

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146866.2 + phase: 0
(504 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
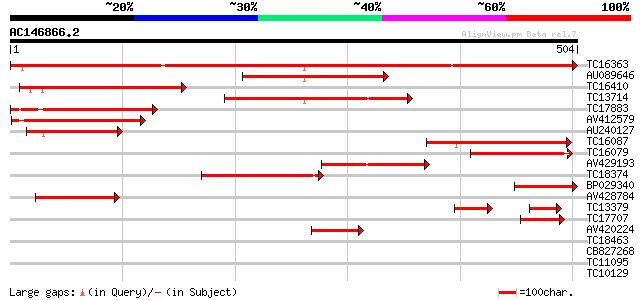
Score E
Sequences producing significant alignments: (bits) Value
TC16363 UP|Q84RQ3 (Q84RQ3) Sucrose transporter 4 protein, complete 829 0.0
AU089646 228 1e-60
TC16410 similar to UP|Q9XHL6 (Q9XHL6) Sucrose transport protein ... 189 7e-49
TC13714 homologue to UP|Q7XA53 (Q7XA53) Sucrose transporter, par... 169 7e-43
TC17883 similar to UP|Q41152 (Q41152) Sucrose carrier, partial (... 162 1e-40
AV412579 149 8e-37
AU240127 139 1e-33
TC16087 similar to UP|Q40583 (Q40583) Sucrose transporter, parti... 124 3e-29
TC16079 similar to UP|O04077 (O04077) Sucrose transport protein,... 107 3e-24
AV429193 105 1e-23
TC18374 similar to UP|Q9FIX9 (Q9FIX9) Sucrose transporter protei... 103 9e-23
BP029340 93 1e-19
AV428784 72 2e-13
TC13379 similar to UP|O65803 (O65803) Sucrose/H+ symporter, part... 43 4e-10
TC17707 similar to UP|Q7XA53 (Q7XA53) Sucrose transporter, parti... 55 2e-08
AV420224 53 1e-07
TC18463 similar to AAR92278 (AAR92278) At4g22580, partial (34%) 32 0.19
CB827268 30 0.71
TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 29 2.1
TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%) 28 3.5
>TC16363 UP|Q84RQ3 (Q84RQ3) Sucrose transporter 4 protein, complete
Length = 2164
Score = 829 bits (2142), Expect = 0.0
Identities = 416/514 (80%), Positives = 453/514 (87%), Gaps = 10/514 (1%)
Frame = +3
Query: 1 MPNPTTTNPH---RSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLS 57
MP H RS+ R STS++ P +P R PLRQLLRVASVASGIQFGWALQLS
Sbjct: 153 MPGDPDNRHHHHPRSKNRPSTSSARPPPSRPPPARVPLRQLLRVASVASGIQFGWALQLS 332
Query: 58 LLTPYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIV 117
LLTPYVQQLGIPH+WASIIWLCGPVSGLFVQPLVGHLSD+C+SRFGRRRPFIL GAASIV
Sbjct: 333 LLTPYVQQLGIPHQWASIIWLCGPVSGLFVQPLVGHLSDKCTSRFGRRRPFILAGAASIV 512
Query: 118 VAVVIIGYAADIGYLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCN 177
VAV+IIGYAADIG+++GD T+++RP AI VFVIGFWILDVANNVTQGPCRALLADLT
Sbjct: 513 VAVLIIGYAADIGWMLGD--TESFRPAAITVFVIGFWILDVANNVTQGPCRALLADLTSK 686
Query: 178 DARRTRVANAYFSLFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVA 237
D RRTRVANAYFSLFMA+GNILGYATG+YSGWY+IFTFTL+PAC+ISCANLKSAFFLDVA
Sbjct: 687 DNRRTRVANAYFSLFMAIGNILGYATGAYSGWYRIFTFTLSPACTISCANLKSAFFLDVA 866
Query: 238 FIVVTTYLSIVSAHEVPLSSSGA-------GESGSAEEAFMWELFGTFKYFSMPVWIVLS 290
FI VTTY+SI +AHEVPL+SSGA GESGS EEAFMWELFGTFKYFS VWI+LS
Sbjct: 867 FIAVTTYVSITAAHEVPLNSSGAAHAGEGAGESGSTEEAFMWELFGTFKYFSSTVWIILS 1046
Query: 291 VTALTWIGWFPFNLFDTDWMGREIYGGDPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLM 350
VTAL W GWFPF LFDTDWMGREIYG DP GG YD GVRMGALGL+LNSVVL VTSLLM
Sbjct: 1047VTALNWTGWFPFILFDTDWMGREIYGADPNGGPNYDAGVRMGALGLMLNSVVLGVTSLLM 1226
Query: 351 ERLCRKRGAGFVWGISNIFMAICFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTI 410
E+LCRKRGAGFVWGISNI MA+CF+AMLV+TY AN+IGYV K PPTGIVIAAL IFTI
Sbjct: 1227EKLCRKRGAGFVWGISNILMAVCFLAMLVVTYVANTIGYVGK-DLPPTGIVIAALIIFTI 1403
Query: 411 LGFPMAITYSVPYALISTHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGN 470
LGFP+AITYSVPYALIS H EPLGLGQGLSMGVLNLAIV+PQIVVSLGSGPWDQLFGGGN
Sbjct: 1404LGFPLAITYSVPYALISKHTEPLGLGQGLSMGVLNLAIVIPQIVVSLGSGPWDQLFGGGN 1583
Query: 471 SPAFAVAAVAALLSGLLALLAIPRTRTQKPRVRI 504
S AFAV AVAA++SGLLA+LAIPRT TQKP++R+
Sbjct: 1584SAAFAVGAVAAIMSGLLAVLAIPRTGTQKPQIRV 1685
>AU089646
Length = 411
Score = 228 bits (582), Expect = 1e-60
Identities = 106/136 (77%), Positives = 118/136 (85%), Gaps = 7/136 (5%)
Frame = +2
Query: 208 GWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSSSGA------- 260
GWYK+F FTLTPAC++SCANLKSAFFLD+ FIV+TTY+S VSAHEVPL+SSGA
Sbjct: 2 GWYKVFPFTLTPACNVSCANLKSAFFLDIIFIVITTYISTVSAHEVPLNSSGAHPVEEGA 181
Query: 261 GESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGGDPE 320
GES SAEEAF+WE+FGTFKYFS P+WI+LSVTALTWIGWFPF LFDTDWMGREIYGG+P
Sbjct: 182 GESXSAEEAFLWEMFGTFKYFSKPIWIILSVTALTWIGWFPFLLFDTDWMGREIYGGEPN 361
Query: 321 GGLIYDTGVRMGALGL 336
G YD GVRMGALGL
Sbjct: 362 EGPNYDIGVRMGALGL 409
>TC16410 similar to UP|Q9XHL6 (Q9XHL6) Sucrose transport protein SUT1,
partial (27%)
Length = 502
Score = 189 bits (481), Expect = 7e-49
Identities = 90/165 (54%), Positives = 117/165 (70%), Gaps = 16/165 (9%)
Frame = +2
Query: 9 PHRSRTRSS--TSTSTSRPVQ--------------PVQPRTPLRQLLRVASVASGIQFGW 52
PH T+ + T T T P+ P +PLR+++ VAS+A+G+QFGW
Sbjct: 8 PHHLNTKHNPKTQTLTMEPLSKHNNLKPSSLHVEAPPPEASPLRKIIVVASIAAGVQFGW 187
Query: 53 ALQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVG 112
ALQLSLLTPYVQ LGIPHKWA+ IWLCGP+SG+ VQP+VG+ SDRC+SRFGRRRPFI G
Sbjct: 188 ALQLSLLTPYVQLLGIPHKWAAYIWLCGPISGMLVQPIVGYHSDRCTSRFGRRRPFIAAG 367
Query: 113 AASIVVAVVIIGYAADIGYLIGDDITQNYRPFAIVVFVIGFWILD 157
A ++ +AV +IGYAAD G+ +GD + + R AI +F++GFWILD
Sbjct: 368 ALAVAIAVFLIGYAADFGHALGDRLDKKIRTRAIAIFIVGFWILD 502
>TC13714 homologue to UP|Q7XA53 (Q7XA53) Sucrose transporter, partial (32%)
Length = 512
Score = 169 bits (429), Expect = 7e-43
Identities = 88/169 (52%), Positives = 106/169 (62%), Gaps = 2/169 (1%)
Frame = +2
Query: 192 FMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAH 251
FMAVGN+LGYA G+YS Y +F FT T AC + CANLKS FFL +A + ++V
Sbjct: 2 FMAVGNVLGYAAGAYSKLYHVFPFTKTTACDVYCANLKSCFFLSIALLTTLATAALVYVK 181
Query: 252 EVPLSSSGA--GESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDW 309
EVPLS A + +L G F+ P+WI+L VT L WI WFPF LFDTDW
Sbjct: 182 EVPLSPEKAVIDSDDNGGMPCFGQLSGAFRELKRPMWILLLVTCLNWIAWFPFLLFDTDW 361
Query: 310 MGREIYGGDPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRG 358
MG+E+YGG+ G YD GV GALGL+LNSVVL TSL +E L R G
Sbjct: 362 MGKEVYGGE-VGDHAYDQGVHAGALGLMLNSVVLGATSLGLEYLARGVG 505
>TC17883 similar to UP|Q41152 (Q41152) Sucrose carrier, partial (19%)
Length = 428
Score = 162 bits (410), Expect = 1e-40
Identities = 80/131 (61%), Positives = 98/131 (74%)
Frame = +3
Query: 1 MPNPTTTNPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLT 60
M +PT +N H SS P Q +PLR+++ VAS+A+GIQFGWALQLSLLT
Sbjct: 51 MESPTKSN-HLDTKGSSLQIEAHIP----QQSSPLRKMIAVASIAAGIQFGWALQLSLLT 215
Query: 61 PYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAV 120
PYVQ LG+PH WAS IWLCGPVSGL VQP+VG+ SDR + RFGRRRPFIL GA ++ ++V
Sbjct: 216 PYVQTLGVPHIWASFIWLCGPVSGLLVQPIVGYYSDRTTCRFGRRRPFILAGAVAVAISV 395
Query: 121 VIIGYAADIGY 131
+IGYAADIG+
Sbjct: 396 FLIGYAADIGH 428
>AV412579
Length = 394
Score = 149 bits (377), Expect = 8e-37
Identities = 70/119 (58%), Positives = 88/119 (73%)
Frame = +3
Query: 2 PNPTTTNPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTP 61
PN TT N + S+ P Q +PL +++ VAS+A+G+QFGWALQLSLLTP
Sbjct: 48 PNSTTQN----NNSIAPSSFQVEPAQHAAGPSPLAKMIAVASIAAGVQFGWALQLSLLTP 215
Query: 62 YVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAV 120
YVQ LG+PHKW+S IWLCGP+SGL VQP+VG+ SDRC+SRFGRRRPFI GA ++ +AV
Sbjct: 216 YVQLLGVPHKWSSFIWLCGPISGLVVQPIVGYSSDRCTSRFGRRRPFIFAGAIAVAIAV 392
>AU240127
Length = 300
Score = 139 bits (350), Expect = 1e-33
Identities = 66/88 (75%), Positives = 77/88 (87%), Gaps = 3/88 (3%)
Frame = +3
Query: 16 SSTSTSTSRPVQP---VQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKW 72
++T+T+T P+QP + R PL QLL+VASVA GIQFGWALQLSLLTPYVQQLGIPH W
Sbjct: 27 TTTTTTTQPPLQPRPKPRARVPLTQLLKVASVAGGIQFGWALQLSLLTPYVQQLGIPHAW 206
Query: 73 ASIIWLCGPVSGLFVQPLVGHLSDRCSS 100
ASIIWLCGP+SGLFVQPLVGH+SDRC++
Sbjct: 207 ASIIWLCGPLSGLFVQPLVGHMSDRCTN 290
>TC16087 similar to UP|Q40583 (Q40583) Sucrose transporter, partial (23%)
Length = 623
Score = 124 bits (312), Expect = 3e-29
Identities = 63/132 (47%), Positives = 86/132 (64%), Gaps = 3/132 (2%)
Frame = +2
Query: 371 AICFIAMLVLTYAANSIGYVSKGQP---PPTGIVIAALAIFTILGFPMAITYSVPYALIS 427
A+C +++T A VS G P G+ AL F +LG P+AITYSVP+AL S
Sbjct: 2 AVCMAMTMLITKVAEHDRRVSGGATIGRPTPGVKAGALMFFAVLGIPLAITYSVPFALAS 181
Query: 428 THIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLL 487
+ G GQGLS+GVLN+AIV+PQ++VS +GPWD LFGGGN PAF V AV A +S +L
Sbjct: 182 IYSSSTGAGQGLSLGVLNVAIVIPQMIVSALNGPWDSLFGGGNLPAFMVGAVMAAVSAVL 361
Query: 488 ALLAIPRTRTQK 499
A++ +P + ++
Sbjct: 362 AMVLLPSPKPEE 397
>TC16079 similar to UP|O04077 (O04077) Sucrose transport protein, partial
(18%)
Length = 573
Score = 107 bits (268), Expect = 3e-24
Identities = 53/91 (58%), Positives = 68/91 (74%)
Frame = +2
Query: 410 ILGFPMAITYSVPYALISTHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGG 469
++G P+AI +SVP+AL S + G GQGLS+GVLNLAIVVPQ++VS SGPWD LFGGG
Sbjct: 5 VMGIPLAINFSVPFALASIYSSAAGAGQGLSLGVLNLAIVVPQMLVSTLSGPWDALFGGG 184
Query: 470 NSPAFAVAAVAALLSGLLALLAIPRTRTQKP 500
N PAF V AV A +S ++A++ +P T KP
Sbjct: 185 NLPAFVVGAVMAAVSAIMAIVLLP---TPKP 268
>AV429193
Length = 295
Score = 105 bits (263), Expect = 1e-23
Identities = 53/97 (54%), Positives = 65/97 (66%), Gaps = 1/97 (1%)
Frame = +1
Query: 278 FKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGGDPEGGLIYDTGVRMGALGLL 337
FK P+WI++ VTA+ W+ WFPF LFDTDWMGRE+YGG G YD GVR G+LGL+
Sbjct: 7 FKELKKPMWILMLVTAVNWVAWFPFFLFDTDWMGREVYGG-VVGEKAYDDGVRAGSLGLM 183
Query: 338 LNSVVLAVTSLLMERLCR-KRGAGFVWGISNIFMAIC 373
+N+VVLA SL E L R G +W I N +AIC
Sbjct: 184 INAVVLAFMSLAAEPLWRILGGVKNLWRIVNFILAIC 294
>TC18374 similar to UP|Q9FIX9 (Q9FIX9) Sucrose transporter protein, partial
(17%)
Length = 330
Score = 103 bits (256), Expect = 9e-23
Identities = 49/109 (44%), Positives = 68/109 (61%)
Frame = +3
Query: 171 LADLTCNDARRTRVANAYFSLFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKS 230
L DL D ++TR A +FS FMAVGN+LGYA G+YSG +KIF FT+T AC CANLKS
Sbjct: 6 LGDLAAGDQKKTRTAMGFFSFFMAVGNVLGYAAGAYSGLHKIFPFTVTDACPSFCANLKS 185
Query: 231 AFFLDVAFIVVTTYLSIVSAHEVPLSSSGAGESGSAEEAFMWELFGTFK 279
FF + +++ + +++ + PL+ A + S F +LFG FK
Sbjct: 186 CFFFSILLLLILSIAALIYVEDTPLTKKPAADVDSPVSCF-GDLFGAFK 329
>BP029340
Length = 402
Score = 92.8 bits (229), Expect = 1e-19
Identities = 44/56 (78%), Positives = 52/56 (92%)
Frame = -2
Query: 449 VVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIPRTRTQKPRVRI 504
V PQ+VVSLGSGPWDQLFGGGNSPAFAVAAVAAL+SGL+A+LAIPR+ Q+PR ++
Sbjct: 401 VFPQMVVSLGSGPWDQLFGGGNSPAFAVAAVAALISGLIAVLAIPRSGGQRPRSQV 234
>AV428784
Length = 334
Score = 72.4 bits (176), Expect = 2e-13
Identities = 36/74 (48%), Positives = 48/74 (64%)
Frame = +2
Query: 24 RPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCGPVS 83
+P Q Q T +++ AS+A +QFGWALQ SLLTP VQ LG+P+K S+I GP+
Sbjct: 107 KPAQIAQGPTQFAKIMVGASIAPEVQFGWALQFSLLTPNVQLLGVPNKGPSLIGFWGPIF 286
Query: 84 GLFVQPLVGHLSDR 97
GL VQP+ + DR
Sbjct: 287 GLVVQPMGAYSRDR 328
>TC13379 similar to UP|O65803 (O65803) Sucrose/H+ symporter, partial (12%)
Length = 534
Score = 43.1 bits (100), Expect(2) = 4e-10
Identities = 17/34 (50%), Positives = 27/34 (79%)
Frame = +2
Query: 396 PPTGIVIAALAIFTILGFPMAITYSVPYALISTH 429
P TG+ +A+ +F++LG P+AITYS+P+AL S +
Sbjct: 2 PSTGVKASAMILFSVLGIPLAITYSIPFALGSIY 103
Score = 37.7 bits (86), Expect(2) = 4e-10
Identities = 16/28 (57%), Positives = 22/28 (78%)
Frame = +1
Query: 463 DQLFGGGNSPAFAVAAVAALLSGLLALL 490
D LFGGGN PAF V A+ A++S +LA++
Sbjct: 205 DALFGGGNLPAFLVGAIMAVVSAILAIV 288
Score = 34.3 bits (77), Expect = 0.049
Identities = 20/44 (45%), Positives = 30/44 (67%), Gaps = 7/44 (15%)
Frame = +3
Query: 425 LISTHI------EPL-GLGQGLSMGVLNLAIVVPQIVVSLGSGP 461
LI++H+ +P+ G GQGL GV N+A+V+PQI++ SGP
Sbjct: 69 LIASHLRLDRYTQPVTGGGQGLYFGVHNIAMVIPQIIIWGLSGP 200
>TC17707 similar to UP|Q7XA53 (Q7XA53) Sucrose transporter, partial (11%)
Length = 554
Score = 55.5 bits (132), Expect = 2e-08
Identities = 24/39 (61%), Positives = 31/39 (78%)
Frame = +2
Query: 455 VSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIP 493
VS+ SG WD LFGGGN PAF V AVAA +SG+L+++ +P
Sbjct: 2 VSVSSGNWDDLFGGGNLPAFVVGAVAAAISGILSMVLLP 118
>AV420224
Length = 241
Score = 53.1 bits (126), Expect = 1e-07
Identities = 23/46 (50%), Positives = 30/46 (65%)
Frame = +1
Query: 269 AFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREI 314
A + L + ++ + VL V ALTW+ WFPF LFDTDWMGRE+
Sbjct: 103 AVLVNLLTSLRHLPPAMHSVLIVMALTWLSWFPFFLFDTDWMGREV 240
>TC18463 similar to AAR92278 (AAR92278) At4g22580, partial (34%)
Length = 555
Score = 32.3 bits (72), Expect = 0.19
Identities = 18/37 (48%), Positives = 23/37 (61%)
Frame = +1
Query: 5 TTTNPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRV 41
TTTNP S+T + S+P QPR+PL QLLR+
Sbjct: 67 TTTNPESSQTLRESLDPHSQPPFTFQPRSPL-QLLRI 174
>CB827268
Length = 533
Score = 30.4 bits (67), Expect = 0.71
Identities = 13/33 (39%), Positives = 18/33 (54%)
Frame = +1
Query: 2 PNPTTTNPHRSRTRSSTSTSTSRPVQPVQPRTP 34
P+ T PH T ++T TS +P + PRTP
Sbjct: 82 PHQTHNQPHPITTPTATQTSKPQPTRTQNPRTP 180
>TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (89%)
Length = 912
Score = 28.9 bits (63), Expect = 2.1
Identities = 15/31 (48%), Positives = 20/31 (64%)
Frame = -1
Query: 5 TTTNPHRSRTRSSTSTSTSRPVQPVQPRTPL 35
TTT + T +ST+TSTSR V+ + P T L
Sbjct: 357 TTTRASTTTTTTSTATSTSRIVRKLNPPTIL 265
>TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%)
Length = 562
Score = 28.1 bits (61), Expect = 3.5
Identities = 12/33 (36%), Positives = 18/33 (54%)
Frame = +1
Query: 2 PNPTTTNPHRSRTRSSTSTSTSRPVQPVQPRTP 34
P+PT PH +S+S ++ P P P+TP
Sbjct: 433 PSPTNQPPHIPSPPNSSSPTSQPPNTPSPPKTP 531
Score = 27.7 bits (60), Expect = 4.6
Identities = 11/33 (33%), Positives = 17/33 (51%)
Frame = +1
Query: 2 PNPTTTNPHRSRTRSSTSTSTSRPVQPVQPRTP 34
P+PT+ PH ++ S + P P P+TP
Sbjct: 337 PSPTSQPPHTPTPPNTPSPISQPPYTPTPPKTP 435
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.140 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,378,609
Number of Sequences: 28460
Number of extensions: 139910
Number of successful extensions: 1205
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 1164
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1194
length of query: 504
length of database: 4,897,600
effective HSP length: 94
effective length of query: 410
effective length of database: 2,222,360
effective search space: 911167600
effective search space used: 911167600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146866.2


