

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146941.4 + phase: 0
(328 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
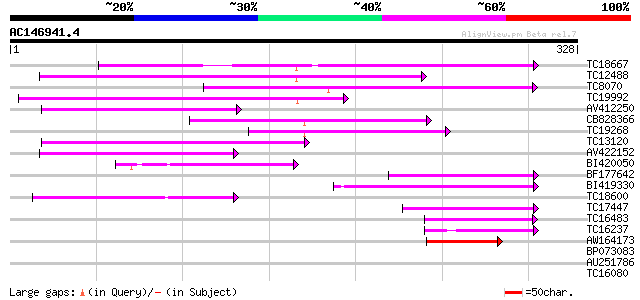
Score E
Sequences producing significant alignments: (bits) Value
TC18667 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein,... 135 9e-33
TC12488 similar to UP|Q9LQZ3 (Q9LQZ3) F10A5.28, partial (47%) 107 3e-24
TC8070 similar to UP|Q9LQZ3 (Q9LQZ3) F10A5.28, partial (73%) 107 4e-24
TC19992 similar to UP|Q8LE45 (Q8LE45) Nodulin-like protein, part... 89 8e-19
AV412250 80 5e-16
CB828366 80 6e-16
TC19268 similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein, partia... 66 7e-12
TC13120 weakly similar to UP|Q8LRB6 (Q8LRB6) Nodulin-like protei... 64 3e-11
AV422152 62 2e-10
BI420050 61 3e-10
BF177642 58 2e-09
BI419330 56 7e-09
TC18600 similar to UP|Q94JU2 (Q94JU2) AT3g28050/MMG15_6, partial... 50 5e-07
TC17447 weakly similar to UP|Q8L9I2 (Q8L9I2) Nodulin MtN21-like ... 48 3e-06
TC16483 homologue to UP|O24091 (O24091) MtN21 protein, partial (... 47 3e-06
TC16237 weakly similar to UP|Q84TV1 (Q84TV1) Nodulin-like protei... 47 4e-06
AW164173 43 6e-05
BP073083 37 0.003
AU251786 35 0.017
TC16080 similar to UP|Q8S0P2 (Q8S0P2) Ribosomal protein L28-like... 30 0.42
>TC18667 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein, partial
(27%)
Length = 911
Score = 135 bits (340), Expect = 9e-33
Identities = 84/260 (32%), Positives = 125/260 (47%), Gaps = 5/260 (1%)
Frame = +3
Query: 52 LVAAVCVAPLAIYFERGQPKNFSCEVLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNF 111
L ++ + PLA + ER + +VL + FL G G ++ L+ + ATY
Sbjct: 6 LFSSAFMIPLAYFAERKSKPKITMKVLFQAFLCGLFGATIQENLFVEAVTWAGATYPSAM 185
Query: 112 LNLIPICTFLTAILFRMENLNIHTWNGRAKCVGAILCVAGTLAARLYKGKEFY----IAQ 167
L E LNI T G+AK VG ++ ++G + YK E + I
Sbjct: 186 L----------------ERLNIETKTGKAKVVGTLIGISGAMILTFYKSIEIHLWPTIVN 317
Query: 168 YHSFHSVAAHKTQMLRGTLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQS 227
VA T + GT G C SYS W +Q ++ FP Y L VMA IQS
Sbjct: 318 QSKPKDVA---TGHVWGTSLAFGTCISYSIWLIIQARMSAKFPWHYTSAALMSVMACIQS 488
Query: 228 AVIGACVDSSK-EAWRLEWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLA 286
+ ++ W+L WN++L+T LY+G +++ V+ L +W + +KGP Y S FNPL
Sbjct: 489 TIFALGMERDNWSQWKLGWNIELLTALYTGVVASGVVWVLMAWCLRLKGPLYASAFNPLL 668
Query: 287 LVFVAFAEAMILGEPLTVGT 306
LV VA A ++ L E L +G+
Sbjct: 669 LVIVAIAGSLFLEEKLYLGS 728
>TC12488 similar to UP|Q9LQZ3 (Q9LQZ3) F10A5.28, partial (47%)
Length = 887
Score = 107 bits (267), Expect = 3e-24
Identities = 66/243 (27%), Positives = 107/243 (43%), Gaps = 19/243 (7%)
Frame = +1
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+M+ +Q G ++SR L G YR ++A + + P A + E+ + +
Sbjct: 154 AMLALQFGYAGFHVVSRAALNMGVSKLVFPVYRNIIALLLLLPFAYFLEKKERPAITLNF 333
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
L + F VG++ Y G+ +TS T+A N +P TFL A + R+E + ++ +
Sbjct: 334 LVQFFFLALVGITANQAFYLLGLDNTSPTFASAIQNSVPAITFLMAAILRIEQVRLNRKD 513
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFY------------------IAQYHSFHSVA-AHK 178
G AK G + CV G LYKG Y I + SF S+ A+
Sbjct: 514 GIAKIAGTVFCVVGASVITLYKGPTIYNPSPAVPHVNSSSITAPVIEESWSFSSLGDANG 693
Query: 179 TQMLRGTLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSK 238
G ++LIG C S+SAW +Q +++ +P R C +Q VI +
Sbjct: 694 KNWTLGCVYLIGHCLSWSAWLVLQAPVLKKYPARLSVTSYTCFFGLLQFLVIALVAERDA 873
Query: 239 EAW 241
+AW
Sbjct: 874 QAW 882
>TC8070 similar to UP|Q9LQZ3 (Q9LQZ3) F10A5.28, partial (73%)
Length = 1758
Score = 107 bits (266), Expect = 4e-24
Identities = 61/210 (29%), Positives = 98/210 (46%), Gaps = 17/210 (8%)
Frame = +1
Query: 113 NLIPICTFLTAILFRMENLNIHTWNGRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFH 172
N +P TFL A + R+E + ++ +G AK G + CV G LYKG Y
Sbjct: 418 NSVPAITFLMAAILRIEQVRLNRKDGIAKVAGTVFCVIGASVITLYKGPTIYSPVPPLNS 597
Query: 173 SVAAH-KTQMLR----------------GTLFLIGACFSYSAWFFMQVKLVEVFPLRYWG 215
+ H +TQ+L G L+LIG C S+S W +Q +++ +P R
Sbjct: 598 IITPHPQTQLLGSVSLSLGDANGKNWTLGCLYLIGHCLSWSGWLVLQAPILKKYPARLSV 777
Query: 216 IMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLITILYSGALSTAAVFCLQSWAMTIKG 275
C IQ VI V+ +AW ++ TILY+G +++ F +Q W + G
Sbjct: 778 TSYTCFFGLIQFLVIALIVERDAQAWIFHSGGEVFTILYAGVVASGIAFAVQIWCIDRGG 957
Query: 276 PTYPSMFNPLALVFVAFAEAMILGEPLTVG 305
P + +++ P+ + VA ++ LGE +G
Sbjct: 958 PVFVAVYQPVQTLVVAIMASLALGEEFYLG 1047
>TC19992 similar to UP|Q8LE45 (Q8LE45) Nodulin-like protein, partial (46%)
Length = 615
Score = 89.4 bits (220), Expect = 8e-19
Identities = 52/197 (26%), Positives = 96/197 (48%), Gaps = 6/197 (3%)
Frame = +2
Query: 6 VKEWFVLSSILWSMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYF 65
V+ F+ S + ++F+Q G +L + L G + L YR VA V +AP A+
Sbjct: 20 VQNGFIKPSHVIGVVFIQSGYAGMDVLCKAALNNGMSNYVLVVYRHAVAFVVMAPFALIL 199
Query: 66 ERGQPKNFSCEVLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAIL 125
++ + + KI + + LYY G++ T+AT+A++ N++P TFL A +
Sbjct: 200 DKKIRPKMTFSIFVKIVALSLLEPVIDQNLYYLGMKYTTATFAVSMYNILPAITFLMAYI 379
Query: 126 FRMENLNIHTWNGRAKCVGAILCVAGTLAARLYKGKEFYI------AQYHSFHSVAAHKT 179
++E + I + +AK G + VAG + L KG E + + H H+ +
Sbjct: 380 LKLEKIKIKSIRSQAKVFGTLATVAGAMVMTLIKGPELELFGTHGGSSTHIQHNGGVNLQ 559
Query: 180 QMLRGTLFLIGACFSYS 196
++G++ + CFS++
Sbjct: 560 HAIKGSVMITIGCFSFA 610
>AV412250
Length = 403
Score = 80.1 bits (196), Expect = 5e-16
Identities = 43/116 (37%), Positives = 63/116 (54%)
Frame = +3
Query: 19 MIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVL 78
M VQ G IL +I++ +G + +T YR +++A LA FER N + ++L
Sbjct: 27 MFVVQAAFAGNNILYKIVIFDGMPVSIITVYRQIISAAFTLSLAFIFERNNMPNLTFKLL 206
Query: 79 TKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIH 134
+ FLNG +G S+ LY+ + SAT+ NLIP TF AILF +E L+ H
Sbjct: 207 SISFLNGLLGGSLYQNLYFESLSLISATFVTALFNLIPAITFALAILFGLETLDFH 374
>CB828366
Length = 568
Score = 79.7 bits (195), Expect = 6e-16
Identities = 42/144 (29%), Positives = 70/144 (48%), Gaps = 4/144 (2%)
Frame = +3
Query: 105 ATYALNFLNLIPICTFLTAILFRMENLNIHTWNGRAKCVGAILCVAGTLAARLYKGKEFY 164
ATY + LNLIP+ T++ + R+E N+ T G+AK +G + V G + LY+G+ +
Sbjct: 135 ATYTITMLNLIPVVTYIMTVSLRLEKPNLGTRAGKAKLLGTLTGVGGAMILTLYEGRRIF 314
Query: 165 IAQYH----SFHSVAAHKTQMLRGTLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQC 220
H + S L G + +G +S W+ +Q + + FP +Y L
Sbjct: 315 NWPLHIDLSKYMSPPPANGSRLWGLILALGTSLCFSLWYIVQANMSQKFPWQYSVAALTS 494
Query: 221 VMAAIQSAVIGACVDSSKEAWRLE 244
VM+AIQS + C + ++E
Sbjct: 495 VMSAIQSIIYALCTERDWSQCKVE 566
>TC19268 similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein, partial (8%)
Length = 505
Score = 66.2 bits (160), Expect = 7e-12
Identities = 38/125 (30%), Positives = 57/125 (45%), Gaps = 8/125 (6%)
Frame = +2
Query: 139 RAKCVGAILCVAGTLAARLYKGKEFYIAQYH--------SFHSVAAHKTQMLRGTLFLIG 190
+AK VG I + G + L KGKE I +H H A + L G L +
Sbjct: 2 KAKIVGTITGIGGAMVLTLVKGKEIKIESFHVNLLHPXNGTHPHAGGTGENLLGALCSLA 181
Query: 191 ACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLI 250
+ SYS W +Q K+ E + Y L A++ S V ++ WRL WN++L+
Sbjct: 182 SGTSYSLWLIIQRKMTERYDGHYTSTALMSFWASVLSFVFALFIERDWSQWRLGWNIRLL 361
Query: 251 TILYS 255
T+ Y+
Sbjct: 362 TVAYA 376
>TC13120 weakly similar to UP|Q8LRB6 (Q8LRB6) Nodulin-like protein, partial
(22%)
Length = 609
Score = 63.9 bits (154), Expect = 3e-11
Identities = 41/155 (26%), Positives = 71/155 (45%)
Frame = +3
Query: 19 MIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVL 78
++ VQ G I + + +G + YR +AA+ ++P A ER S V
Sbjct: 105 LLAVQFGSAGMFIFAMDAMKKGMSHYVFIVYRNAIAAITLSPFAFVLERKIRPKMSVRVF 284
Query: 79 TKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWNG 138
+I F + + G++ TSA++ +N P TF+ A++ +ME++ I
Sbjct: 285 AEIMALAFFEIILDQCFALLGMKLTSASFLSAVMNAAPSITFVMAVVLKMEHMKIKEVAC 464
Query: 139 RAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFHS 173
+AK +G + GTL LYKG + +FH+
Sbjct: 465 QAKMIGTAVTFGGTLLMALYKGPIVSGVRSSTFHA 569
>AV422152
Length = 465
Score = 61.6 bits (148), Expect = 2e-10
Identities = 32/115 (27%), Positives = 62/115 (53%)
Frame = +2
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+++ +Q +G +++ + G + L+ YR +VA + +AP A ER + +
Sbjct: 119 AIVSLQFGYSGMYVITMVSFKHGMSHWILSVYRHVVALIIIAPFAFVLERKIRPKMTLPI 298
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLN 132
+I GF+ + LY G++ TS T+A +N++P TF+ A++FR+E +N
Sbjct: 299 FLRIVALGFLEPVLDQNLYNMGMKMTSTTFASATVNVLPAITFIMALIFRLETVN 463
>BI420050
Length = 613
Score = 60.8 bits (146), Expect = 3e-10
Identities = 40/109 (36%), Positives = 60/109 (54%), Gaps = 3/109 (2%)
Frame = +3
Query: 62 AIYFERGQ---PKNFSCEVLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPIC 118
A+ ER + P +FS V+ KI L G +G + ++ Y GI +S T + NL+P
Sbjct: 165 AVMAERSRVLPPLSFS--VMCKIGLLGLIGCTS-QIVGYTGISYSSPTLSSAISNLVPAF 335
Query: 119 TFLTAILFRMENLNIHTWNGRAKCVGAILCVAGTLAARLYKGKEFYIAQ 167
TFL A++FRME + I + + +AK +G I+ + G LYKG I Q
Sbjct: 336 TFLFAVIFRMEKVAIKSSSTQAKVLGTIVSITGAFIVTLYKGPPIIILQ 482
>BF177642
Length = 482
Score = 57.8 bits (138), Expect = 2e-09
Identities = 26/87 (29%), Positives = 44/87 (49%)
Frame = +1
Query: 220 CVMAAIQSAVIGACVDSSKEAWRLEWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYP 279
C M + SA++G + + AW L + +L+ + YS + +WA KGP Y
Sbjct: 43 CSMVVVLSAIVGLIAEGTSRAWILRPDKELVAVCYSAIFVVTMRSVVYAWAFRKKGPIYV 222
Query: 280 SMFNPLALVFVAFAEAMILGEPLTVGT 306
+MF+PL +V + LG+ L +G+
Sbjct: 223 AMFSPLGMVIAVVLGVLFLGDILYLGS 303
>BI419330
Length = 470
Score = 56.2 bits (134), Expect = 7e-09
Identities = 33/120 (27%), Positives = 60/120 (49%), Gaps = 1/120 (0%)
Frame = +1
Query: 188 LIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDS-SKEAWRLEWN 246
LIG +SA++ +Q + +P C + A+QS+ + ++ AW L +
Sbjct: 4 LIGGA-GFSAFYILQAITLRKYPAEMSLATWICFVGALQSSALTIFMEHHDPAAWSLGLD 180
Query: 247 LQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
+L YSG +++A F +Q + KGP + + FNPL ++ V ++L E L +G+
Sbjct: 181 SKLFACAYSGIVTSAVQFYVQGTVIKTKGPVFVTAFNPLRMIIVTALACILLSEKLYLGS 360
>TC18600 similar to UP|Q94JU2 (Q94JU2) AT3g28050/MMG15_6, partial (19%)
Length = 438
Score = 50.1 bits (118), Expect = 5e-07
Identities = 35/120 (29%), Positives = 57/120 (47%), Gaps = 1/120 (0%)
Frame = +3
Query: 14 SILWSMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQ-PKN 72
S+ +M+ Q + G L + G + Y L+A + P I + R + P
Sbjct: 81 SVTVAMVTTQFLDVGLNTLVKSATNSGMSNYVFVVYSNLLALCFLLPSTILYHRKRAPPP 260
Query: 73 FSCEVLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLN 132
+L ++FL + + V L Y GI +S T A ++L+P TF+ A++ RMENLN
Sbjct: 261 IPSSILCRMFLISCLSTA-VQTLVYTGIGYSSPTLASAAVDLVPAFTFILAVISRMENLN 437
>TC17447 weakly similar to UP|Q8L9I2 (Q8L9I2) Nodulin MtN21-like protein,
partial (24%)
Length = 590
Score = 47.8 bits (112), Expect = 3e-06
Identities = 22/79 (27%), Positives = 39/79 (48%)
Frame = +3
Query: 228 AVIGACVDSSKEAWRLEWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLAL 287
AV+G ++ W++ N L +IL SG + + + + +KG Y +MF PL++
Sbjct: 3 AVVGVITETKASVWKIGLNTALASILCSGIFGSILNLAVHTHMLHLKGAVYVAMFKPLSI 182
Query: 288 VFVAFAEAMILGEPLTVGT 306
M LG+ L +G+
Sbjct: 183 AIAVVLGVMFLGDTLHLGS 239
>TC16483 homologue to UP|O24091 (O24091) MtN21 protein, partial (24%)
Length = 739
Score = 47.4 bits (111), Expect = 3e-06
Identities = 18/65 (27%), Positives = 38/65 (57%)
Frame = +1
Query: 241 WRLEWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGE 300
W + W++ L+ Y+G ++++ + +Q + KGP + + F+PL ++ VA + IL E
Sbjct: 1 WTIGWDMNLLAAAYAGIVTSSISYYVQGLVIRKKGPVFATAFSPLMMIIVAIMGSFILAE 180
Query: 301 PLTVG 305
+ +G
Sbjct: 181 QIFLG 195
>TC16237 weakly similar to UP|Q84TV1 (Q84TV1) Nodulin-like protein, partial
(11%)
Length = 695
Score = 47.0 bits (110), Expect = 4e-06
Identities = 24/66 (36%), Positives = 38/66 (57%)
Frame = +3
Query: 241 WRLEWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGE 300
W ++++LQ + + S + L +W + +KGP Y S FNPL LV VA A ++ L E
Sbjct: 12 WNIKFSLQFMWVC-----SFWNTWVLTAWVLRLKGPLYASSFNPLFLVIVAIAGSLFLDE 176
Query: 301 PLTVGT 306
L +G+
Sbjct: 177 KLYLGS 194
>AW164173
Length = 377
Score = 43.1 bits (100), Expect = 6e-05
Identities = 19/44 (43%), Positives = 28/44 (63%)
Frame = +1
Query: 242 RLEWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPL 285
R+ ++ LI+ILYSG ST + +W + +KGP Y S+F PL
Sbjct: 13 RINPDVTLISILYSGFFSTGLSCLVHTWGLRLKGPVYISLFKPL 144
>BP073083
Length = 352
Score = 37.4 bits (85), Expect = 0.003
Identities = 17/51 (33%), Positives = 29/51 (56%)
Frame = -2
Query: 256 GALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
G +++ V+ L +W + +KGP S F+PL LV VA ++ L +G+
Sbjct: 348 GGVASGVVWVLMAWCLRLKGPL*ASAFHPLLLVLVALPGSLFFEGKLYLGS 196
>AU251786
Length = 349
Score = 35.0 bits (79), Expect = 0.017
Identities = 28/114 (24%), Positives = 46/114 (39%), Gaps = 10/114 (8%)
Frame = +1
Query: 127 RMENLNIHTWNGRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFHSVAAHKTQMLRGT- 185
RME L + +AK +G+I+ +AG YKG I + S+ + + L+
Sbjct: 10 RMEKLAAKRRSSQAKVLGSIISIAGAFELTFYKGPS-VINGHTRLTSLLDQRFRFLKSDS 186
Query: 186 ---------LFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVI 230
+ L F S W+ +QV ++ FP + V A I S +
Sbjct: 187 EDAIWAIAGILLTADYFLASLWYILQVHILNEFPDELTIVFFYNVTATIISVTV 348
>TC16080 similar to UP|Q8S0P2 (Q8S0P2) Ribosomal protein L28-like, complete
Length = 776
Score = 30.4 bits (67), Expect = 0.42
Identities = 11/22 (50%), Positives = 19/22 (86%)
Frame = +2
Query: 13 SSILWSMIFVQLIVTGTQILSR 34
S++LWSMI++QLI+ Q++S+
Sbjct: 680 STVLWSMIYIQLIMRSWQLVSK 745
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.332 0.141 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,562,217
Number of Sequences: 28460
Number of extensions: 94727
Number of successful extensions: 672
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 667
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 668
length of query: 328
length of database: 4,897,600
effective HSP length: 91
effective length of query: 237
effective length of database: 2,307,740
effective search space: 546934380
effective search space used: 546934380
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146941.4


