

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
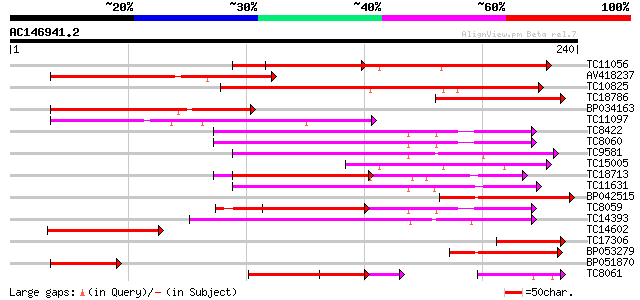
Query= AC146941.2 - phase: 0 /pseudo
(240 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC11056 similar to UP|O81390 (O81390) Calcium-dependent protein ... 117 1e-27
AV418237 107 2e-24
TC10825 similar to GB|AAP68339.1|31711966|BT008900 At1g74740 {Ar... 106 3e-24
TC18786 similar to UP|Q42479 (Q42479) Calcium-dependent protein ... 82 1e-16
BP034163 75 1e-14
TC11097 similar to UP|P93520 (P93520) Calcium/calmodulin-depende... 72 6e-14
TC8422 UP|AAQ63461 (AAQ63461) Calmodulin 4 (Fragment), complete 69 5e-13
TC8060 UP|O49184 (O49184) Calmodulin (ESTS AU081349) (Auxin-regu... 69 9e-13
TC9581 UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, complete 67 2e-12
TC15005 homologue to UP|Q9AR92 (Q9AR92) Protein kinase , partia... 65 1e-11
TC18713 similar to GB|AAL90989.1|19699246|AY090328 AT3g03000/F13... 64 3e-11
TC11631 weakly similar to UP|TCH2_ARATH (P25070) Calmodulin-rela... 60 3e-10
BP042515 58 1e-09
TC8059 GB|AAB68399.1|1773321|HAU79736 calmodulin {Helianthus ann... 57 4e-09
TC14393 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, p... 51 2e-07
TC14602 UP|Q8GS56 (Q8GS56) Phosphoenolpyruvate carboxylase kinas... 50 3e-07
TC17306 similar to UP|Q42479 (Q42479) Calcium-dependent protein ... 49 8e-07
BP053279 47 2e-06
BP051870 47 3e-06
TC8061 UP|AAA34013 (AAA34013) Calmodulin, complete 45 3e-06
>TC11056 similar to UP|O81390 (O81390) Calcium-dependent protein kinase ,
partial (22%)
Length = 675
Score = 117 bits (294), Expect = 1e-27
Identities = 70/124 (56%), Positives = 83/124 (66%), Gaps = 3/124 (2%)
Frame = +3
Query: 109 TITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGIIDYDEFITAS--QQLCTRTD*TE 166
TIT EELK GL + G+RLSE +VKQLMEAAD DGNG IDY EFI+A+ + R +
Sbjct: 3 TITYEELKSGLARIGSRLSETEVKQLMEAADVDGNGSIDYLEFISATMHRHRLERDEHLY 182
Query: 167 KNMFILPSNTSIKIT-TELEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRK 225
K + S IT ELE A+ ++ M D IKEIISEVD DNDGRINY+EF AMMR
Sbjct: 183 KAFQYFDKDNSGHITIDELETAMTQHGMGDEASIKEIISEVDTDNDGRINYEEFCAMMRS 362
Query: 226 GNPE 229
G P+
Sbjct: 363 GMPQ 374
Score = 44.3 bits (103), Expect = 2e-05
Identities = 22/57 (38%), Positives = 35/57 (60%)
Frame = +3
Query: 95 LKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGIIDYDEF 151
L + F+ D DNSG ITI+EL+ + + G E +K+++ D D +G I+Y+EF
Sbjct: 177 LYKAFQYFDKDNSGHITIDELETAMTQHGMG-DEASIKEIISEVDTDNDGRINYEEF 344
>AV418237
Length = 361
Score = 107 bits (267), Expect = 2e-24
Identities = 58/103 (56%), Positives = 75/103 (72%), Gaps = 7/103 (6%)
Frame = +2
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T 77
WPSIS SAKDL+RKML+ +PK R++A++VL HPWI +D APD PLD+AVL+R L+Q +
Sbjct: 59 WPSISDSAKDLIRKMLDRNPKTRLTAHQVLCHPWIVDDNIAPDKPLDSAVLSR--LKQFS 232
Query: 78 DSRKL-------L*SCLSEEEIMGLKQMFKGMDTDNSGTITIE 113
KL + LSEEEI GLK++FK +D DNSGTIT +
Sbjct: 233 AMNKLKKMALRVIAESLSEEEIGGLKELFKMIDADNSGTITFD 361
>TC10825 similar to GB|AAP68339.1|31711966|BT008900 At1g74740 {Arabidopsis
thaliana;}, partial (32%)
Length = 1091
Score = 106 bits (265), Expect = 3e-24
Identities = 62/141 (43%), Positives = 87/141 (60%), Gaps = 4/141 (2%)
Frame = +1
Query: 90 EEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGIIDYD 149
EE+ +K MF MDTD G ++ EELK GL K G++L++Q++K LME AD DGNG++DY
Sbjct: 4 EEVEIIKDMFTLMDTDKDGRVSYEELKAGLRKVGSQLADQEIKMLMEVADVDGNGVLDYG 183
Query: 150 EF--ITASQQLCTRTD*TEKNMFILPSNTSIKITT-ELEQAL-HEYNMHDGRYIKEIISE 205
EF +T Q + K + S I + EL++AL E + D + +I+ E
Sbjct: 184 EFVAVTIHLQKMENDEHFHKAFKFFDKDGSGYIESGELQEALADESGVTDADVLNDIMRE 363
Query: 206 VDADNDGRINYDEFVAMMRKG 226
VD D DGRI+Y+EFVAMM+ G
Sbjct: 364 VDTDKDGRISYEEFVAMMKAG 426
>TC18786 similar to UP|Q42479 (Q42479) Calcium-dependent protein kinase
CDPK6 (AT4g23650/F9D16_120) , partial (11%)
Length = 405
Score = 81.6 bits (200), Expect = 1e-16
Identities = 39/55 (70%), Positives = 46/55 (82%)
Frame = +1
Query: 181 TTELEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGNPEAHTKKR 235
T ELE AL +YNM D + IKEII+EVD D+DGRINYDEFVAMMRKGNP+ T++R
Sbjct: 13 TEELESALKKYNMGDEKTIKEIIAEVDTDHDGRINYDEFVAMMRKGNPDIITQRR 177
Score = 33.9 bits (76), Expect = 0.026
Identities = 18/45 (40%), Positives = 27/45 (60%)
Frame = +1
Query: 108 GTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGIIDYDEFI 152
G IT EEL+ L K E+ +K+++ D D +G I+YDEF+
Sbjct: 1 GYITTEELESALKKYNMG-DEKTIKEIIAEVDTDHDGRINYDEFV 132
>BP034163
Length = 477
Score = 74.7 bits (182), Expect = 1e-14
Identities = 40/94 (42%), Positives = 60/94 (63%), Gaps = 7/94 (7%)
Frame = +1
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNR------- 70
WP +S +AKDLV+KMLN DPK+R++A EVL+HPW+ +AP+ L V R
Sbjct: 202 WPKVSDNAKDLVKKMLNPDPKRRLTAQEVLDHPWLVHAKKAPNVSLGETVKARLKQFSVM 381
Query: 71 NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDT 104
N L++ + +++ L+ EE GL++ F+GMDT
Sbjct: 382 NKLKK--RALRVIAEHLTVEEAAGLREGFQGMDT 477
>TC11097 similar to UP|P93520 (P93520) Calcium/calmodulin-dependent protein
kinase homolog|CaM kinase homolog|MCK1, partial (36%)
Length = 949
Score = 72.4 bits (176), Expect = 6e-14
Identities = 42/144 (29%), Positives = 80/144 (55%), Gaps = 6/144 (4%)
Frame = +2
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAV--LNRNSLEQ 75
WPS++P AKD V+++LN D ++R++A + L HPW+++D PLD + L ++ L
Sbjct: 131 WPSVTPEAKDFVKRLLNKDYRKRMTAAQALTHPWLRDDSR--PIPLDILIYKLVKSYLHA 304
Query: 76 *TDSR---KLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGT-RLSEQQV 131
R K L L+EE+++ L+ F+ ++ + G I+++ K L + T + E +V
Sbjct: 305 TPFKRAAVKALSKALTEEQLVYLRAQFRLLEPNRDGHISLDNFKMALARHATDAMRESRV 484
Query: 132 KQLMEAADADGNGIIDYDEFITAS 155
++ + +D++EF A+
Sbjct: 485 LDIIHMMEPLAYRKMDFEEFCAAA 556
>TC8422 UP|AAQ63461 (AAQ63461) Calmodulin 4 (Fragment), complete
Length = 832
Score = 69.3 bits (168), Expect = 5e-13
Identities = 46/148 (31%), Positives = 73/148 (49%), Gaps = 11/148 (7%)
Frame = +1
Query: 87 LSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGII 146
L++++I K+ F D D G+IT +EL + G +E +++ ++ DADGNG I
Sbjct: 148 LTDDQISEFKEAFSLFDKDGDGSITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTI 327
Query: 147 DYDEFITASQQLCTRTD*TEK-----NMFILPSNTSIK------ITTELEQALHEYNMHD 195
D+ EF+ + TD E+ +F N I + T L + L +
Sbjct: 328 DFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTD----- 492
Query: 196 GRYIKEIISEVDADNDGRINYDEFVAMM 223
+ E+I E D D DG+INY+EFV +M
Sbjct: 493 -EEVDEMIREADVDGDGQINYEEFVKVM 573
>TC8060 UP|O49184 (O49184) Calmodulin (ESTS AU081349) (Auxin-regulated
calmodulin) (Calmodulin TACAM1-1) (Calmodulin TACAM1-2)
(Calmodulin TACAM1-3) (Calmodulin TACAM3-1) (Calmodulin
TACAM3-2) (Calmodulin TACAM3-3) (Calmodulin TACAM4-1)
(CaM protein), complete
Length = 615
Score = 68.6 bits (166), Expect = 9e-13
Identities = 46/148 (31%), Positives = 72/148 (48%), Gaps = 11/148 (7%)
Frame = +3
Query: 87 LSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGII 146
L++++I K+ F D D G IT +EL + G +E +++ ++ DADGNG I
Sbjct: 81 LTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTI 260
Query: 147 DYDEFITASQQLCTRTD*TEK-----NMFILPSNTSIK------ITTELEQALHEYNMHD 195
D+ EF+ + TD E+ +F N I + T L + L +
Sbjct: 261 DFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTD----- 425
Query: 196 GRYIKEIISEVDADNDGRINYDEFVAMM 223
+ E+I E D D DG+INY+EFV +M
Sbjct: 426 -EEVDEMIREADVDGDGQINYEEFVKVM 506
>TC9581 UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, complete
Length = 1049
Score = 67.4 bits (163), Expect = 2e-12
Identities = 46/146 (31%), Positives = 73/146 (49%), Gaps = 8/146 (5%)
Frame = +3
Query: 95 LKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGIIDYDEFITA 154
LK++F+ D + G IT +EL L G + ++++ Q++E D +G+G +D DEF
Sbjct: 351 LKRVFQMFDRNGDGRITKKELNDSLENLGIFIPDKELTQMIERIDVNGDGCVDIDEFGEL 530
Query: 155 SQQLCTRTD*TEK-----NMFILPSNTSIKITTELEQALHEYNMHDGRYI---KEIISEV 206
Q + D E N+F + I + EL L + GR + K++I +V
Sbjct: 531 YQSIMDERDEEEDMREAFNVFDQNGDGFITV-EELRTVLASLGIKQGRTVEDCKKMIMKV 707
Query: 207 DADNDGRINYDEFVAMMRKGNPEAHT 232
D D DG ++Y EF MM+ G A T
Sbjct: 708 DVDGDGMVDYKEFKQMMKGGGFSALT 785
>TC15005 homologue to UP|Q9AR92 (Q9AR92) Protein kinase , partial (19%)
Length = 657
Score = 65.1 bits (157), Expect = 1e-11
Identities = 45/97 (46%), Positives = 55/97 (56%), Gaps = 10/97 (10%)
Frame = +2
Query: 143 NGIIDYDEFITAS--QQLCTRTD*TEKNMFILPSNTSIKITT-ELEQALHEYNMHDGRYI 199
+G IDY EF+TA+ + R D ++S IT ELE A+ EY M D I
Sbjct: 14 DGTIDYIEFVTATMHRHKLERDDHLN*AFPYFDKDSSGFITRDELEIAMKEYGMGDDATI 193
Query: 200 KEIISEVDA-------DNDGRINYDEFVAMMRKGNPE 229
KEIISEVD D+DGRINY+EF AMMR GN +
Sbjct: 194 KEIISEVDTIISEVDTDHDGRINYEEFCAMMRSGNQQ 304
>TC18713 similar to GB|AAL90989.1|19699246|AY090328 AT3g03000/F13E7_5
{Arabidopsis thaliana;}, partial (88%)
Length = 868
Score = 63.5 bits (153), Expect = 3e-11
Identities = 43/144 (29%), Positives = 76/144 (51%), Gaps = 11/144 (7%)
Frame = +1
Query: 87 LSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGII 146
L EE+I L+++F+ D +N G++T EL L G + S +QV+ ++ AD + NG+I
Sbjct: 193 LDEEQIAELREIFRSFDRNNDGSLTQLELSSLLRSLGLKPSAEQVEGFIQKADTNSNGLI 372
Query: 147 DYDEF--ITASQQLCTRTD*TEKNM------FILPSN---TSIKITTELEQALHEYNMHD 195
++ EF + A + L R+ TE+ + F N T+ ++ + + H + +
Sbjct: 373 EFSEFVAVVAPELLPARSPYTEEQLRQLFRVFDRDGNGFITAAELAHSMAKLGHALSAEE 552
Query: 196 GRYIKEIISEVDADNDGRINYDEF 219
+ +I E DAD DG I++ EF
Sbjct: 553 ---LTGMIKEADADGDGMISFQEF 615
Score = 50.1 bits (118), Expect = 3e-07
Identities = 24/60 (40%), Positives = 38/60 (63%)
Frame = +1
Query: 95 LKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGIIDYDEFITA 154
L+Q+F+ D D +G IT EL + K G LS +++ +++ ADADG+G+I + EF A
Sbjct: 445 LRQLFRVFDRDGNGFITAAELAHSMAKLGHALSAEELTGMIKEADADGDGMISFQEFAHA 624
>TC11631 weakly similar to UP|TCH2_ARATH (P25070) Calmodulin-related protein
2, touch-induced, partial (68%)
Length = 530
Score = 60.1 bits (144), Expect = 3e-10
Identities = 42/138 (30%), Positives = 72/138 (51%), Gaps = 7/138 (5%)
Frame = +3
Query: 95 LKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGIIDYDEFITA 154
++++F D + G I+ ELK L GT+ S ++VK++M D +G+G ID EF
Sbjct: 102 VRKIFNKFDKNGDGKISCTELKEMLCALGTKTSSEEVKRMMAEIDQNGDGYIDLKEFAEF 281
Query: 155 SQQLCTRTD*TEK----NMFILPSNTSI---KITTELEQALHEYNMHDGRYIKEIISEVD 207
D E +++ L N I ++ + L++ + ++ D R ++IS VD
Sbjct: 282 HLHDAGEDDSKELRDACDLYDLDKNGLISANELHSVLKKLGEKCSLGDCR---KMISNVD 452
Query: 208 ADNDGRINYDEFVAMMRK 225
AD DG +N++EF MM +
Sbjct: 453 ADGDGNVNFEEFKKMMAR 506
Score = 36.2 bits (82), Expect = 0.005
Identities = 19/63 (30%), Positives = 34/63 (53%)
Frame = +3
Query: 89 EEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGIIDY 148
E++ L+ D D +G I+ EL L K G + S ++++ DADG+G +++
Sbjct: 300 EDDSKELRDACDLYDLDKNGLISANELHSVLKKLGEKCSLGDCRKMISNVDADGDGNVNF 479
Query: 149 DEF 151
+EF
Sbjct: 480 EEF 488
>BP042515
Length = 539
Score = 58.2 bits (139), Expect = 1e-09
Identities = 29/57 (50%), Positives = 41/57 (71%)
Frame = -1
Query: 183 ELEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGNPEAHTKKRHDST 239
EL+QA E+ + D R ++E+I E D DNDGRI+Y+EFVAMM++GN + K R S+
Sbjct: 503 ELQQACEEFGLGDVR-LEEMIREADQDNDGRIDYNEFVAMMQRGNADLGKKGRKGSS 336
Score = 38.1 bits (87), Expect = 0.001
Identities = 20/55 (36%), Positives = 34/55 (61%)
Frame = -1
Query: 103 DTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGIIDYDEFITASQQ 157
D D SG IT +EL+ + G L + ++++++ AD D +G IDY+EF+ Q+
Sbjct: 536 DKDGSGYITQDELQQACEEFG--LGDVRLEEMIREADQDNDGRIDYNEFVAMMQR 378
>TC8059 GB|AAB68399.1|1773321|HAU79736 calmodulin {Helianthus annuus;} ,
partial (83%)
Length = 836
Score = 56.6 bits (135), Expect = 4e-09
Identities = 40/127 (31%), Positives = 61/127 (47%), Gaps = 11/127 (8%)
Frame = +2
Query: 108 GTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGIIDYDEFITASQQLCTRTD*TEK 167
G IT +EL + G +E +++ ++ DADGNG ID+ EF+ + TD E+
Sbjct: 164 GCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNLMARKMKDTDSEEE 343
Query: 168 -----NMFILPSNTSIK------ITTELEQALHEYNMHDGRYIKEIISEVDADNDGRINY 216
+F N I + T L + L + + E+I E D D DG+INY
Sbjct: 344 LKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTD------EEVDEMIREADVDGDGQINY 505
Query: 217 DEFVAMM 223
+EFV +M
Sbjct: 506 EEFVKVM 526
Score = 51.6 bits (122), Expect = 1e-07
Identities = 25/65 (38%), Positives = 43/65 (65%)
Frame = +2
Query: 88 SEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGIID 147
SEEE LK+ F+ D D +G I+ EL+ + G +L++++V +++ AD DG+G I+
Sbjct: 332 SEEE---LKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVDGDGQIN 502
Query: 148 YDEFI 152
Y+EF+
Sbjct: 503 YEEFV 517
>TC14393 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, partial
(85%)
Length = 786
Score = 50.8 bits (120), Expect = 2e-07
Identities = 39/152 (25%), Positives = 74/152 (48%), Gaps = 5/152 (3%)
Frame = +3
Query: 77 TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLME 136
T+S+ + + +++ LK++F DT+ G I++ EL L G+ + +++K++M
Sbjct: 99 TNSKASIKPSVYLDDMEELKRVFTRFDTNGDGKISVNELDNVLRSLGSGVPPEELKRVMV 278
Query: 137 AADADGNGIIDYDEFITASQQLCTRTD*TEKN----MFILPSNTSIKITTELEQALHEYN 192
D D +G I+ EF + +E + ++ N I EL + L+
Sbjct: 279 DLDGDHDGFINLSEFAAFCRSDTADGGASELHDAFELYDQDKNGLIS-AAELCKVLNRLG 455
Query: 193 MH-DGRYIKEIISEVDADNDGRINYDEFVAMM 223
M+ ++I VD+D DG +N++EF MM
Sbjct: 456 MNCSEEECHKMIKSVDSDGDGNVNFEEFKKMM 551
Score = 33.1 bits (74), Expect = 0.044
Identities = 13/40 (32%), Positives = 22/40 (54%)
Frame = +3
Query: 199 IKEIISEVDADNDGRINYDEFVAMMRKGNPEAHTKKRHDS 238
+K ++ ++D D+DG IN EF A R + + HD+
Sbjct: 261 LKRVMVDLDGDHDGFINLSEFAAFCRSDTADGGASELHDA 380
>TC14602 UP|Q8GS56 (Q8GS56) Phosphoenolpyruvate carboxylase kinase, complete
Length = 1256
Score = 50.1 bits (118), Expect = 3e-07
Identities = 22/49 (44%), Positives = 32/49 (64%)
Frame = +2
Query: 17 IWPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDN 65
I+ ++SP+AKDL+RKM+ DP RISA + L HPW G+ + +N
Sbjct: 737 IFRNVSPAAKDLLRKMICRDPSNRISAEQALRHPWFLSAGDTVNVT*EN 883
>TC17306 similar to UP|Q42479 (Q42479) Calcium-dependent protein kinase
CDPK6 (AT4g23650/F9D16_120) , partial (6%)
Length = 511
Score = 48.9 bits (115), Expect = 8e-07
Identities = 22/29 (75%), Positives = 24/29 (81%)
Frame = +3
Query: 207 DADNDGRINYDEFVAMMRKGNPEAHTKKR 235
D DNDGRIN +EFVAMMRKGNPE T +R
Sbjct: 3 DTDNDGRINCEEFVAMMRKGNPELVTDRR 89
>BP053279
Length = 442
Score = 47.4 bits (111), Expect = 2e-06
Identities = 24/48 (50%), Positives = 33/48 (68%)
Frame = -3
Query: 187 ALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGNPEAHTKK 234
A E+ + D R ++EII E+D DNDGR++Y+EFV M+ KGN KK
Sbjct: 440 ACDEFXLKDVR-LEEIIGEIDPDNDGRLDYNEFVDMIPKGNRSLVGKK 300
>BP051870
Length = 519
Score = 47.0 bits (110), Expect = 3e-06
Identities = 21/30 (70%), Positives = 27/30 (90%)
Frame = -3
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVL 47
WP+IS SAKDLVRKML DPK+R++A++VL
Sbjct: 310 WPNISESAKDLVRKMLIRDPKKRMTAHQVL 221
>TC8061 UP|AAA34013 (AAA34013) Calmodulin, complete
Length = 568
Score = 44.7 bits (104), Expect = 1e-05
Identities = 18/51 (35%), Positives = 35/51 (68%)
Frame = +2
Query: 102 MDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGIIDYDEFI 152
+D D +G I+ EL+ + G +L++++V +++ AD DG+G I+Y+EF+
Sbjct: 323 LDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVDGDGQINYEEFV 475
Score = 36.2 bits (82), Expect(2) = 3e-06
Identities = 20/45 (44%), Positives = 27/45 (59%), Gaps = 8/45 (17%)
Frame = +2
Query: 199 IKEIISEVDADNDGRINYDEFV-AMMRKGNP-------EAHTKKR 235
+ E+I E D D DG+INY+EFV MM K P +AH +K+
Sbjct: 410 VDEMIREADVDGDGQINYEEFVKVMMAK*GPIHNTNI*KAHEQKK 544
Score = 30.0 bits (66), Expect(2) = 3e-06
Identities = 13/36 (36%), Positives = 20/36 (55%)
Frame = +1
Query: 132 KQLMEAADADGNGIIDYDEFITASQQLCTRTD*TEK 167
+ ++ DADGNG ID+ EF+ + TD E+
Sbjct: 193 RHMINEVDADGNGTIDFPEFLNLMARKMKDTDSEEE 300
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.137 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,043,862
Number of Sequences: 28460
Number of extensions: 30844
Number of successful extensions: 229
Number of sequences better than 10.0: 95
Number of HSP's better than 10.0 without gapping: 198
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 216
length of query: 240
length of database: 4,897,600
effective HSP length: 88
effective length of query: 152
effective length of database: 2,393,120
effective search space: 363754240
effective search space used: 363754240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 54 (25.4 bits)
Medicago: description of AC146941.2


