

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146940.4 + phase: 1 /pseudo
(674 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
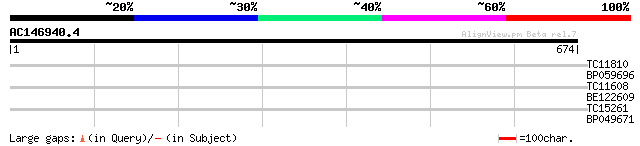
Score E
Sequences producing significant alignments: (bits) Value
TC11810 33 0.20
BP059696 29 2.2
TC11608 similar to GB|AAF68117.1|7715599|AC010793 F20B17.17 {Ara... 28 6.3
BE122609 27 8.2
TC15261 weakly similar to UP|Q9ZQ55 (Q9ZQ55) At2g22570 protein, ... 27 8.2
BP049671 27 8.2
>TC11810
Length = 640
Score = 32.7 bits (73), Expect = 0.20
Identities = 15/31 (48%), Positives = 20/31 (64%)
Frame = +3
Query: 572 NRLEHQRLNDLVFVRYNLMLENRNNKIRNYD 602
NRL ++LND+VFV YNL L + R+ D
Sbjct: 3 NRLSQKKLNDIVFVHYNLRLRECQVRKRSRD 95
>BP059696
Length = 206
Score = 29.3 bits (64), Expect = 2.2
Identities = 16/38 (42%), Positives = 18/38 (47%), Gaps = 2/38 (5%)
Frame = -1
Query: 240 CCSDSDR*CCKLCCCW*VIGERVPWP--VLVSLCNSLH 275
CC *CC LCCCW W L+ LC +LH
Sbjct: 164 CC-----*CCCLCCCW--------WKCLCLMPLCLALH 90
>TC11608 similar to GB|AAF68117.1|7715599|AC010793 F20B17.17 {Arabidopsis
thaliana;} , partial (22%)
Length = 739
Score = 27.7 bits (60), Expect = 6.3
Identities = 12/29 (41%), Positives = 17/29 (58%)
Frame = +1
Query: 568 KKKRNRLEHQRLNDLVFVRYNLMLENRNN 596
++ RL LNDLV++ YNL L + N
Sbjct: 190 QRSGTRLIENTLNDLVYINYNLKLARQMN 276
>BE122609
Length = 356
Score = 27.3 bits (59), Expect = 8.2
Identities = 22/56 (39%), Positives = 31/56 (55%)
Frame = +3
Query: 510 LWKEICYRSSKFTISR*MVRTLRVSSTTFAKIGDSGSKSNL*LFWLREKLECV*AY 565
LW E +++ + IS *+V L + T FA I + +L*L L E+LE *AY
Sbjct: 168 LWNESEHKTKDYHISC*VVEELLTAYT*FAIICC*SALLDL*LIRL*EELENF*AY 335
>TC15261 weakly similar to UP|Q9ZQ55 (Q9ZQ55) At2g22570 protein, partial
(75%)
Length = 1031
Score = 27.3 bits (59), Expect = 8.2
Identities = 10/31 (32%), Positives = 15/31 (48%)
Frame = +3
Query: 112 CKMVHCCIYSLQCSKFTIFSVCGRCSLLHGS 142
C C++S C K + S G+C +H S
Sbjct: 519 CMRAGFCLFSFVCEKSPVPSASGKCDCVHRS 611
>BP049671
Length = 488
Score = 27.3 bits (59), Expect = 8.2
Identities = 9/17 (52%), Positives = 10/17 (57%), Gaps = 1/17 (5%)
Frame = +3
Query: 240 CCSDS-DR*CCKLCCCW 255
CC D R C +CCCW
Sbjct: 306 CCQDHHSRNTCPICCCW 356
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.366 0.162 0.645
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,588,417
Number of Sequences: 28460
Number of extensions: 245702
Number of successful extensions: 4188
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 2316
Number of HSP's successfully gapped in prelim test: 119
Number of HSP's that attempted gapping in prelim test: 1578
Number of HSP's gapped (non-prelim): 2724
length of query: 674
length of database: 4,897,600
effective HSP length: 96
effective length of query: 578
effective length of database: 2,165,440
effective search space: 1251624320
effective search space used: 1251624320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146940.4


