

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146972.10 - phase: 0
(106 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
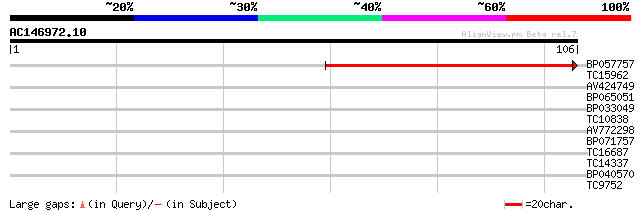
Sequences producing significant alignments: (bits) Value
BP057757 100 5e-23
TC15962 similar to UP|Q9FHC6 (Q9FHC6) Emb|CAB83315.1, partial (13%) 25 1.5
AV424749 25 2.0
BP065051 25 2.0
BP033049 25 2.6
TC10838 weakly similar to PIR|T04232|T04232 pathogenesis-related... 25 2.6
AV772298 24 4.5
BP071757 23 7.6
TC16687 weakly similar to GB|AAP12866.1|30017265|BT006217 At1g10... 23 7.6
TC14337 similar to UP|Q94JQ8 (Q94JQ8) AT3g57880/T10K17_90, parti... 23 9.9
BP040570 23 9.9
TC9752 similar to UP|Q7VR98 (Q7VR98) Glutaminyl-tRNA synthetase ... 23 9.9
>BP057757
Length = 510
Score = 100 bits (248), Expect = 5e-23
Identities = 45/47 (95%), Positives = 47/47 (99%)
Frame = -1
Query: 60 KGHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSKLLSQ 106
+GHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSKLL+Q
Sbjct: 378 QGHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSKLLTQ 238
>TC15962 similar to UP|Q9FHC6 (Q9FHC6) Emb|CAB83315.1, partial (13%)
Length = 602
Score = 25.4 bits (54), Expect = 1.5
Identities = 14/37 (37%), Positives = 18/37 (47%), Gaps = 1/37 (2%)
Frame = -1
Query: 46 TEALETQVK-SKVSIKGHLHTYRFCDNVWTFILQDAL 81
T LET K+ IKG +H C N+ T I A+
Sbjct: 410 THGLETNTSLKKMEIKGTIHEI*ECKNIHTLIKDGAI 300
>AV424749
Length = 416
Score = 25.0 bits (53), Expect = 2.0
Identities = 12/29 (41%), Positives = 17/29 (58%)
Frame = +2
Query: 70 DNVWTFILQDALFKNEDNQENVGRVKIVA 98
DN+ + +D FK ED EN+ R+ VA
Sbjct: 266 DNIRHGLNRDLSFKAEDRSENIRRIGEVA 352
>BP065051
Length = 496
Score = 25.0 bits (53), Expect = 2.0
Identities = 18/66 (27%), Positives = 31/66 (46%), Gaps = 1/66 (1%)
Frame = -1
Query: 17 TETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVS-IKGHLHTYRFCDNVWTF 75
T T E++ G L+ + ++ F+ + E Q KS+ S + +RFCD F
Sbjct: 253 TATRQEVLHIGNLTTRLNLESPCPFNLKVKE*SLVQHKSQHSQLSLPKFRFRFCDEFVLF 74
Query: 76 ILQDAL 81
++ D L
Sbjct: 73 LVWDGL 56
>BP033049
Length = 460
Score = 24.6 bits (52), Expect = 2.6
Identities = 15/58 (25%), Positives = 27/58 (45%), Gaps = 1/58 (1%)
Frame = -2
Query: 23 MVQNGTLSPE-IAIQVLVQFDKSMTEALETQVKSKVSIKGHLHTYRFCDNVWTFILQD 79
+V G + P + I L F + EAL + +K +H + FC+ V +++ D
Sbjct: 249 LVYVGEVRPHAVEIIELEAFVSGVFEALSLHCQYXSDLKFFVHCFTFCNMVIVYLIWD 76
>TC10838 weakly similar to PIR|T04232|T04232 pathogenesis-related protein
homolog F14M19.60 - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (78%)
Length = 914
Score = 24.6 bits (52), Expect = 2.6
Identities = 6/16 (37%), Positives = 12/16 (74%)
Frame = -1
Query: 60 KGHLHTYRFCDNVWTF 75
+ HLH++ F +++W F
Sbjct: 818 ESHLHSFNFSNSIWAF 771
>AV772298
Length = 528
Score = 23.9 bits (50), Expect = 4.5
Identities = 17/64 (26%), Positives = 27/64 (41%), Gaps = 5/64 (7%)
Frame = -2
Query: 17 TETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL-----HTYRFCDN 71
TE D GT+ Q +QF + + +T V + + I+ HL H+ R C
Sbjct: 380 TEEFDYSPWLGTIGCRWYHQTRIQFGEHLATDSDTCVPAHIHIRNHLCSPCHHSRRQCVK 201
Query: 72 VWTF 75
+ F
Sbjct: 200 MEAF 189
>BP071757
Length = 391
Score = 23.1 bits (48), Expect = 7.6
Identities = 8/25 (32%), Positives = 14/25 (56%)
Frame = -3
Query: 62 HLHTYRFCDNVWTFILQDALFKNED 86
H Y+F ++W F LQ ++ +D
Sbjct: 368 HQKPYKFWPHIWNFGLQKNIWLTQD 294
>TC16687 weakly similar to GB|AAP12866.1|30017265|BT006217 At1g10820
{Arabidopsis thaliana;}, partial (11%)
Length = 488
Score = 23.1 bits (48), Expect = 7.6
Identities = 9/17 (52%), Positives = 12/17 (69%)
Frame = +2
Query: 59 IKGHLHTYRFCDNVWTF 75
I+ HL TY+F DN+ F
Sbjct: 26 IRTHLSTYKFQDNIDLF 76
>TC14337 similar to UP|Q94JQ8 (Q94JQ8) AT3g57880/T10K17_90, partial (16%)
Length = 1005
Score = 22.7 bits (47), Expect = 9.9
Identities = 10/20 (50%), Positives = 11/20 (55%), Gaps = 4/20 (20%)
Frame = -2
Query: 58 SIKGHLHTYRFC----DNVW 73
S K H H +RFC D VW
Sbjct: 785 SAKSHPHCFRFCGSCIDMVW 726
>BP040570
Length = 542
Score = 22.7 bits (47), Expect = 9.9
Identities = 10/38 (26%), Positives = 19/38 (49%)
Frame = +2
Query: 36 QVLVQFDKSMTEALETQVKSKVSIKGHLHTYRFCDNVW 73
Q+ + S E ++ ++S + HLH R+C+ W
Sbjct: 377 QIESPVESSSLEPQHSKAHPRIS-QTHLHPGRYCNRFW 487
>TC9752 similar to UP|Q7VR98 (Q7VR98) Glutaminyl-tRNA synthetase , partial
(3%)
Length = 523
Score = 22.7 bits (47), Expect = 9.9
Identities = 10/31 (32%), Positives = 19/31 (61%)
Frame = +1
Query: 67 RFCDNVWTFILQDALFKNEDNQENVGRVKIV 97
+ C+++W F+L L E+ ++N R+K V
Sbjct: 403 KICNSLWFFVLNSRLSWKEE-RDNKFRIKKV 492
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.133 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,471,747
Number of Sequences: 28460
Number of extensions: 13817
Number of successful extensions: 70
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 70
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 70
length of query: 106
length of database: 4,897,600
effective HSP length: 82
effective length of query: 24
effective length of database: 2,563,880
effective search space: 61533120
effective search space used: 61533120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 47 (22.7 bits)
Medicago: description of AC146972.10


