

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
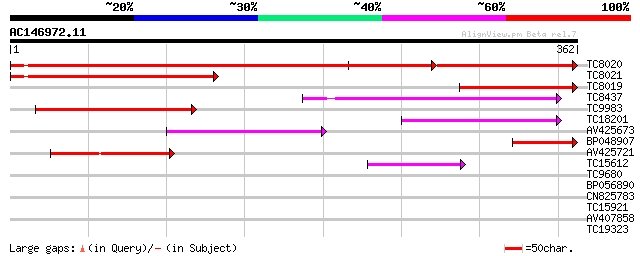
Query= AC146972.11 + phase: 0
(362 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8020 homologue to UP|Q9ZQY2 (Q9ZQY2) Pyruvate dehydrogenase E1... 491 e-140
TC8021 similar to UP|ODPB_PEA (P52904) Pyruvate dehydrogenase E1... 222 7e-59
TC8019 homologue to UP|Q9ZQY2 (Q9ZQY2) Pyruvate dehydrogenase E1... 144 3e-35
TC8437 homologue to GB|AAB86804.1|2454184|ATU80186 pyruvate dehy... 116 7e-27
TC9983 homologue to GB|AAB86804.1|2454184|ATU80186 pyruvate dehy... 98 2e-21
TC18201 homologue to GB|AAB86804.1|2454184|ATU80186 pyruvate deh... 83 6e-17
AV425673 75 1e-14
BP048907 72 1e-13
AV425721 63 7e-11
TC15612 homologue to UP|Q8L693 (Q8L693) 1-deoxy-D-xylulose 5-pho... 43 7e-05
TC9680 homologue to UP|Q9LDY2 (Q9LDY2) Branched chain alpha-keto... 32 0.13
BP056890 29 1.1
CN825783 28 1.8
TC15921 weakly similar to GB|AAB64701.1|603415|SCE9163 Yer174cp ... 28 2.4
AV407858 27 7.0
TC19323 homologue to UP|Q9XH50 (Q9XH50) 1-D-deoxyxylulose 5-phos... 26 9.1
>TC8020 homologue to UP|Q9ZQY2 (Q9ZQY2) Pyruvate dehydrogenase E1 beta
subunit isoform 2 , partial (90%)
Length = 1643
Score = 491 bits (1265), Expect = e-140
Identities = 250/272 (91%), Positives = 259/272 (94%)
Frame = +2
Query: 1 MLGVIRNKVNLLRPSFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQ 60
M GV R K +RP+FS+ R SS AK+MTVR+ALNSALDEEMSADPKVFLMGEEVGEYQ
Sbjct: 170 MWGVTRLKT--IRPAFSSLRQFSSVAKEMTVREALNSALDEEMSADPKVFLMGEEVGEYQ 343
Query: 61 GAYKISKGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHII 120
GAYKI+KGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGL+PVVEFMTFNFSMQAIDHII
Sbjct: 344 GAYKITKGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLRPVVEFMTFNFSMQAIDHII 523
Query: 121 NSAAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDAR 180
NSAAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHS CYASWYGSCPGLKVL+PYSSEDAR
Sbjct: 524 NSAAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHSQCYASWYGSCPGLKVLSPYSSEDAR 703
Query: 181 GLLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSK 240
GLLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSK
Sbjct: 704 GLLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSK 883
Query: 241 MVGFALKAAETLEKEGISAEVINLRSIRPLDR 272
MVG+ALKAAE L KEGISAEVINLRSIRPLDR
Sbjct: 884 MVGYALKAAEILAKEGISAEVINLRSIRPLDR 979
Score = 174 bits (441), Expect = 2e-44
Identities = 95/147 (64%), Positives = 111/147 (74%), Gaps = 1/147 (0%)
Frame = +3
Query: 217 FCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLE-KEGISAEVINLRSIRPLDRATI 275
F P K ++ + K + + F K + + + KE + I +RS R +D +TI
Sbjct: 813 FAFP*EKQRLREKEKMLLLQPFQKWLVMLSRLLRY*QRKESVPRLSICVRSGRLID-STI 989
Query: 276 NASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERIAGADVPMPYAANLER 335
N SVRKTNRLVTVEEGFPQHGVGAEIC SVIEESFGYLDAPVERIAGADVPMPYAANLER
Sbjct: 990 NTSVRKTNRLVTVEEGFPQHGVGAEICTSVIEESFGYLDAPVERIAGADVPMPYAANLER 1169
Query: 336 LAVPQIEDIVRAAKRACHRSVPMAATA 362
+AVPQ+EDIVRAAKRAC+RSVP+AA A
Sbjct: 1170MAVPQVEDIVRAAKRACYRSVPLAAAA 1250
>TC8021 similar to UP|ODPB_PEA (P52904) Pyruvate dehydrogenase E1 component
beta subunit, mitochondrial precursor (PDHE1-B) ,
partial (36%)
Length = 642
Score = 222 bits (566), Expect = 7e-59
Identities = 113/133 (84%), Positives = 122/133 (90%)
Frame = +1
Query: 1 MLGVIRNKVNLLRPSFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQ 60
M GV R K +RP+FS+ R SS AK+MTVR+ALNSALDEEMSADPKVFLMGEEVGEYQ
Sbjct: 250 MWGVTRLKT--IRPAFSSLRQFSSVAKEMTVREALNSALDEEMSADPKVFLMGEEVGEYQ 423
Query: 61 GAYKISKGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHII 120
GAYKI+KGLLEKYGPERVLDTPITEAG TGIGVGAAYYGL+PVVE+MTFNFSMQAIDHII
Sbjct: 424 GAYKITKGLLEKYGPERVLDTPITEAGLTGIGVGAAYYGLRPVVEYMTFNFSMQAIDHII 603
Query: 121 NSAAKSNYMSAGQ 133
NSAAKSNY+SAGQ
Sbjct: 604 NSAAKSNYVSAGQ 642
>TC8019 homologue to UP|Q9ZQY2 (Q9ZQY2) Pyruvate dehydrogenase E1 beta
subunit isoform 2 , partial (20%)
Length = 603
Score = 144 bits (362), Expect = 3e-35
Identities = 69/75 (92%), Positives = 73/75 (97%)
Frame = +1
Query: 288 VEEGFPQHGVGAEICASVIEESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRA 347
VEEGFPQHGVGAEIC SVIEESFGYLDAPVERIAGADVPMPYAANLER+AVPQ+EDIVRA
Sbjct: 1 VEEGFPQHGVGAEICTSVIEESFGYLDAPVERIAGADVPMPYAANLERMAVPQVEDIVRA 180
Query: 348 AKRACHRSVPMAATA 362
AKRAC+RSVP+AA A
Sbjct: 181 AKRACYRSVPLAAAA 225
>TC8437 homologue to GB|AAB86804.1|2454184|ATU80186 pyruvate dehydrogenase
E1 beta subunit {Arabidopsis thaliana;} , partial (40%)
Length = 915
Score = 116 bits (290), Expect = 7e-27
Identities = 63/165 (38%), Positives = 98/165 (59%)
Frame = +1
Query: 188 RDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALK 247
R +PV+ E+ LLY + + D + L + +A++ R G+ VTI +S+M ++
Sbjct: 4 RSDNPVILFEHVLLYN----LKERIPDEEYVLSLEEAEMVRPGEHVTILTYSRMRYHVMQ 171
Query: 248 AAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIE 307
AA+TL +G EVI++RS++P D TI SV+KT+R++ VEE G+GA + A++ E
Sbjct: 172 AAKTLVNKGYDPEVIDIRSLKPFDLHTIGNSVKKTHRVLIVEECMRTGGIGASLTAAISE 351
Query: 308 ESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAKRAC 352
YLDAP+ ++ DVP PYA LE + V Q IV A ++ C
Sbjct: 352 NFNDYLDAPIVCLSSQDVPTPYAGPLEEMTVVQPAQIVTAVEQLC 486
>TC9983 homologue to GB|AAB86804.1|2454184|ATU80186 pyruvate dehydrogenase
E1 beta subunit {Arabidopsis thaliana;} , partial (23%)
Length = 640
Score = 98.2 bits (243), Expect = 2e-21
Identities = 48/103 (46%), Positives = 65/103 (62%)
Frame = +2
Query: 17 SAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPE 76
S+ S ++ + +AL L+EEM DP V +MGE VG Y G+YK++K L K+G
Sbjct: 329 SSVASTSKPGHELLLFEALREGLEEEMERDPNVCVMGENVGHYGGSYKVTKDLANKFGDL 508
Query: 77 RVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHI 119
RVLDTPI E FTG+G+GAA GL+PV+E M F + A + I
Sbjct: 509 RVLDTPIAENAFTGMGIGAAMTGLRPVIEGMNMGFLLLAFNQI 637
>TC18201 homologue to GB|AAB86804.1|2454184|ATU80186 pyruvate dehydrogenase
E1 beta subunit {Arabidopsis thaliana;} , partial (25%)
Length = 636
Score = 83.2 bits (204), Expect = 6e-17
Identities = 44/102 (43%), Positives = 61/102 (59%)
Frame = +2
Query: 251 TLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESF 310
TL +G EVI++RS++P D TI SV+KT+R++ VEE G+GA + A++ E
Sbjct: 2 TLVNKGYDPEVIDIRSLKPFDLHTIGNSVKKTHRVLIVEECMRTGGIGASLTAAITENFN 181
Query: 311 GYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAKRAC 352
YLDAPV ++ DVP PY LE V Q IV A ++ C
Sbjct: 182 DYLDAPVVCLSSQDVPTPYTGPLEEWTVVQPAQIVTAVEQLC 307
>AV425673
Length = 332
Score = 75.5 bits (184), Expect = 1e-14
Identities = 40/103 (38%), Positives = 57/103 (54%), Gaps = 1/103 (0%)
Frame = +1
Query: 101 KPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVP-IVFRGPNGAAAGVGAQHSHCYA 159
+ + E ++ A D I+N AAK Y S Q N + R P GA G HS
Sbjct: 1 RAIAEIQFADYIFPAFDQIVNEAAKFRYRSGNQFNCGGLTIRAPYGAVGHGGHYHSQSPE 180
Query: 160 SWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLY 202
+++ PG+KV+ P S + A+GLL ++IRDP+PVVF E + LY
Sbjct: 181 AFFCHVPGIKVVIPRSPQQAKGLLLSSIRDPNPVVFFEPKWLY 309
>BP048907
Length = 436
Score = 72.4 bits (176), Expect = 1e-13
Identities = 33/41 (80%), Positives = 39/41 (94%)
Frame = -2
Query: 322 GADVPMPYAANLERLAVPQIEDIVRAAKRACHRSVPMAATA 362
GADVP+PYAA+LER+AVP +EDIVRAAKRAC+RSVP+A TA
Sbjct: 435 GADVPLPYAAHLERMAVPPVEDIVRAAKRACYRSVPLAPTA 313
>AV425721
Length = 392
Score = 63.2 bits (152), Expect = 7e-11
Identities = 28/79 (35%), Positives = 48/79 (60%)
Frame = +3
Query: 27 KQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTPITEA 86
K M + A+N AL + +DP+ ++ GE+V + G ++ + GL +++G RVL+TP+ E
Sbjct: 135 KSMNLYSAINQALHIALDSDPRAYVFGEDVS-FGGVFRCTTGLADRFGKNRVLNTPLCEQ 311
Query: 87 GFTGIGVGAAYYGLKPVVE 105
G G G+G A G + + E
Sbjct: 312 GIIGFGIGLAAMGNRAIAE 368
>TC15612 homologue to UP|Q8L693 (Q8L693) 1-deoxy-D-xylulose 5-phosphate
synthase 1 precursor, partial (19%)
Length = 703
Score = 43.1 bits (100), Expect = 7e-05
Identities = 22/63 (34%), Positives = 34/63 (53%)
Frame = +3
Query: 229 EGKDVTITAFSKMVGFALKAAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTV 288
EG+ V + + V L AA +E+ G+ V + R +PLDR+ I + + L+TV
Sbjct: 3 EGERVALLGYGSAVQNCLAAASLVERHGLRITVADARFCKPLDRSLIRSLAKSHEVLITV 182
Query: 289 EEG 291
EEG
Sbjct: 183 EEG 191
>TC9680 homologue to UP|Q9LDY2 (Q9LDY2) Branched chain alpha-keto acid
dehydrogenase E1 beta subunit, partial (13%)
Length = 488
Score = 32.3 bits (72), Expect = 0.13
Identities = 17/46 (36%), Positives = 23/46 (49%)
Frame = +3
Query: 304 SVIEESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAK 349
S++E F L+APV R+ G D P P E +P I+ A K
Sbjct: 3 SIVERCFSRLEAPVARVCGLDTPFPLV--FEPFYMPTKNKILDAIK 134
>BP056890
Length = 581
Score = 29.3 bits (64), Expect = 1.1
Identities = 18/48 (37%), Positives = 26/48 (53%)
Frame = +1
Query: 8 KVNLLRPSFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEE 55
+V LLR F+ F SSS+ T D ++ L+E S FL+GE+
Sbjct: 229 QVLLLRRPFTVFCKSSSSSSDPTSDDYVSRVLEENPSQVQPKFLIGEK 372
>CN825783
Length = 625
Score = 28.5 bits (62), Expect = 1.8
Identities = 20/68 (29%), Positives = 35/68 (51%), Gaps = 1/68 (1%)
Frame = +2
Query: 23 SSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAY-KISKGLLEKYGPERVLDT 81
S S +Q +DE + DPK+ +G ++GE GA+ K+ +G +YG +R++
Sbjct: 263 SKSVRQNGSLTTTQLTIDENLLVDPKLLFIGSKIGE--GAHGKVYEG---RYG-DRIVAI 424
Query: 82 PITEAGFT 89
+ G T
Sbjct: 425 KVLNRGRT 448
>TC15921 weakly similar to GB|AAB64701.1|603415|SCE9163 Yer174cp
{Saccharomyces cerevisiae;} , partial (10%)
Length = 580
Score = 28.1 bits (61), Expect = 2.4
Identities = 25/82 (30%), Positives = 38/82 (45%), Gaps = 6/82 (7%)
Frame = +1
Query: 226 IEREGKDVTITAFSK------MVGFALKAAETLEKEGISAEVINLRSIRPLDRATINASV 279
I++ KD + AF K M GF+ + LE EG+ E +N+ LD N +
Sbjct: 112 IDKLVKDNKVVAFIKGSRSAPMCGFSQRVIGILESEGVDYESVNV-----LDE-DYNYGL 273
Query: 280 RKTNRLVTVEEGFPQHGVGAEI 301
R+T + + FPQ V E+
Sbjct: 274 RETLKKYSNWPTFPQIFVDGEL 339
>AV407858
Length = 420
Score = 26.6 bits (57), Expect = 7.0
Identities = 15/47 (31%), Positives = 22/47 (45%)
Frame = -3
Query: 22 LSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKG 68
L + +VRD D+E + G EVGE+ G K+S+G
Sbjct: 211 LGVGVDEWSVRDG-----DDENGGGEMTVVFG*EVGEFDGGDKVSEG 86
>TC19323 homologue to UP|Q9XH50 (Q9XH50) 1-D-deoxyxylulose 5-phosphate
synthase, partial (15%)
Length = 530
Score = 26.2 bits (56), Expect = 9.1
Identities = 12/31 (38%), Positives = 18/31 (57%)
Frame = +1
Query: 261 VINLRSIRPLDRATINASVRKTNRLVTVEEG 291
V + R +PLD + I + + L+TVEEG
Sbjct: 10 VADARFCKPLDHSLIRSLAKSHEILITVEEG 102
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.134 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,020,284
Number of Sequences: 28460
Number of extensions: 58850
Number of successful extensions: 317
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 315
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 315
length of query: 362
length of database: 4,897,600
effective HSP length: 91
effective length of query: 271
effective length of database: 2,307,740
effective search space: 625397540
effective search space used: 625397540
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146972.11


