

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146971.7 - phase: 0 /pseudo
(2360 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
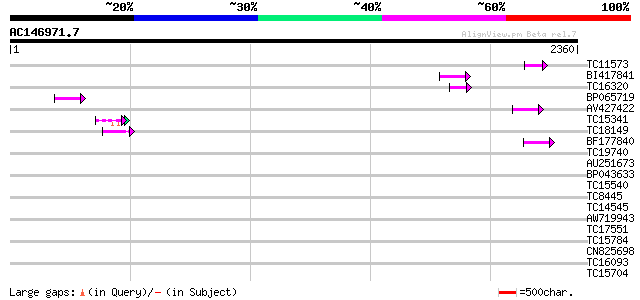
Score E
Sequences producing significant alignments: (bits) Value
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 69 8e-12
BI417841 67 2e-11
TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial ... 56 6e-08
BP065719 55 2e-07
AV427422 53 5e-07
TC15341 similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich pro... 50 4e-06
TC18149 weakly similar to UP|Q91BL3 (Q91BL3) Essential structura... 49 1e-05
BF177840 46 8e-05
TC19740 similar to UP|FKH_DROME (P14734) Fork head protein, part... 41 0.002
AU251673 41 0.002
BP043633 40 0.004
TC15540 similar to UP|Q9NGS5 (Q9NGS5) Prespore protein MF12 (Fra... 40 0.004
TC8445 similar to UP|O80910 (O80910) At2g38410 protein, partial ... 39 0.012
TC14545 similar to UP|Q9FT78 (Q9FT78) P23 co-chaperone, partial ... 38 0.016
AW719943 38 0.021
TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, part... 37 0.036
TC15784 similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precurso... 36 0.079
CN825698 35 0.10
TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transc... 35 0.14
TC15704 similar to UP|O22265 (O22265) Expressed protein (At2g474... 35 0.14
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 68.9 bits (167), Expect = 8e-12
Identities = 38/96 (39%), Positives = 52/96 (53%), Gaps = 1/96 (1%)
Frame = +2
Query: 2143 DNGTNLNNKMMKELCDDFKIEHHNSSPYRPQMNGAVEAANKNI-KRIVQKMVVTYKDWHE 2201
+NG +K ++ CD I+ SS PQ NG EAANK I K I +++ W +
Sbjct: 8 ENGIQFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWID 187
Query: 2202 MLPFALHGYRTSVRTSIGATPFSLVYGMEAVLPVEV 2237
LP L Y T ++SI TPF L YG + +LPVE+
Sbjct: 188 ELPIVLWSYNTMPQSSIKETPF*LTYGADTMLPVEI 295
>BI417841
Length = 617
Score = 67.4 bits (163), Expect = 2e-11
Identities = 41/131 (31%), Positives = 66/131 (50%), Gaps = 2/131 (1%)
Frame = +1
Query: 1789 LIFDGAV--NVYGNGIGAVLLTPKGTHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRI 1846
L FDG+ N G GAVL G+ + + + TNN AEY I+G++ A +
Sbjct: 142 LEFDGSSKGNPGSAGAGAVLRAEDGSKVYLREGVG-NQTNNQAEYRGLILGLKHAHEQGY 318
Query: 1847 KNIEIYGDSALVINQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADAL 1906
++I + GDS LV Q++G W+ + + + A+ L + F +++H+PR N AD
Sbjct: 319 QHINVKGDSQLVCKQVEGSWKARNPNIASLCNEAKELKSKFQSFDINHVPRQYNSEADVQ 498
Query: 1907 ATLSSMIKVNH 1917
A L + H
Sbjct: 499 ANLGVNLPAGH 531
>TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (11%)
Length = 632
Score = 56.2 bits (134), Expect = 6e-08
Identities = 28/91 (30%), Positives = 47/91 (50%)
Frame = +2
Query: 1831 YEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYARRLLTFFNKV 1890
Y I+G++ AI K+I++ GDS LV NQ++G W+ + + A+ L F
Sbjct: 2 YRGLILGLKHAIKEGYKHIQVKGDSMLVCNQVQGLWKIKNQNIASLCSEAKELKNKFLSF 181
Query: 1891 ELHHIPRDENQMADALATLSSMIKVNHHNDV 1921
+++HIPR+ N AD A ++ +V
Sbjct: 182 KINHIPREYNSEADVQANFGISLRAGQVEEV 274
>BP065719
Length = 567
Score = 54.7 bits (130), Expect = 2e-07
Identities = 35/131 (26%), Positives = 60/131 (45%), Gaps = 2/131 (1%)
Frame = +3
Query: 185 QVPHKFKIPDFEKYKGSSCPE--EHLKMYVRRMPAYAQDDQILIYYFQESLTGPASKWYT 242
++P +K+P F K+ G S EH+ Y A ++ + + YF SLT A W+T
Sbjct: 159 ELPRGWKVPKFTKFSGDSGESTVEHIARYQIEAGDLAINENLKMKYFPSSLTKNAFTWFT 338
Query: 243 NLDKTRVQTFRDLCEAFVEQYSYNVDMTPDRSDLQAMTQGDKETFKEYAQRWRDTAAQVS 302
L V T+ L F EQ+ + + DL ++ + E+ +Y R+R ++
Sbjct: 339 TLAPRSVHTWAQLERIFHEQF-FRGECKVSXKDLASVKRKPAESIDDYLNRFRMLKSRCF 515
Query: 303 PRIEEKEMTKL 313
+ E E+ L
Sbjct: 516 THVSEHELVVL 548
>AV427422
Length = 417
Score = 53.1 bits (126), Expect = 5e-07
Identities = 32/133 (24%), Positives = 60/133 (45%), Gaps = 2/133 (1%)
Frame = +1
Query: 2092 SNGHRFILVAIDYFTKWVEAASYAN-VTKQVVVKFIKNHIICRYGIPNRIITDNGTNLNN 2150
S G+ +LV +D +K+ + T +V+ ++ +G+P I++D +
Sbjct: 19 SKGYEAVLVVVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDPLFMS 198
Query: 2151 KMMKELCDDFKIEHHNSSPYRPQMNGAVEAANKNIKRIVQKMVVTY-KDWHEMLPFALHG 2209
KEL + S+ Y P+ +G E N+ ++ ++ + K W +P+A +
Sbjct: 199 NFWKELFKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWVPWAEYW 378
Query: 2210 YRTSVRTSIGATP 2222
Y TS S G TP
Sbjct: 379 YNTSYHVSTGQTP 417
>TC15341 similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich protein,
partial (33%)
Length = 1002
Score = 50.1 bits (118), Expect = 4e-06
Identities = 42/132 (31%), Positives = 54/132 (40%), Gaps = 2/132 (1%)
Frame = +1
Query: 357 ATPANGTKKTGNHFPRKKEQEVGMVTHGMPQQTY--PAYQHIAAITPTSHPFQQTNNHSQ 414
+TP G +T P+ + + P Q Y P Q ++ + P Q N S
Sbjct: 676 STPLPGNTQTVAQLPQNQ--------YLPPDQQYRTPQMQDMSRVAP-----QPCTNXSG 816
Query: 415 IPQYPQIPQYPQIPQYPQFPQNPSPQNTQQQNFQQQPYQQFPYQQYPQQNFQQQPYQQRP 474
P + QYPQ Q QQQ QQQ QQ+P QQ PQQ QP +
Sbjct: 817 XSPPPPVQQYPQY------------QQPQQQQQQQQQQQQWP-QQLPQQVQPTQPPSMQS 957
Query: 475 QQPRPPRMPINP 486
Q RPP P+ P
Sbjct: 958 PQIRPPSSPVYP 993
Score = 42.4 bits (98), Expect = 8e-04
Identities = 44/169 (26%), Positives = 64/169 (37%), Gaps = 27/169 (15%)
Frame = +1
Query: 355 KNATPANGTKKTGNHFPRKKEQEVGMVTHGMPQQTYPAYQHIAAITPTSHPFQQTNNHSQ 414
++ +P+ K ++ Q++ + +P Q P Q AA P N +Q
Sbjct: 490 EDKSPSTTDSKKADNASDASNQQLALA---LPNQIAPQPQPAAA-----QPQAPAPNVTQ 645
Query: 415 IPQYPQ--IPQYP---------QIPQYPQFPQNPSPQNTQQQNFQQ-------------- 449
PQ P IP P Q+PQ P + + Q Q+ +
Sbjct: 646 APQQPAYYIPSTPLPGNTQTVAQLPQNQYLPPDQQYRTPQMQDMSRVAPQPCTNXSGXSP 825
Query: 450 -QPYQQFP-YQQYPQQNFQQQPYQQRPQQPRPPRMPINPIPVTYAELLP 496
P QQ+P YQQ QQ QQQ QQ PQQ P P + ++ P
Sbjct: 826 PPPVQQYPQYQQPQQQQQQQQQQQQWPQQLPQQVQPTQPPSMQSPQIRP 972
Score = 32.7 bits (73), Expect = 0.67
Identities = 25/68 (36%), Positives = 27/68 (38%)
Frame = +1
Query: 386 PQQTYPAYQHIAAITPTSHPFQQTNNHSQIPQYPQIPQYPQIPQYPQFPQNPSPQNTQQQ 445
P Q YP YQ P QQ Q Q+PQ Q PQ Q Q P SPQ
Sbjct: 832 PVQQYPQYQQ---------PQQQQQQQQQQQQWPQ--QLPQQVQPTQPPSMQSPQIRPPS 978
Query: 446 NFQQQPYQ 453
+ PYQ
Sbjct: 979 SPVYPPYQ 1002
>TC18149 weakly similar to UP|Q91BL3 (Q91BL3) Essential structural protein
pp78/81, partial (5%)
Length = 710
Score = 48.5 bits (114), Expect = 1e-05
Identities = 49/149 (32%), Positives = 65/149 (42%), Gaps = 17/149 (11%)
Frame = +3
Query: 387 QQTYPAY----QHIAAITPTSHPFQQTNNHSQIPQYPQIPQYPQ--IPQYPQ----FPQN 436
QQ +P+ Q I + P + P Q P +P Q+ Q IP PQ FP
Sbjct: 66 QQQFPSPVNLPQSIPVVPPPNAPPQLPPQQGLPPSFPPPNQFSQTQIPAVPQRDPYFPPP 245
Query: 437 PSPQNTQQQNFQQ----QPYQQF---PYQQYPQQNFQQQPYQQRPQQPRPPRMPINPIPV 489
Q T Q +QQ QP+ Q P+QQY QQ Q Q P Q +PP INP P
Sbjct: 246 VQSQETPNQQYQQPLAPQPHPQPGAPPHQQY-QQTPHPQYSQPAPPQHQPPIPSINP-PQ 419
Query: 490 TYAELLPGLLKKNLVQTRTAPPIPEKLPS 518
+ L + + V ++T PP + PS
Sbjct: 420 LQSSLGHHVEEPPYVPSQTYPPNLRQPPS 506
>BF177840
Length = 410
Score = 45.8 bits (107), Expect = 8e-05
Identities = 30/129 (23%), Positives = 62/129 (47%), Gaps = 3/129 (2%)
Frame = +2
Query: 2140 IITDNGTNLNNKMMKELCDDFKIEHHNSSPYRPQMNGAVEAANKNIKRIVQKMVV-TYKD 2198
I++D T + + L + S+ PQ +G E NK + +++ ++ K
Sbjct: 17 IVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNLKM 196
Query: 2199 WHEMLPFALHGYRTSVRTSIGATPFSLVYGMEAVLPVE-VEIPSLRVLMEVD-LSEAEWV 2256
W LP Y V ++ +PF +VYG + P++ + +P+ +L D ++AE+V
Sbjct: 197 WETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDLLPMPNTYLLKHKDGKAKAEFV 376
Query: 2257 QNRYDQLNL 2265
+ ++++ L
Sbjct: 377 KRLHEKIKL 403
>TC19740 similar to UP|FKH_DROME (P14734) Fork head protein, partial (4%)
Length = 622
Score = 41.2 bits (95), Expect = 0.002
Identities = 29/81 (35%), Positives = 35/81 (42%)
Frame = +3
Query: 406 FQQTNNHSQIPQYPQIPQYPQIPQYPQFPQNPSPQNTQQQNFQQQPYQQFPYQQYPQQNF 465
FQ + S + I P PQ Q+ Q+ Q QQQP Q F +QQ P F
Sbjct: 39 FQI*TSSSSFLHFTPIAMMYNDPHNPQHQQHQHQQHQHQFQQQQQP-QHFHHQQ-PGPEF 212
Query: 466 QQQPYQQRPQQPRPPRMPINP 486
+ P PQQP PP M P
Sbjct: 213 HRGPPPPPPQQPHPPPMMRQP 275
>AU251673
Length = 413
Score = 41.2 bits (95), Expect = 0.002
Identities = 26/119 (21%), Positives = 53/119 (43%), Gaps = 5/119 (4%)
Frame = +2
Query: 233 LTGPASKWYTNLDKTRVQ-----TFRDLCEAFVEQYSYNVDMTPDRSDLQAMTQGDKETF 287
L G A WY L + + T+ D F+ ++ D + + Q + T
Sbjct: 44 LEGVARDWYNVLTRAKPVGSPPWTWADFSAEFMNRFLPQSVRDGFVRDFERLEQAEGMTV 223
Query: 288 KEYAQRWRDTAAQVSPRIEEKEMTKLFLKTLNHFYYKKMVGSTPKSFAEMVGMGVQLEE 346
EY+ + + V + E+E K F++ L + +K +VGS + +E++ + + +E+
Sbjct: 224 SEYSAHFTHLSRYVPYPLLEEERVKRFVRGLKEYLFKSVVGSKSSTLSEVLSLALLVEQ 400
>BP043633
Length = 542
Score = 40.0 bits (92), Expect = 0.004
Identities = 31/94 (32%), Positives = 36/94 (37%), Gaps = 6/94 (6%)
Frame = +1
Query: 401 PTSHPFQQTNNHSQIPQYPQIPQYPQIPQY-PQFP-----QNPSPQNTQQQNFQQQPYQQ 454
P P N+H PQ P P PQ P Y PQF Q PSP P
Sbjct: 256 PPPPPPPDPNHHHHFPQIPP-PHPPQAPWYSPQFQYHHPSQTPSPPPQWGPPPPPPPPPS 432
Query: 455 FPYQQYPQQNFQQQPYQQRPQQPRPPRMPINPIP 488
PY +P+Q P + P P P P + IP
Sbjct: 433 NPYSYHPKQFPPPPPPPRPPHPPHPQFPPHSHIP 534
>TC15540 similar to UP|Q9NGS5 (Q9NGS5) Prespore protein MF12 (Fragment),
partial (6%)
Length = 806
Score = 40.0 bits (92), Expect = 0.004
Identities = 32/73 (43%), Positives = 34/73 (45%), Gaps = 3/73 (4%)
Frame = +2
Query: 407 QQTNNHSQIPQYPQIPQYPQIPQYPQFPQNPSPQNTQQQ--NFQQQPYQQF-PYQQYPQQ 463
QQ Q PQ Q+ Q Q Q PQ Q Q QQQ QQQ Q P QQ PQ
Sbjct: 23 QQQQLQQQPPQLTQLQQQQQ-QQLPQVQQQQLSQMQQQQLPQLQQQQLSQLQPQQQLPQ- 196
Query: 464 NFQQQPYQQRPQQ 476
QQ +QQ PQQ
Sbjct: 197 -LQQLQHQQLPQQ 232
Score = 40.0 bits (92), Expect = 0.004
Identities = 29/66 (43%), Positives = 32/66 (47%), Gaps = 3/66 (4%)
Frame = +2
Query: 417 QYPQIPQYPQIPQYPQFPQNPSPQNTQQQNF---QQQPYQQFPYQQYPQQNFQQQPYQQR 473
Q PQ+ Q Q+ Q P PQ Q QQQ QQQ Q QQ PQ QQQ Q +
Sbjct: 5 QLPQLQQQQQLQQQP--PQLTQLQQQQQQQLPQVQQQQLSQMQQQQLPQLQ-QQQLSQLQ 175
Query: 474 PQQPRP 479
PQQ P
Sbjct: 176 PQQQLP 193
Score = 38.9 bits (89), Expect = 0.009
Identities = 28/63 (44%), Positives = 31/63 (48%), Gaps = 1/63 (1%)
Frame = +2
Query: 414 QIPQYPQIPQYP-QIPQYPQFPQNPSPQNTQQQNFQQQPYQQFPYQQYPQQNFQQQPYQQ 472
Q+ Q Q+ Q P Q+ Q Q Q PQ QQQ Q QQ P Q QQ Q QP QQ
Sbjct: 14 QLQQQQQLQQQPPQLTQLQQQQQQQLPQ-VQQQQLSQMQQQQLP-QLQQQQLSQLQPQQQ 187
Query: 473 RPQ 475
PQ
Sbjct: 188 LPQ 196
>TC8445 similar to UP|O80910 (O80910) At2g38410 protein, partial (7%)
Length = 841
Score = 38.5 bits (88), Expect = 0.012
Identities = 28/78 (35%), Positives = 33/78 (41%), Gaps = 7/78 (8%)
Frame = +1
Query: 408 QTNNHSQIPQYPQIPQYPQIPQY-PQFPQNPSPQNTQQ------QNFQQQPYQQFPYQQY 460
Q +N Q QY P PQ+ P F +P PQ Q Q Q QP Q + Q
Sbjct: 10 QLHNEPQPLQYQPQPHLQHQPQHHPHFQPHPQPQLQSQPQPQHIQQSQPQPQSQPQHIQQ 189
Query: 461 PQQNFQQQPYQQRPQQPR 478
PQ Q Q QQ QP+
Sbjct: 190 PQPQPQLQHIQQSQPQPQ 243
Score = 32.3 bits (72), Expect = 0.88
Identities = 27/112 (24%), Positives = 34/112 (30%)
Frame = +1
Query: 369 HFPRKKEQEVGMVTHGMPQQTYPAYQHIAAITPTSHPFQQTNNHSQIPQYPQIPQYPQIP 428
H + + Q H Q P QHI P Q N H Q P P + P
Sbjct: 136 HIQQSQPQPQSQPQHIQQPQPQPQLQHIQQSQPQP---QFQNQHFQYPGTYPPPPWAATP 306
Query: 429 QYPQFPQNPSPQNTQQQNFQQQPYQQFPYQQYPQQNFQQQPYQQRPQQPRPP 480
YP + + S N PY + Q P Q+P P
Sbjct: 307 GYPNYQNHLSSTNALSTPQANTPYTSAVRGDPVASSGQSNPPNATGQRPFVP 462
Score = 31.2 bits (69), Expect = 2.0
Identities = 25/90 (27%), Positives = 36/90 (39%), Gaps = 4/90 (4%)
Frame = +1
Query: 428 PQYPQFPQNPSPQNTQQQNFQQQPYQQFPYQQYPQQNFQQQPYQQRPQQPRPPRMPINPI 487
PQ Q+ P Q+ Q + QP+ Q Q PQ QQ Q QP+ + P P
Sbjct: 25 PQPLQYQPQPHLQHQPQHHPHFQPHPQPQLQSQPQPQHIQQSQPQPQSQPQHIQQP-QPQ 201
Query: 488 P----VTYAELLPGLLKKNLVQTRTAPPIP 513
P + ++ P ++ T PP P
Sbjct: 202 PQLQHIQQSQPQPQFQNQHFQYPGTYPPPP 291
>TC14545 similar to UP|Q9FT78 (Q9FT78) P23 co-chaperone, partial (46%)
Length = 1127
Score = 38.1 bits (87), Expect = 0.016
Identities = 25/93 (26%), Positives = 39/93 (41%), Gaps = 2/93 (2%)
Frame = -3
Query: 401 PTSHPFQQTNNHS--QIPQYPQIPQYPQIPQYPQFPQNPSPQNTQQQNFQQQPYQQFPYQ 458
P + P +H + P P +P P +P +P P +PS Q Q + + P+ Q
Sbjct: 663 PQNRPLHHHPSHQI*RNPFLPILPFLPILPFHPSLPFHPSLQVLQDLHHRPYPFHQI--- 493
Query: 459 QYPQQNFQQQPYQQRPQQPRPPRMPINPIPVTY 491
PYQ + Q P PI P P+++
Sbjct: 492 ----SRSPSLPYQVQQQHLLHPHPPIYPNPLSH 406
>AW719943
Length = 508
Score = 37.7 bits (86), Expect = 0.021
Identities = 21/42 (50%), Positives = 25/42 (59%), Gaps = 3/42 (7%)
Frame = +2
Query: 438 SPQNTQQQNFQQQ---PYQQFPYQQYPQQNFQQQPYQQRPQQ 476
+PQ QQ QQQ P QQ P+QQ N QQQ +QQ+ QQ
Sbjct: 80 TPQQQQQPQQQQQLFQPQQQSPFQQNSLFNSQQQQFQQQQQQ 205
Score = 35.8 bits (81), Expect = 0.079
Identities = 20/45 (44%), Positives = 22/45 (48%)
Frame = +2
Query: 431 PQFPQNPSPQNTQQQNFQQQPYQQFPYQQYPQQNFQQQPYQQRPQ 475
PQ Q P Q Q QQ P+QQ QQ FQQQ QQ+ Q
Sbjct: 83 PQQQQQPQQQQQLFQPQQQSPFQQNSLFNSQQQQFQQQQQQQQQQ 217
Score = 30.8 bits (68), Expect = 2.6
Identities = 20/42 (47%), Positives = 21/42 (49%), Gaps = 6/42 (14%)
Frame = +2
Query: 419 PQIPQYPQIPQYPQFPQNPSPQ------NTQQQNFQQQPYQQ 454
PQ Q PQ Q PQ SP N+QQQ FQQQ QQ
Sbjct: 83 PQQQQQPQQQQQLFQPQQQSPFQQNSLFNSQQQQFQQQQQQQ 208
>TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, partial (22%)
Length = 505
Score = 37.0 bits (84), Expect = 0.036
Identities = 36/138 (26%), Positives = 56/138 (40%)
Frame = +1
Query: 398 AITPTSHPFQQTNNHSQIPQYPQIPQYPQIPQYPQFPQNPSPQNTQQQNFQQQPYQQFPY 457
A T H +HS++P +P+ +P P P P P + P P
Sbjct: 109 ATTSHHHFLPP*RHHSRLP-HPRPTLFPPFPLPRLPPLVPPPPLPRLLRPPPPPPPPPPL 285
Query: 458 QQYPQQNFQQQPYQQRPQQPRPPRMPINPIPVTYAELLPGLLKKNLVQTRTAPPIPEKLP 517
+ P + ++ RP PPR + P+P LP LL++ L + PP+P + P
Sbjct: 286 RPRPHR------HRLRPLLRHPPRRQMEPLP------LPPLLQRRLPRDLPLPPLP-RAP 426
Query: 518 SWYRLDQTCDFHEGGRGH 535
++R FH R H
Sbjct: 427 RFHRPQ---GFHRRKRRH 471
>TC15784 similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precursor (Atperox
P31) (ATP41) , partial (38%)
Length = 475
Score = 35.8 bits (81), Expect = 0.079
Identities = 24/91 (26%), Positives = 38/91 (41%), Gaps = 13/91 (14%)
Frame = +1
Query: 34 AVTEAQARSQAPPPPPPVRTQAEASSSWTLCADTPTRSAPQ-------------RSAPWF 80
+ T + PPPP P R+ + ++++ T A TP S+PQ +P
Sbjct: 157 SATPSPTNKSPPPPPAPPRSASSSTTASTPTAATPPSSSPQLPSTKPNATPTSTSPSPAT 336
Query: 81 PPFTAGEIFRPITCEAQMPTHQYTGQVPLPA 111
P ++ E+ P A P+ T P PA
Sbjct: 337 PSTSSSELKPPSNSLAPTPSPAPTSSPPPPA 429
Score = 32.7 bits (73), Expect = 0.67
Identities = 21/56 (37%), Positives = 26/56 (45%)
Frame = +1
Query: 26 AQAQTQAQAVTEAQARSQAPPPPPPVRTQAEASSSWTLCADTPTRSAPQRSAPWFP 81
A ++ A +S PPP PP ASSS T A TPT + P S+P P
Sbjct: 136 AHNSPKSSATPSPTNKSPPPPPAPP----RSASSSTT--ASTPTAATPPSSSPQLP 285
>CN825698
Length = 704
Score = 35.4 bits (80), Expect = 0.10
Identities = 17/40 (42%), Positives = 22/40 (54%), Gaps = 2/40 (5%)
Frame = +3
Query: 450 QPYQQFPYQQYPQQNFQQ--QPYQQRPQQPRPPRMPINPI 487
+P QQ P Q P Q Q QP P +PRPP +P +P+
Sbjct: 273 RPQQQLPPAQRPGQGLQAPPQPLTSFPTRPRPPSLPSSPL 392
>TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transcription
factor PosF21 (AtbZIP59), partial (17%)
Length = 570
Score = 35.0 bits (79), Expect = 0.14
Identities = 24/53 (45%), Positives = 28/53 (52%)
Frame = +1
Query: 423 QYPQIPQYPQFPQNPSPQNTQQQNFQQQPYQQFPYQQYPQQNFQQQPYQQRPQ 475
Q+ Q+ +PQ Q Q QQ QQQ QQ P QQ PQQ QQQ +Q Q
Sbjct: 88 QFQQLQIHPQKHQQHQHQ-FQQPIQQQQQQQQQPIQQ-PQQQMQQQQQEQHQQ 240
Score = 34.7 bits (78), Expect = 0.18
Identities = 19/48 (39%), Positives = 23/48 (47%)
Frame = +1
Query: 429 QYPQFPQNPSPQNTQQQNFQQQPYQQFPYQQYPQQNFQQQPYQQRPQQ 476
Q+ Q +P Q FQQ QQ QQ P Q QQQ QQ+ +Q
Sbjct: 88 QFQQLQIHPQKHQQHQHQFQQPIQQQQQQQQQPIQQPQQQMQQQQQEQ 231
Score = 32.7 bits (73), Expect = 0.67
Identities = 31/89 (34%), Positives = 36/89 (39%)
Frame = +1
Query: 384 GMPQQTYPAYQHIAAITPTSHPFQQTNNHSQIPQYPQIPQYPQIPQYPQFPQNPSPQNTQ 443
G QQ YP + H + FQQ H PQ Q Q+ Q Q
Sbjct: 28 GGGQQFYP-HNHSMHTLLAAQQFQQLQIH---------------PQKHQQHQHQFQQPIQ 159
Query: 444 QQNFQQQPYQQFPYQQYPQQNFQQQPYQQ 472
QQ QQQ Q P QQ QQ QQ+ +QQ
Sbjct: 160 QQQQQQQQPIQQPQQQMQQQ--QQEQHQQ 240
>TC15704 similar to UP|O22265 (O22265) Expressed protein
(At2g47450/T30B22.25), partial (25%)
Length = 664
Score = 35.0 bits (79), Expect = 0.14
Identities = 24/84 (28%), Positives = 31/84 (36%)
Frame = -1
Query: 405 PFQQTNNHSQIPQYPQIPQYPQIPQYPQFPQNPSPQNTQQQNFQQQPYQQFPYQQYPQQN 464
P T+ H Q+ P P P+ P P+NPSP Q + + P N
Sbjct: 310 PTAPTSPHPQLSGQPPPPP----PRTPAPPRNPSPDPPPQTSHSPTRFHHRPSTARDTPN 143
Query: 465 FQQQPYQQRPQQPRPPRMPINPIP 488
P P P PPR P+P
Sbjct: 142 --SHPSPSAPISPPPPRTRTPPLP 77
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 41,165,100
Number of Sequences: 28460
Number of extensions: 624593
Number of successful extensions: 6784
Number of sequences better than 10.0: 150
Number of HSP's better than 10.0 without gapping: 5252
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 6233
length of query: 2360
length of database: 4,897,600
effective HSP length: 105
effective length of query: 2255
effective length of database: 1,909,300
effective search space: 4305471500
effective search space used: 4305471500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146971.7


