

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146866.12 + phase: 0
(273 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
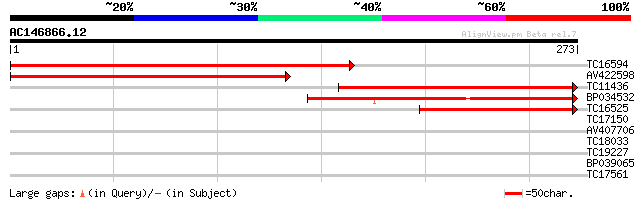
Score E
Sequences producing significant alignments: (bits) Value
TC16594 similar to GB|AAG38144.1|21700765|AY004240 phosphoglycer... 258 8e-70
AV422598 231 1e-61
TC11436 similar to GB|AAG38145.1|21700767|AY004241 phosphoglycer... 206 3e-54
BP034532 182 7e-47
TC16525 similar to GB|AAG38145.1|21700767|AY004241 phosphoglycer... 125 1e-29
TC17150 similar to UP|Q9FPW6 (Q9FPW6) POZ/BTB containing-protein... 28 1.3
AV407706 28 1.7
TC18033 28 2.2
TC19227 27 2.9
BP039065 26 8.3
TC17561 26 8.3
>TC16594 similar to GB|AAG38144.1|21700765|AY004240 phosphoglycerate
mutase-like protein {Glycine max;} , partial (56%)
Length = 578
Score = 258 bits (659), Expect = 8e-70
Identities = 119/166 (71%), Positives = 142/166 (84%)
Frame = +1
Query: 1 MDTAPGQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENL 60
MD Q LYPLH SKT+HLVRHAQG HNV GEK+ +AYLSYD+ DA+LTPLGW QV+NL
Sbjct: 79 MDANVSQGLYPLHRSKTLHLVRHAQGFHNVAGEKDPEAYLSYDYLDASLTPLGWNQVDNL 258
Query: 61 QKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVS 120
++HVK+ GLSK IELV+ SPL RTMQTAVGVFGGEA+TDG++ PPLMI+N G S PA+S
Sbjct: 259 REHVKSSGLSKGIELVITSPLTRTMQTAVGVFGGEASTDGIDSPPLMIDNAGDSARPAIS 438
Query: 121 SLNCPPFVAVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETD 166
SLN PPF+AVELCRE +G+HPCDKRR+++EYR++FP IDFSLIE D
Sbjct: 439 SLNSPPFLAVELCREHLGVHPCDKRRSITEYRNIFPAIDFSLIEID 576
>AV422598
Length = 475
Score = 231 bits (589), Expect = 1e-61
Identities = 109/135 (80%), Positives = 121/135 (88%)
Frame = +1
Query: 1 MDTAPGQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENL 60
MD+A GQ LYPLHH KT+HLVRHAQGVHNVEGEKNHDAYLS D FDA+LTPLGW QV+NL
Sbjct: 70 MDSAAGQILYPLHHCKTVHLVRHAQGVHNVEGEKNHDAYLSNDLFDAHLTPLGWNQVDNL 249
Query: 61 QKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVS 120
+KHVK GLS+ +ELVVVSPLLRTMQTAVGVFGGE +DG+N+PPLMIE+VG SDHPAVS
Sbjct: 250 RKHVKGHGLSRSVELVVVSPLLRTMQTAVGVFGGEIYSDGINEPPLMIESVGSSDHPAVS 429
Query: 121 SLNCPPFVAVELCRE 135
SLNCP F+A ELCRE
Sbjct: 430 SLNCPLFIAKELCRE 474
>TC11436 similar to GB|AAG38145.1|21700767|AY004241 phosphoglycerate
mutase-like protein {Glycine max;} , partial (35%)
Length = 632
Score = 206 bits (525), Expect = 3e-54
Identities = 95/115 (82%), Positives = 104/115 (89%)
Frame = +3
Query: 159 DFSLIETDDDTWWKPEREKKEEVTGRGLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGN 218
DFSL+ETD+DT W PEREKKE V RGLKFLEWL TRKEKEIAVVTHSSFLFNTLSAFGN
Sbjct: 3 DFSLVETDEDTLWAPEREKKEAVAARGLKFLEWLSTRKEKEIAVVTHSSFLFNTLSAFGN 182
Query: 219 DCHPNIKTEMCAHFANCELRSMVIVDKCMIGSNNSTTNYPGKIPHGPDLPSDATD 273
DCHPNIK+E+C HFANCELRSMVI+D+ +IGSN STTNYPGKIP GPDLPS++ D
Sbjct: 183 DCHPNIKSEICKHFANCELRSMVIIDRGVIGSNESTTNYPGKIPRGPDLPSESAD 347
>BP034532
Length = 533
Score = 182 bits (461), Expect = 7e-47
Identities = 87/131 (66%), Positives = 105/131 (79%), Gaps = 1/131 (0%)
Frame = -3
Query: 144 KRRTVSEYRHMFPGIDFSLIETDDDTWWKPE-REKKEEVTGRGLKFLEWLCTRKEKEIAV 202
+RR+VSEY+ +FP +DFSLIE+D+D WW+ + RE KEE+ RG KF+ WL TRKEKEIA+
Sbjct: 531 RRRSVSEYQFLFPAVDFSLIESDEDVWWEADVRETKEELAARGQKFMNWLWTRKEKEIAI 352
Query: 203 VTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRSMVIVDKCMIGSNNSTTNYPGKIP 262
VTHS FL +TL+A NDC P +K E+ HFANCELRSMVIVD+ MIGS STTNYPGKIP
Sbjct: 351 VTHSGFLSHTLNAITNDC-PLMKKEISKHFANCELRSMVIVDRGMIGSETSTTNYPGKIP 175
Query: 263 HGPDLPSDATD 273
GPDLPS+ D
Sbjct: 174 SGPDLPSEVAD 142
>TC16525 similar to GB|AAG38145.1|21700767|AY004241 phosphoglycerate
mutase-like protein {Glycine max;} , partial (30%)
Length = 502
Score = 125 bits (313), Expect = 1e-29
Identities = 55/76 (72%), Positives = 66/76 (86%)
Frame = +3
Query: 198 KEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRSMVIVDKCMIGSNNSTTNY 257
KEIAVV+H+ FLF+ LSAFGNDCHP +K E+C HFANCELRSMVIVD+ +IGS++ ++NY
Sbjct: 3 KEIAVVSHTGFLFHALSAFGNDCHPTVKNEICTHFANCELRSMVIVDRGLIGSDDPSSNY 182
Query: 258 PGKIPHGPDLPSDATD 273
PGKIPHG DLPSD D
Sbjct: 183 PGKIPHGLDLPSDIAD 230
>TC17150 similar to UP|Q9FPW6 (Q9FPW6) POZ/BTB containing-protein AtPOB1,
partial (28%)
Length = 709
Score = 28.5 bits (62), Expect = 1.3
Identities = 10/27 (37%), Positives = 15/27 (55%)
Frame = +3
Query: 228 MCAHFANCELRSMVIVDKCMIGSNNST 254
+C+H A C LR + +D+C N T
Sbjct: 126 LCSHLAMCGLRHSICLDRCFFYQQNVT 206
>AV407706
Length = 428
Score = 28.1 bits (61), Expect = 1.7
Identities = 16/65 (24%), Positives = 29/65 (44%)
Frame = -1
Query: 190 EWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRSMVIVDKCMIG 249
+W+C + A+ F+ N+ +FG N +M H A+C + +D+C+
Sbjct: 419 KWICATESTYFAI----KFIKNSWDSFGRFIRNNDTVDML*HMASC----IFTLDQCLRF 264
Query: 250 SNNST 254
ST
Sbjct: 263 QRTST 249
>TC18033
Length = 560
Score = 27.7 bits (60), Expect = 2.2
Identities = 11/13 (84%), Positives = 11/13 (84%)
Frame = -3
Query: 7 QSLYPLHHSKTIH 19
QSLYPLHHS T H
Sbjct: 396 QSLYPLHHSHTKH 358
>TC19227
Length = 410
Score = 27.3 bits (59), Expect = 2.9
Identities = 12/23 (52%), Positives = 17/23 (73%), Gaps = 1/23 (4%)
Frame = -3
Query: 168 DTWWKPEREKKEEV-TGRGLKFL 189
DT W+P +EKKEE+ + +KFL
Sbjct: 357 DTTWRPLKEKKEELGVQKSMKFL 289
>BP039065
Length = 564
Score = 25.8 bits (55), Expect = 8.3
Identities = 19/51 (37%), Positives = 25/51 (48%), Gaps = 6/51 (11%)
Frame = +1
Query: 141 PCDKRRTVSEY--RHMFPGIDFSLIETDDDTWWKP----EREKKEEVTGRG 185
P +R TVS R M G+ ++ W+P ER+K EVTGRG
Sbjct: 307 PSQRRSTVSSSLCRLMVCGLGKIVLRRRQ*VSWRP*QL*ERKKGLEVTGRG 459
>TC17561
Length = 581
Score = 25.8 bits (55), Expect = 8.3
Identities = 8/16 (50%), Positives = 9/16 (56%)
Frame = -1
Query: 220 CHPNIKTEMCAHFANC 235
CHPN C+HF C
Sbjct: 191 CHPNCYCLTCSHFCGC 144
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,456,620
Number of Sequences: 28460
Number of extensions: 79347
Number of successful extensions: 389
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 386
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 387
length of query: 273
length of database: 4,897,600
effective HSP length: 89
effective length of query: 184
effective length of database: 2,364,660
effective search space: 435097440
effective search space used: 435097440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 54 (25.4 bits)
Medicago: description of AC146866.12


