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQRE 151
A+ E E EEEEEEEEEE + EE + +E + +E
Sbjct: 277 ADGEGNEEEEEEEEEEEEEEEEEEEEAEPEEEKNEEENVNTQE 149
Score = 32.3 bits (72), Expect = 0.056
Identities = 18/43 (41%), Positives = 20/43 (45%)
Frame = -1
Query: 97 NGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESS 139
NG D E EE EEEEEEEEEE + E E +
Sbjct: 304 NGKLISPDNADGEGNEEEEEEEEEEEEEEEEEEEEAEPEEEKN 176
Score = 30.0 bits (66), Expect = 0.28
Identities = 15/35 (42%), Positives = 19/35 (53%)
Frame = -1
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEESWYDSEES 134
+ E+ E E EE EEEEEE E E + EE+
Sbjct: 268 EGNEEEEEEEEEEEEEEEEEEEEAEPEEEKNEEEN 164
Score = 29.6 bits (65), Expect = 0.36
Identities = 16/41 (39%), Positives = 23/41 (56%)
Frame = -1
Query: 117 EEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQN 157
EEEEEEEEEE + EE +E ++E+ + E +N
Sbjct: 256 EEEEEEEEEEEEEEEEEEEAEPEEEKNEEENVNTQEGEGEN 134
Score = 27.7 bits (60), Expect = 1.4
Identities = 14/36 (38%), Positives = 19/36 (51%)
Frame = -1
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSE 137
+E+ E E EE EEEE E EEE + ++ E
Sbjct: 256 EEEEEEEEEEEEEEEEEEEAEPEEEKNEEENVNTQE 149
Score = 27.3 bits (59), Expect = 1.8
Identities = 13/38 (34%), Positives = 19/38 (49%)
Frame = -1
Query: 115 ESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREV 152
+ E EEEEEEE + EE E +E+ ++ V
Sbjct: 274 DGEGNEEEEEEEEEEEEEEEEEEEEAEPEEEKNEEENV 161
Score = 26.6 bits (57), Expect = 3.1
Identities = 13/39 (33%), Positives = 19/39 (48%)
Frame = -1
Query: 118 EEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
E EEEEEE + EE E +E+ ++ E + Q
Sbjct: 268 EGNEEEEEEEEEEEEEEEEEEEEAEPEEEKNEEENVNTQ 152
>TC16493 similar to UP|Q7XTG2 (Q7XTG2) OJ991214_12.5 protein, partial (33%)
Length = 769
Score = 36.2 bits (82), Expect = 0.004
Identities = 25/94 (26%), Positives = 40/94 (41%), Gaps = 4/94 (4%)
Frame = +2
Query: 99 MKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQ----YDQREVIE 154
+K P R R E S S ++S ST +++E+ ++ E
Sbjct: 41 LKLNLSPPRVNHRRLESSPNRSATVSPTS-----PTTSCVSTELNQEEGNNNHNHSTSPE 205
Query: 155 KQNVLVVAGCKICLMYFMVPKQVEDCPKCSGQLL 188
+ +V+ GC CLMY M+ + CPKC +L
Sbjct: 206 EATSMVLVGCPRCLMYVMLSEDDPKCPKCKSTVL 307
>TC14994 similar to UP|O82469 (O82469) Protein phosphatase-2C , partial
(89%)
Length = 1672
Score = 35.4 bits (80), Expect = 0.007
Identities = 18/37 (48%), Positives = 23/37 (61%)
Frame = +2
Query: 107 RAAERECEESEEEEEEEEEESWYDSEESSSESSTIIS 143
R E E EE EEEEEEEEEE ++E++ S +S
Sbjct: 158 REEEEEVEEEEEEEEEEEEEEEEEAEKNGG*SRNRVS 268
Score = 32.3 bits (72), Expect = 0.056
Identities = 15/25 (60%), Positives = 17/25 (68%)
Frame = +2
Query: 101 AKEDPRRAAERECEESEEEEEEEEE 125
A+E+ E E EE EEEEEEEEE
Sbjct: 155 AREEEEEVEEEEEEEEEEEEEEEEE 229
Score = 32.0 bits (71), Expect = 0.073
Identities = 16/26 (61%), Positives = 18/26 (68%)
Frame = +2
Query: 109 AERECEESEEEEEEEEEESWYDSEES 134
A E EE EEEEEEEEEE + EE+
Sbjct: 155 AREEEEEVEEEEEEEEEEEEEEEEEA 232
Score = 27.7 bits (60), Expect = 1.4
Identities = 13/28 (46%), Positives = 18/28 (63%)
Frame = +2
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEES 127
+ +E+ E E EE EEEEEEE E++
Sbjct: 158 REEEEEVEEEEEEEEEEEEEEEEEAEKN 241
>BP038960
Length = 524
Score = 35.0 bits (79), Expect = 0.009
Identities = 31/125 (24%), Positives = 45/125 (35%)
Frame = +3
Query: 34 AGEGRGGGGGGIGISSSSSSSDENHISNNSDATLELNSNISLPYHWEQCLDLKTGEIYYI 93
+G G GGGGGG +S + E+ L+ GE
Sbjct: 192 SGSGSGGGGGGYAADGASFAEGEH-------------------------TQLQKGEAE-- 290
Query: 94 NWRNGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVI 153
ED A ++ EEEEEEEEE+ D S I +E + + E I
Sbjct: 291 -----QPRVEDSEEEAADYDDDEEEEEEEEEEDEHADRVAVSGPGKGPILREGFYEVEAI 455
Query: 154 EKQNV 158
++ +
Sbjct: 456 RRKRI 470
>TC19449 similar to PIR|E86300|E86300 protein F3O9.30 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(24%)
Length = 600
Score = 34.3 bits (77), Expect = 0.015
Identities = 18/51 (35%), Positives = 26/51 (50%)
Frame = +1
Query: 134 SSSESSTIISKEQYDQREVIEKQNVLVVAGCKICLMYFMVPKQVEDCPKCS 184
SSS SS I E+ + +++ V GC CL Y +V K CP+C+
Sbjct: 193 SSSSSSIITQAEEECEVKILASP---VATGCPGCLTYVLVTKNNPKCPRCN 336
>BP048314
Length = 394
Score = 33.9 bits (76), Expect = 0.019
Identities = 22/55 (40%), Positives = 27/55 (49%)
Frame = +1
Query: 98 GMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREV 152
G K K D A + EE EEEEEEEEEE + EE + + E + EV
Sbjct: 136 GRKIKGDVSGGAGDKHEEEEEEEEEEEEE---EEEEEEAMCKDCLRNELFVSPEV 291
>TC10643 similar to UP|BAC84847 (BAC84847) Acidic nuclear
phosphoprotein-like protein, partial (5%)
Length = 789
Score = 33.9 bits (76), Expect = 0.019
Identities = 19/56 (33%), Positives = 31/56 (54%), Gaps = 3/56 (5%)
Frame = +2
Query: 114 EESEEEEEEEEEESWYDSEESSSES---STIISKEQYDQREVIEKQNVLVVAGCKI 166
E+ +EEEEE+ EES D E+ S+E T++ +++ D K+ AGC +
Sbjct: 41 EDEDEEEEEDSEESSDDDEDESAEGGSRGTLVDEDENDNGRSSRKR-TRKSAGCSV 205
>BF177554
Length = 246
Score = 33.5 bits (75), Expect = 0.025
Identities = 16/51 (31%), Positives = 25/51 (48%)
Frame = +1
Query: 101 AKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQRE 151
+++ + E E ES EEEEEEEEE + EE E+ ++ +
Sbjct: 13 SEQQDEDSCEEEAGESSEEEEEEEEEGEEEEEEDDGPGDRDAQPEEEEEED 165
Score = 29.6 bits (65), Expect = 0.36
Identities = 15/36 (41%), Positives = 19/36 (52%)
Frame = +1
Query: 98 GMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEE 133
G ++D E E SEEEEEEEEE + E+
Sbjct: 7 GTSEQQDEDSCEEEAGESSEEEEEEEEEGEEEEEED 114
Score = 28.5 bits (62), Expect = 0.81
Identities = 16/50 (32%), Positives = 22/50 (44%)
Frame = +1
Query: 110 ERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQNVL 159
E CEE E EEEEE + EE E ++ + E E ++L
Sbjct: 28 EDSCEEEAGESSEEEEEEEEEGEEEEEEDDGPGDRDAQPEEEEEEDFSLL 177
Score = 25.0 bits (53), Expect = 8.9
Identities = 9/10 (90%), Positives = 9/10 (90%)
Frame = +3
Query: 35 GEGRGGGGGG 44
G GRGGGGGG
Sbjct: 72 GRGRGGGGGG 101
>AV780096
Length = 564
Score = 33.5 bits (75), Expect = 0.025
Identities = 16/41 (39%), Positives = 23/41 (56%)
Frame = +1
Query: 104 DPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISK 144
D R A R +SEEEEEE++EE +D E E +++
Sbjct: 16 DLRDMARRNGSDSEEEEEEQQEEEVHDQIEEDDEELEAVAR 138
>TC9623 homologue to UP|Q8H1K8 (Q8H1K8) Stromal ascorbate peroxidase,
partial (42%)
Length = 640
Score = 33.5 bits (75), Expect = 0.025
Identities = 21/63 (33%), Positives = 33/63 (52%)
Frame = -1
Query: 83 LDLKTGEIYYINWRNGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTII 142
L+ +T + + R M+ +E ERE EE EEE E++E E + EE SE ++
Sbjct: 271 LETRTSALP*LRKR*EMRGEERKHWKEEREEEEEEEENEKKEREEY---EEEESERVALV 101
Query: 143 SKE 145
+ E
Sbjct: 100 AVE 92
>AV408845
Length = 429
Score = 33.1 bits (74), Expect = 0.033
Identities = 19/42 (45%), Positives = 23/42 (54%)
Frame = -3
Query: 98 GMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESS 139
G K + R +E EE EEEEEEEEEE + E S +S
Sbjct: 181 GCGVKNEKRVRG*KEEEEEEEEEEEEEEEGEFVGLEKSPVNS 56
Score = 28.5 bits (62), Expect = 0.81
Identities = 15/31 (48%), Positives = 18/31 (57%)
Frame = -3
Query: 115 ESEEEEEEEEEESWYDSEESSSESSTIISKE 145
+ EEEEEEEEEE + E E S + S E
Sbjct: 142 KEEEEEEEEEEEEEEEGEFVGLEKSPVNSAE 50
Score = 25.0 bits (53), Expect = 8.9
Identities = 14/30 (46%), Positives = 18/30 (59%)
Frame = -3
Query: 110 ERECEESEEEEEEEEEESWYDSEESSSESS 139
E E EE EEEEEE E S +S+E++
Sbjct: 133 EEEEEEEEEEEEEGEFVGLEKSPVNSAETN 44
>BP043633
Length = 542
Score = 32.7 bits (73), Expect = 0.043
Identities = 14/33 (42%), Positives = 19/33 (57%)
Frame = -2
Query: 110 ERECEESEEEEEEEEEESWYDSEESSSESSTII 142
E E E +EEEEEEEE +W + S+E +
Sbjct: 508 EDEAGEEDEEEEEEEETAWGGKNKGSTEEEAAV 410
>TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-binding,
abscisic acid-inducible protein, complete
Length = 972
Score = 32.7 bits (73), Expect = 0.043
Identities = 16/44 (36%), Positives = 27/44 (61%), Gaps = 1/44 (2%)
Frame = +2
Query: 4 ITASLERSLQNCSLNNNNQNEEG-SATIDVVAGEGRGGGGGGIG 46
+T + E+S+++ N Q+ +G + T++ GRGGGGGG G
Sbjct: 494 VTFASEQSMKDAIEGMNGQDIDGRNVTVNEAQARGRGGGGGGGG 625
>AV426624
Length = 422
Score = 32.7 bits (73), Expect = 0.043
Identities = 15/24 (62%), Positives = 19/24 (78%)
Frame = +3
Query: 103 EDPRRAAERECEESEEEEEEEEEE 126
E+ + ++RE EE EEEEEEEEEE
Sbjct: 321 EEE*KGSQREEEEEEEEEEEEEEE 392
Score = 29.6 bits (65), Expect = 0.36
Identities = 14/25 (56%), Positives = 18/25 (72%)
Frame = +3
Query: 102 KEDPRRAAERECEESEEEEEEEEEE 126
+E+ + + E EE EEEEEEEEEE
Sbjct: 321 EEE*KGSQREEEEEEEEEEEEEEEE 395
Score = 28.9 bits (63), Expect = 0.62
Identities = 13/17 (76%), Positives = 14/17 (81%)
Frame = +3
Query: 110 ERECEESEEEEEEEEEE 126
E E EE EEEEEEEEE+
Sbjct: 348 EEEEEEEEEEEEEEEED 398
Score = 28.9 bits (63), Expect = 0.62
Identities = 13/17 (76%), Positives = 14/17 (81%)
Frame = +3
Query: 110 ERECEESEEEEEEEEEE 126
E E EE EEEEEEEE+E
Sbjct: 351 EEEEEEEEEEEEEEEDE 401
Score = 28.5 bits (62), Expect = 0.81
Identities = 13/20 (65%), Positives = 15/20 (75%)
Frame = +3
Query: 106 RRAAERECEESEEEEEEEEE 125
+R E E EE EEEEEEE+E
Sbjct: 342 QREEEEEEEEEEEEEEEEDE 401
Score = 28.1 bits (61), Expect = 1.1
Identities = 13/17 (76%), Positives = 13/17 (76%)
Frame = +3
Query: 114 EESEEEEEEEEEESWYD 130
EE EEEEEEEEEE D
Sbjct: 348 EEEEEEEEEEEEEEEED 398
Score = 26.9 bits (58), Expect = 2.4
Identities = 13/24 (54%), Positives = 16/24 (66%)
Frame = +3
Query: 110 ERECEESEEEEEEEEEESWYDSEE 133
E E + S+ EEEEEEEE + EE
Sbjct: 321 EEE*KGSQREEEEEEEEEEEEEEE 392
Score = 26.9 bits (58), Expect = 2.4
Identities = 13/23 (56%), Positives = 14/23 (60%)
Frame = +3
Query: 102 KEDPRRAAERECEESEEEEEEEE 124
K R E E EE EEEEEE+E
Sbjct: 333 KGSQREEEEEEEEEEEEEEEEDE 401
Score = 26.6 bits (57), Expect = 3.1
Identities = 12/18 (66%), Positives = 13/18 (71%)
Frame = +3
Query: 115 ESEEEEEEEEEESWYDSE 132
E EEEEEEEEEE + E
Sbjct: 348 EEEEEEEEEEEEEEEEDE 401
Score = 26.2 bits (56), Expect = 4.0
Identities = 11/20 (55%), Positives = 14/20 (70%)
Frame = +3
Query: 114 EESEEEEEEEEEESWYDSEE 133
+ EEEEEEEEEE + +E
Sbjct: 342 QREEEEEEEEEEEEEEEEDE 401
>TC16950 similar to UP|RGP1_HUMAN (P46060) Ran GTPase-activating protein 1,
partial (3%)
Length = 643
Score = 32.7 bits (73), Expect = 0.043
Identities = 23/64 (35%), Positives = 32/64 (49%)
Frame = +1
Query: 71 SNISLPYHWEQCLDLKTGEIYYINWRNGMKAKEDPRRAAERECEESEEEEEEEEEESWYD 130
S + P WE+ + +I+ + MK P A E +E EEE++EEEEE D
Sbjct: 211 SPLDFPIEWERPKPGRRPDIF--PQFSPMKTPLPPPIADPPEEDEEEEEKKEEEEEEDPD 384
Query: 131 SEES 134
EES
Sbjct: 385 KEES 396
Score = 28.1 bits (61), Expect = 1.1
Identities = 12/24 (50%), Positives = 17/24 (70%)
Frame = +1
Query: 114 EESEEEEEEEEEESWYDSEESSSE 137
EE EEEEE++EEE D ++ S+
Sbjct: 328 EEDEEEEEKKEEEEEEDPDKEESD 399
Score = 27.3 bits (59), Expect = 1.8
Identities = 13/30 (43%), Positives = 17/30 (56%)
Frame = +1
Query: 104 DPRRAAERECEESEEEEEEEEEESWYDSEE 133
DP E E E+ EEEEEE+ ++ D E
Sbjct: 319 DPPEEDEEEEEKKEEEEEEDPDKEESDKPE 408
>TC13011 similar to UP|Q9JM93 (Q9JM93) SRp25 nuclear protein, partial (10%)
Length = 593
Score = 32.0 bits (71), Expect = 0.073
Identities = 15/38 (39%), Positives = 23/38 (60%)
Frame = -3
Query: 101 AKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSES 138
A +R +E +SEEEEEE+ EES + +E + E+
Sbjct: 552 ADHSKKRKKAKEPSDSEEEEEEDAEESDHSEDEVNGEA 439
>TC15833 similar to UP|Q9FJ12 (Q9FJ12) Genomic DNA, chromosome 5, TAC
clone:K21P3 (Extensin like protein), partial (25%)
Length = 542
Score = 32.0 bits (71), Expect = 0.073
Identities = 18/38 (47%), Positives = 21/38 (54%)
Frame = -1
Query: 22 QNEEGSATIDVVAGEGRGGGGGGIGISSSSSSSDENHI 59
+ E G + VV G GRGGGGGG G + S E HI
Sbjct: 443 EGEGGGGQLLVVEGGGRGGGGGGEG-TGLQGFSHELHI 333
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.306 0.126 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,967,187
Number of Sequences: 28460
Number of extensions: 63156
Number of successful extensions: 1934
Number of sequences better than 10.0: 239
Number of HSP's better than 10.0 without gapping: 957
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1408
length of query: 198
length of database: 4,897,600
effective HSP length: 86
effective length of query: 112
effective length of database: 2,450,040
effective search space: 274404480
effective search space used: 274404480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (22.0 bits)
S2: 53 (25.0 bits)
Medicago: description of AC146972.8


