

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146866.11 - phase: 0 /pseudo
(685 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
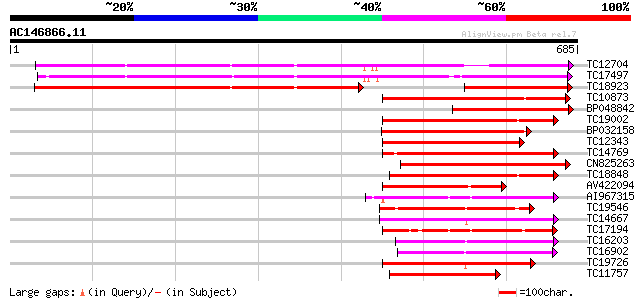
Score E
Sequences producing significant alignments: (bits) Value
TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3,... 556 e-159
TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2,... 534 e-152
TC18923 UP|CAE45594 (CAE45594) S-receptor kinase-like protein 1,... 327 4e-90
TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partia... 207 5e-54
BP048842 201 2e-52
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 192 1e-49
BP032158 192 2e-49
TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine prote... 184 4e-47
TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete 181 3e-46
CN825263 178 2e-45
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 177 7e-45
AV422094 174 4e-44
AI967315 171 4e-43
TC19546 similar to UP|Q40543 (Q40543) Protein-serine/threonine k... 160 6e-40
TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, com... 158 3e-39
TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete 154 3e-38
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 151 4e-37
TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-... 147 6e-36
TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase... 147 6e-36
TC11757 UP|Q91W27 (Q91W27) Tuberoinfundibular 39 residue protein... 140 7e-34
>TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3, complete
Length = 2481
Score = 556 bits (1434), Expect = e-159
Identities = 306/708 (43%), Positives = 409/708 (57%), Gaps = 58/708 (8%)
Frame = +1
Query: 32 ITKSASLSNGSTLVSKDGTFEMGFFRPGKSLNRYVGIWYKNIPVRRVVWVANRNNPTKDD 91
+ + S+ + TLVS +GTFE GFFR G SL RY GIWYK+I R +VWVANR+ P ++
Sbjct: 1 MAQKQSIQDDETLVSPEGTFEAGFFRFGNSLRRYFGIWYKSISPRTIVWVANRDAPVQNS 180
Query: 92 SSKLIISQDGNLVLLNHNDSLVWSTNASRKASSPVVQLLNNGNLVLRDEKDNNEESFLWQ 151
++ L ++ GNL++L+ +VWS+NASR P++QLL++GN V++D + EE+ +W+
Sbjct: 181 TATLKLTDQGNLLILDGLKGIVWSSNASRTKDKPLMQLLDSGNFVVKD--GDKEENLIWE 354
Query: 152 GFDHPCDTLLPGMTFGYNRKLDFYWNLTAWKNEDDPSSGDLYASVVFTSNPESMIWKGST 211
FD+P DT L GM N LT+W+N +DP+SG+ + P+ ++ KG+T
Sbjct: 355 SFDYPGDTFLAGMKIKSNLATGPTSYLTSWRNAEDPASGEFSYHIDTHGYPQLVVTKGAT 534
Query: 212 KICRSGPW-NPLSSGVVGMKPNPLYDYKVVNNEDEVYYQFVLRNSSVTSIAVLNQTLLIR 270
R+GPW SG G++ + + + + EV ++ N S+ + V+ +
Sbjct: 535 VTLRAGPWIGNKFSGASGLRLQKILTFSMQFTDKEVSLEYETANRSIITRTVITPS-GTT 711
Query: 271 QRLVYVPESKIWSVYQIMPSDTCEYYNVCGANAQCTIDGSPMCQCLPGFKPKSPQQWNSM 330
QRL++ S+ W + P D C YY CGAN+ C +P+C CL GF PK QWNS+
Sbjct: 712 QRLLWSDRSQSWEIISTHPMDQCAYYAFCGANSMCDTSNNPICDCLEGFTPKFQAQWNSL 891
Query: 331 DWTQGCVRGGNWSCGIKNRDGFQKFVRMKLPDTTNSWINLNMTLQDCKTKCLQNCSCTAY 390
DWT GCV N SC +N DGF K ++ PDT++SW + +L +C T CLQNCSCTAY
Sbjct: 892 DWTGGCVPIKNLSC--QNGDGFPKHTGVQFPDTSSSWYGNSKSLDECGTICLQNCSCTAY 1065
Query: 391 TYLDPNGAVSGCSLWFNDLIDL-RLSQSSEGDDLYIRV---DRDSNFGKK---------- 436
YLD G S C WF D++D+ +G ++Y+RV + D KK
Sbjct: 1066AYLDNVGGRSVCLNWFGDILDMSEHPDPDQGQEIYLRVVASELDHRRNKKSINIKKLAGS 1245
Query: 437 -----------------------------ERDGG-------------EHEDFDL-PFFDL 453
E +GG ED DL FD
Sbjct: 1246LAGSIAFIICITILGLATVTCIRRKKNEREDEGGIETRIINHWKDKRGDEDIDLATIFDF 1425
Query: 454 ATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVILCVK 513
+TI T++FS +NKLGEGGFGPVYK L +G IAVKRLS S QG +EFKNEV L +
Sbjct: 1426STISSTTNHFSESNKLGEGGFGPVYKGVLANGQEIAVKRLSNTSGQGMEEFKNEVKLIAR 1605
Query: 514 LQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIARGI 573
LQHRNLVK+LGC I DE LLIYE+M N+SLD F+F
Sbjct: 1606LQHRNLVKLLGCSIHHDEMLLIYEFMHNRSLDYFIF------------------------ 1713
Query: 574 QYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMA 633
DSRLRIIHRDLK SNILLD+EM+PKISDFG+AR+ GDQ+E KT+R++GTYGYM+
Sbjct: 1714-----DSRLRIIHRDLKTSNILLDSEMNPKISDFGLARIFTGDQVEAKTKRVMGTYGYMS 1878
Query: 634 PEYVIHGLFSIKSDVFSFGVLLLETISGKKNRTLTYHEHDHNLIWHVS 681
PEY +HG FS+KSDVFSFGV++LE ISGKK H NL+ H S
Sbjct: 1879PEYAVHGSFSVKSDVFSFGVIVLEIISGKKIGRFCDPHHHRNLLSHSS 2022
>TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2, complete
Length = 2946
Score = 534 bits (1375), Expect = e-152
Identities = 296/703 (42%), Positives = 408/703 (57%), Gaps = 57/703 (8%)
Frame = +1
Query: 34 KSASLSNGSTLVSKDGTFEMGFFRPGKSLNRYVGIWYKNIPVRRVVWVANRNNPTKDDSS 93
+ +LS G T+ DGTFE GFF + Y G+WYK+I R +VWVANR+ P ++ ++
Sbjct: 235 EQCTLSQGMTV--HDGTFEAGFFHFENPQHHYFGVWYKSISPRTIVWVANRDAPLRNSTA 408
Query: 94 KLI-ISQDGNLVLLNHNDSLVWSTNASRKASSPVVQLLNNGNLVLRDEKDNNEESFLWQG 152
+ ++ G++++ + ++WSTN SR P +QLL++GNLV +D + E+ +W+
Sbjct: 409 PTLKVTHKGSILIRDGAKGVIWSTNTSRAKEQPFMQLLDSGNLVAKD--GDKGENVIWES 582
Query: 153 FDHPCDTLLPGMTFGYNRKLDFYWNLTAWKNEDDPSSGDLYASVVFTSNPESMIWKGSTK 212
F++P DT L GM N + LT+W+N +DP+SG+ + P+ ++ KG+
Sbjct: 583 FNYPGDTFLAGMKIKSNLAIGPTSYLTSWRNSEDPASGEFSYHIDIRGFPQLVVTKGAAI 762
Query: 213 ICRSGPWNPLS-SGVVGMKPNPLYDYKVVNNEDEVYYQFVLRNSSVTSIAVLNQTLLIRQ 271
R+GPW SG G + + + + E+ ++ N S+ + V+ I Q
Sbjct: 763 TLRAGPWTGNKFSGAFGQVLQKILTFFMQFTDQEISLEYETVNRSIITREVITPLGTI-Q 939
Query: 272 RLVYVPESKIWSVYQIMPSDTCEYYNVCGANAQCTIDGSPMCQCLPGFKPKSPQQWNSMD 331
RL++ ++ W + P D C Y CGAN+ C +P+C CL GF P+ +WNS+D
Sbjct: 940 RLLWSVRNQSWEIIATRPVDQCADYVFCGANSLCDTSKNPICDCLEGFMPQFQAKWNSLD 1119
Query: 332 WTQGCVRGGNWSCGIKNRDGFQKFVRMKLPDTTNSWINLNMTLQDCKTKCLQNCSCTAYT 391
W GCV SC +N DGF K +KLPDT++SW NM+L +C+T CLQNCSCTAY
Sbjct: 1120WAGGCVSMEKLSC--QNGDGFMKHTGVKLPDTSSSWFGKNMSLDECRTLCLQNCSCTAYA 1293
Query: 392 YLDPNGAVSGCSLWFNDLIDL-RLSQSSEGDDLYIRV-----DRDSN------------- 432
LD + S C +WF D++D+ + +G ++YIRV DR N
Sbjct: 1294GLDNDVDRSVCLIWFGDILDMSKHPDPDQGQEIYIRVVASKLDRTRNKKSINTKKLAGSL 1473
Query: 433 ---------------------FGKKERDGGE--------------HEDFDL-PFFDLATI 456
KK + G E ED DL FD +TI
Sbjct: 1474VVIIAFVIFITILGLAISTCIQRKKNKRGDEGEIGIINHWKDKRGDEDIDLATIFDFSTI 1653
Query: 457 IKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVILCVKLQH 516
AT++FS +NKLGEGGFGPVYK L +G IAVKRLS S QG +EFKNE+ L +LQH
Sbjct: 1654SSATNHFSLSNKLGEGGFGPVYKGLLANGQEIAVKRLSNTSGQGMEEFKNEIKLIARLQH 1833
Query: 517 RNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIARGIQYL 576
RNLVK+ GC + DE NK + L D T+SKL+ W+ RL I++ IARG+ YL
Sbjct: 1834RNLVKLFGCSVHQDE-----NSHANKKM-KILLDSTRSKLVDWNKRLQIIDGIARGLLYL 1995
Query: 577 HQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMAPEY 636
HQDSRLRIIHRDLK SNILLD+EM+PKISDFG+AR+ GDQ+E +T+R++GTYGYM PEY
Sbjct: 1996HQDSRLRIIHRDLKTSNILLDDEMNPKISDFGLARIFIGDQVEARTKRVMGTYGYMPPEY 2175
Query: 637 VIHGLFSIKSDVFSFGVLLLETISGKKNRTLTYHEHDHNLIWH 679
+HG FSIKSDVFSFGV++LE ISGKK H NL+ H
Sbjct: 2176AVHGSFSIKSDVFSFGVIVLEIISGKKIGRFYDPHHHLNLLSH 2304
>TC18923 UP|CAE45594 (CAE45594) S-receptor kinase-like protein 1, complete
Length = 2061
Score = 327 bits (838), Expect = 4e-90
Identities = 159/399 (39%), Positives = 245/399 (60%), Gaps = 2/399 (0%)
Frame = +1
Query: 31 TITKSASLSNGSTLVSKDGTFEMGFFRPGKSLNRYVGIWYKNIPVRRVVWVANRNNPTKD 90
T+T++ S+ + TLVS +GTFE GFF G S +Y GIWYK+I R +VWVANR+ P ++
Sbjct: 31 TVTQNQSIQDDETLVSAEGTFEAGFFGLGNSQRQYFGIWYKSISPRTIVWVANRDAPVQN 210
Query: 91 DSSKLIISQDGNLVLLNHNDSLVWSTNASRKASSPVVQLLNNGNLVLRDEKDNNEESFLW 150
++ + ++ GNL++L+ + ++WS+N SR A P +QLL++GNLV++D +++ +W
Sbjct: 211 STATIKLTDKGNLLILDGSKGIIWSSNGSRAAEKPYMQLLDSGNLVVKD-GGKRKKNLIW 387
Query: 151 QGFDHPCDTLLPGMTFGYNRKLDFYWNLTAWKNEDDPSSGDLYASVVFTSNPESMIWKGS 210
+ FD+P DTLL GM N LT+W+N +DP+SG+ + P+ +I + +
Sbjct: 388 ESFDYPGDTLLAGMKIKSNLVKGPTSYLTSWRNTEDPASGEFSYLIDTRGFPQLVITRNA 567
Query: 211 TKICRSGPW-NPLSSGVVGMKPNPLYDYKVVNNEDEVYYQFVLRNSSVTSIAVLNQTLLI 269
T R+GPW L SG ++ + + + E+ ++ N S+ + AV+N +
Sbjct: 568 TAYYRAGPWTGKLFSGSSWLRLRKILTFSMQFTSQEISLEYETANRSIITRAVINPS-GT 744
Query: 270 RQRLVYVPESKIWSVYQIMPSDTCEYYNVCGANAQCTIDGSPMCQCLPGFKPKSPQQWNS 329
QRL++ S+ W + P+D C YY +CGAN+ C I +P+C CL GF+PK +WNS
Sbjct: 745 TQRLLWSDRSQSWEIISTHPTDQCTYYGLCGANSMCDISNNPICHCLEGFRPKFQAKWNS 924
Query: 330 MDWTQGCVRGGNWSCGIKNRDGFQKFVRMKLPDTTNSWINLNMTLQDCKTKCLQNCSCTA 389
DW GCV N SC +N DGF K +KLPDT++SW N +L +C T CLQNCSCT+
Sbjct: 925 FDWPGGCVPMKNLSC--QNGDGFLKHTGVKLPDTSSSWYGKNKSLDECGTLCLQNCSCTS 1098
Query: 390 YTYLDPNGAVSGCSLWFNDLIDLRLSQS-SEGDDLYIRV 427
Y YLD + S C +WF D++DL + + +G ++YI+V
Sbjct: 1099YAYLDNDIGGSACLIWFGDILDLSIHPNPDQGQEIYIKV 1215
Score = 184 bits (468), Expect = 3e-47
Identities = 89/130 (68%), Positives = 106/130 (81%)
Frame = +1
Query: 550 DPTQSKLLSWSMRLNILNAIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGM 609
D T+SKLL W+ RL I++ IARG+ YLHQDSRLRIIHRDLK SNILLDNEM+PKISDFG+
Sbjct: 1378 DSTRSKLLDWNKRLQIIDGIARGLLYLHQDSRLRIIHRDLKTSNILLDNEMNPKISDFGL 1557
Query: 610 ARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKKNRTLTY 669
AR+ GDQ+E +T+R++GTYGYM PEY +HG FSIKSDVFSFGV++LE ISGKK R
Sbjct: 1558 ARIFIGDQVEARTKRVMGTYGYMPPEYAVHGSFSIKSDVFSFGVIVLEIISGKKIRKFYD 1737
Query: 670 HEHDHNLIWH 679
H NL+ H
Sbjct: 1738 PHHHLNLLSH 1767
>TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partial (77%)
Length = 1478
Score = 207 bits (527), Expect = 5e-54
Identities = 105/229 (45%), Positives = 153/229 (65%), Gaps = 2/229 (0%)
Frame = +1
Query: 451 FDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVIL 510
F + AT NF N +GEGGFG VYK L G +AVK+LS + QG +EF EV++
Sbjct: 415 FGFRELADATRNFKEANLIGEGGFGKVYKGRLTTGEAVAVKQLSHDGRQGFQEFVMEVLM 594
Query: 511 CVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSK-LLSWSMRLNILNAI 569
L H NLV+++G C +GD++LL+YEYMP SL+ LF+ + K L+WS R+ +
Sbjct: 595 LSLLHHTNLVRLIGYCTDGDQRLLVYEYMPMGSLEDHLFELSHDKEPLNWSTRMKVAVGA 774
Query: 570 ARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCG-GDQIEGKTRRIVGT 628
ARG++YLH + +I+RDLK++NILLDNE +PK+SDFG+A++ GD T R++GT
Sbjct: 775 ARGLEYLHCTADPPVIYRDLKSANILLDNEFNPKLSDFGLAKLGPVGDNTHVST-RVMGT 951
Query: 629 YGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKKNRTLTYHEHDHNLI 677
YGY APEY + G ++KSD++SFGV+LLE ++G++ + + NL+
Sbjct: 952 YGYCAPEYAMSGKLTLKSDIYSFGVVLLELLTGRRAIDTSRRPGEQNLV 1098
>BP048842
Length = 524
Score = 201 bits (512), Expect = 2e-52
Identities = 97/147 (65%), Positives = 120/147 (80%)
Frame = -3
Query: 535 IYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIARGIQYLHQDSRLRIIHRDLKASNI 594
IYE+M N+SL+ F+FD T+SKL+ W+ RL I++ IARG+ YLHQDSRLRIIHRDLK SNI
Sbjct: 522 IYEFMHNRSLNYFIFDSTRSKLVDWNKRLQIIDGIARGLLYLHQDSRLRIIHRDLKTSNI 343
Query: 595 LLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFSIKSDVFSFGVL 654
LLD+EM+PKISDFG+AR+ GDQ+E +T+R++GTYGYM PEY +HG FSIKSDVFSFGV+
Sbjct: 342 LLDDEMNPKISDFGLARIFIGDQVEARTKRVMGTYGYMPPEYAVHGSFSIKSDVFSFGVI 163
Query: 655 LLETISGKKNRTLTYHEHDHNLIWHVS 681
+LE ISGKK H NL+ HVS
Sbjct: 162 VLEIISGKKIGRFYDPHHHLNLLSHVS 82
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 192 bits (489), Expect = 1e-49
Identities = 104/214 (48%), Positives = 138/214 (63%), Gaps = 1/214 (0%)
Frame = +1
Query: 451 FDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVIL 510
F L + AT+NF+ +NKLGEGGFG VY L DG IAVKRL S + EF EV +
Sbjct: 28 FSLKELHSATNNFNYDNKLGEGGFGSVYWGQLWDGSQIAVKRLKVWSNKADMEFAVEVEI 207
Query: 511 CVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSK-LLSWSMRLNILNAI 569
+++H+NL+ + G C EG E+L++Y+YMPN SL S L S+ LL W+ R+NI
Sbjct: 208 LARVRHKNLLSLRGYCAEGQERLIVYDYMPNLSLLSHLHGQHSSECLLDWNRRMNIAIGS 387
Query: 570 ARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTY 629
A GI YLH + IIHRD+KASN+LLD++ +++DFG A++ D T R+ GT
Sbjct: 388 AEGIVYLHHQATPHIIHRDIKASNVLLDSDFQARVADFGFAKLI-PDGATHVTTRVKGTL 564
Query: 630 GYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKK 663
GY+APEY + G + DVFSFG+LLLE SGKK
Sbjct: 565 GYLAPEYAMLGKANECCDVFSFGILLLELASGKK 666
>BP032158
Length = 555
Score = 192 bits (487), Expect = 2e-49
Identities = 97/182 (53%), Positives = 129/182 (70%), Gaps = 1/182 (0%)
Frame = +2
Query: 450 FFDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVI 509
+F L I AT+NF NK+GEGGFGPVYK L +G VIAVK+LS S+QG++EF NE+
Sbjct: 11 YFSLRQIKAATNNFDPANKIGEGGFGPVYKGVLSEGDVIAVKQLSSKSKQGNREFINEIG 190
Query: 510 LCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKL-LSWSMRLNILNA 568
+ LQH NLVK+ GCCIEG++ LL+YEYM N SL LF + KL L+W R+ I
Sbjct: 191 MISALQHPNLVKLYGCCIEGNQLLLVYEYMENNSLARALFGNEEQKLNLNWRTRMKICVG 370
Query: 569 IARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGT 628
IA+G+ YLH++SRL+I+HRD+KA+N+LLD ++ KISDFG+A++ + T RI GT
Sbjct: 371 IAKGLAYLHEESRLKIVHRDIKATNVLLDKDLKAKISDFGLAKLDEEENTHIST-RIAGT 547
Query: 629 YG 630
G
Sbjct: 548 IG 553
>TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine protein kinase
Cdk9 (Cyclin-dependent kinase Cdk9) , partial (7%)
Length = 723
Score = 184 bits (467), Expect = 4e-47
Identities = 90/172 (52%), Positives = 116/172 (67%)
Frame = +3
Query: 451 FDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVIL 510
F T++ AT NF NKLGEGGFGPV+K L DG IAVK+LS S QG +F NE L
Sbjct: 207 FSYETLVAATKNFHAVNKLGEGGFGPVFKGKLNDGREIAVKKLSRRSNQGRTQFINEAKL 386
Query: 511 CVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIA 570
++QHRN+V + G C G EKLL+YEY+P +SLD LF + + L W R +I++ +A
Sbjct: 387 LTRVQHRNVVSLFGYCAHGSEKLLVYEYVPRESLDKLLFRSQKKEQLDWKRRFDIISGVA 566
Query: 571 RGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKT 622
RG+ YLH+DS IIHRD+KA+NILLD + PKI+DFG+AR+ DQ T
Sbjct: 567 RGLLYLHEDSHDCIIHRDIKAANILLDEKWVPKIADFGLARIFPEDQTHVNT 722
>TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete
Length = 3495
Score = 181 bits (460), Expect = 3e-46
Identities = 98/214 (45%), Positives = 135/214 (62%), Gaps = 1/214 (0%)
Frame = +3
Query: 451 FDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVIL 510
F L I AT+ + T +GEGGFG VY+ TL DG +AVK S S QG++EF NE+ L
Sbjct: 2133 FTLEYIEVATERYKT--LIGEGGFGSVYRGTLNDGQEVAVKVRSSTSTQGTREFDNELNL 2306
Query: 511 CVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLF-DPTQSKLLSWSMRLNILNAI 569
+QH NLV +LG C E D+++L+Y +M N SL L+ +P + K+L W RL+I
Sbjct: 2307 LSAIQHENLVPLLGYCNESDQQILVYPFMSNGSLQDRLYGEPAKRKILDWPTRLSIALGA 2486
Query: 570 ARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTY 629
ARG+ YLH +IHRD+K+SNILLD+ M K++DFG ++ + + + GT
Sbjct: 2487 ARGLAYLHTFPGRSVIHRDIKSSNILLDHSMCAKVADFGFSKYAPQEGDSYVSLEVRGTA 2666
Query: 630 GYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKK 663
GY+ PEY S KSDVFSFGV+LLE +SG++
Sbjct: 2667 GYLDPEYYKTQQLSEKSDVFSFGVVLLEIVSGRE 2768
>CN825263
Length = 663
Score = 178 bits (452), Expect = 2e-45
Identities = 92/206 (44%), Positives = 133/206 (63%), Gaps = 1/206 (0%)
Frame = +1
Query: 473 GFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVILCVKLQHRNLVKVLGCCIEGDEK 532
GFG VYK L DG +AVK L + ++G +EF EV + +L HRNLVK++G CIE +
Sbjct: 1 GFGLVYKGILNDGRDVAVKILKRDDQRGGREFLAEVEMLSRLHHRNLVKLIGICIEKQTR 180
Query: 533 LLIYEYMPNKSLDSFLFDPT-QSKLLSWSMRLNILNAIARGIQYLHQDSRLRIIHRDLKA 591
LIYE +PN S++S L ++ L W+ R+ I ARG+ YLH+DS +IHRD K+
Sbjct: 181 CLIYELVPNGSVESHLHGADKETGPLDWNARMKIALGAARGLAYLHEDSNPCVIHRDFKS 360
Query: 592 SNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFSIKSDVFSF 651
SNILL+ + PK+SDFG+AR + + + ++GT+GY+APEY + G +KSDV+S+
Sbjct: 361 SNILLECDFTPKVSDFGLARTALDEGNKHISTHVMGTFGYLAPEYAMTGHLLVKSDVYSY 540
Query: 652 GVLLLETISGKKNRTLTYHEHDHNLI 677
GV+LLE ++G K L+ NL+
Sbjct: 541 GVVLLELLTGTKPVDLSQPPGQENLV 618
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 177 bits (448), Expect = 7e-45
Identities = 91/206 (44%), Positives = 133/206 (64%), Gaps = 1/206 (0%)
Frame = +2
Query: 459 ATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVILCVKLQHRN 518
AT FS +NKLGEGGFG VY DG IAVK+L + + EF EV + +++H+N
Sbjct: 290 ATGGFSDDNKLGEGGFGSVYWGRTSDGLQIAVKKLKAMNSKAEMEFAVEVEVLGRVRHKN 469
Query: 519 LVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKL-LSWSMRLNILNAIARGIQYLH 577
L+ + G C+ D++L++Y+YMPN SL S L ++ L+W R+ I A GI YLH
Sbjct: 470 LLGLRGYCVGDDQRLIVYDYMPNLSLLSHLHGQFAVEVQLNWQKRMKIAIGSAEGILYLH 649
Query: 578 QDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYV 637
+ IIHRD+KASN+LL+++ +P ++DFG A++ + + T R+ GT GY+APEY
Sbjct: 650 HEVTPHIIHRDIKASNVLLNSDFEPLVADFGFAKLI-PEGVSHMTTRVKGTLGYLAPEYA 826
Query: 638 IHGLFSIKSDVFSFGVLLLETISGKK 663
+ G S DV+SFG+LLLE ++G+K
Sbjct: 827 MWGKVSESCDVYSFGILLLELVTGRK 904
>AV422094
Length = 462
Score = 174 bits (441), Expect = 4e-44
Identities = 86/150 (57%), Positives = 111/150 (73%)
Frame = +2
Query: 451 FDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVIL 510
F + + AT++F+ +NKLGEGGFGPVYK L DG VIAVK+LS S QG +F E+
Sbjct: 14 FSYSELKNATNDFNIDNKLGEGGFGPVYKGILNDGTVIAVKQLSLGSHQGKSQFIAEIAT 193
Query: 511 CVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIA 570
+QHRNLVK+ GCCIEG ++LL+YEY+ NKSLD LF S L+WS R +I +A
Sbjct: 194 ISAVQHRNLVKLYGCCIEGSKRLLVYEYLENKSLDQALFGKALS--LNWSTRYDICLGVA 367
Query: 571 RGIQYLHQDSRLRIIHRDLKASNILLDNEM 600
RG+ YLH++SRLRI+HRD+KASNILLD+E+
Sbjct: 368 RGLTYLHEESRLRIVHRDVKASNILLDHEL 457
>AI967315
Length = 1308
Score = 171 bits (433), Expect = 4e-43
Identities = 100/249 (40%), Positives = 144/249 (57%), Gaps = 16/249 (6%)
Frame = +1
Query: 431 SNFGKKERDGGEHEDFDL---PF-----------FDLATIIKATDNFSTNNKLGEGGFGP 476
S+FG++ GGE++D L P+ F + AT+ FS+ N +G+GG+
Sbjct: 13 SSFGRRSF-GGENDDSSLVVPPYEEHPPRPSWKCFSYEELFHATNGFSSENMVGKGGYAE 189
Query: 477 VYKATLQDGHVIAVKRLSGN--SEQGSKEFKNEVILCVKLQHRNLVKVLGCCIEGDEKLL 534
VYK L+ G IAVKRL+ E+ KEF E+ + H N++ +LGCCI+ L
Sbjct: 190 VYKGRLESGDEIAVKRLTRTCRDERKEKEFLTEIGTIGHVCHSNVMPLLGCCIDNG-LYL 366
Query: 535 IYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIARGIQYLHQDSRLRIIHRDLKASNI 594
++E S+ S + D + L W R I+ ARG+ YLH+ + RIIHRD+KASNI
Sbjct: 367 VFELSTVGSVASLIHDEKMAPL-DWKTRYKIVLGTARGLHYLHKGCQRRIIHRDIKASNI 543
Query: 595 LLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFSIKSDVFSFGVL 654
LL + +P+ISDFG+A+ I GT+G++APEY +HG+ K+DVF+FGV
Sbjct: 544 LLTEDFEPQISDFGLAKWLPSQWTHHSIAPIEGTFGHLAPEYYMHGVVDEKTDVFAFGVF 723
Query: 655 LLETISGKK 663
LLE ISG+K
Sbjct: 724 LLEVISGRK 750
>TC19546 similar to UP|Q40543 (Q40543) Protein-serine/threonine kinase ,
partial (46%)
Length = 731
Score = 160 bits (405), Expect = 6e-40
Identities = 84/187 (44%), Positives = 124/187 (65%)
Frame = +1
Query: 448 LPFFDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNE 507
+P + I KAT NF+T LG+G FG VYKAT+ G V+AVK L+ NS+QG EF+ E
Sbjct: 190 IPKYSYKDIQKATQNFTTI--LGQGSFGTVYKATMPTGEVVAVKVLAPNSKQGEHEFQTE 363
Query: 508 VILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILN 567
V L +L HRNLV ++G C++ +++L+Y++M N SL + L+ + K LSW RL I
Sbjct: 364 VHLLGRLHHRNLVNLVGFCVDKGQRILVYQFMSNGSLANLLYG--EEKELSWDERLQIAM 537
Query: 568 AIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVG 627
I+ GI+YLH+ + +IHRDLK++NILLD+ M ++DFG+++ + +G+ + G
Sbjct: 538 DISHGIEYLHEGAVPPVIHRDLKSANILLDDSMRAMVADFGLSK---EEIFDGRNSGLKG 708
Query: 628 TYGYMAP 634
TYGYM P
Sbjct: 709 TYGYMDP 729
>TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1591
Score = 158 bits (399), Expect = 3e-39
Identities = 84/224 (37%), Positives = 134/224 (59%), Gaps = 7/224 (3%)
Frame = +2
Query: 447 DLPFFDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSE-QGSKEFK 505
++P L + + TDNF + +GEG +G VY ATL DG+ +AVK+L +SE + + EF
Sbjct: 311 EVPALSLDELKEKTDNFGSKALIGEGSYGRVYYATLNDGNAVAVKKLDVSSEPETNNEFL 490
Query: 506 NEVILCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFD------PTQSKLLSW 559
+V + +L++ N V++ G C+EG+ ++L YE+ SL L L W
Sbjct: 491 TQVSMVSRLKNDNFVELHGYCVEGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDW 670
Query: 560 SMRLNILNAIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIE 619
R+ I ARG++YLH+ + IIHRD+++SN+L+ + KI+DF ++
Sbjct: 671 IQRVRIAVDAARGLEYLHEKVQPAIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAAR 850
Query: 620 GKTRRIVGTYGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGKK 663
+ R++GT+GY APEY + G + KSDV+SFGV+LLE ++G+K
Sbjct: 851 LHSTRVLGTFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRK 982
>TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete
Length = 2193
Score = 154 bits (390), Expect = 3e-38
Identities = 87/214 (40%), Positives = 133/214 (61%), Gaps = 2/214 (0%)
Frame = +1
Query: 451 FDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVIL 510
F + KAT+NFS +NK+G+GGFG VY A L+ G A+K++ Q S EF E+ +
Sbjct: 1051 FSYQELAKATNNFSLDNKIGQGGFGAVYYAELR-GKKTAIKKMD---VQASTEFLCELKV 1218
Query: 511 CVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIA 570
+ H NLV+++G C+EG L+YE++ N +L +L + L WS R+ I A
Sbjct: 1219 LTHVHHLNLVRLIGYCVEGS-LFLVYEHIDNGNLGQYLHGSGKEPL-PWSSRVQIALDAA 1392
Query: 571 RGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARM--CGGDQIEGKTRRIVGT 628
RG++Y+H+ + IHRD+K++NIL+D + K++DFG+ ++ G ++ R+VGT
Sbjct: 1393 RGLEYIHEHTVPVYIHRDVKSANILIDKNLRGKVADFGLTKLIEVGNSTLQ---TRLVGT 1563
Query: 629 YGYMAPEYVIHGLFSIKSDVFSFGVLLLETISGK 662
+GYM PEY +G S K DV++FGV+L E IS K
Sbjct: 1564 FGYMPPEYAQYGDISPKIDVYAFGVVLFELISAK 1665
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 151 bits (381), Expect = 4e-37
Identities = 83/198 (41%), Positives = 119/198 (59%), Gaps = 1/198 (0%)
Frame = +2
Query: 467 NKLGEGGFGPVYKATLQDGHVIAVKRLSGN-SEQGSKEFKNEVILCVKLQHRNLVKVLGC 525
N +G+GG G VY+ ++ +G +A+KRL G S + F+ E+ K++HRN++++LG
Sbjct: 2204 NIIGKGGAGIVYRGSMPNGTDVAIKRLVGQGSGRNDYGFRAEIETLGKIRHRNIMRLLGY 2383
Query: 526 CIEGDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIARGIQYLHQDSRLRII 585
D LL+YEYMPN SL +L + L W MR I ARG+ Y+H D II
Sbjct: 2384 VSNKDTNLLLYEYMPNGSLGEWLHG-AKGGHLRWEMRYKIAVEAARGLCYMHHDCSPLII 2560
Query: 586 HRDLKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFSIK 645
HRD+K++NILLD + + ++DFG+A+ I G+YGY+APEY K
Sbjct: 2561 HRDVKSNNILLDADFEAHVADFGLAKFLYDPGASQSMSSIAGSYGYIAPEYAYTLKVDEK 2740
Query: 646 SDVFSFGVLLLETISGKK 663
SDV+SFGV+LLE I G+K
Sbjct: 2741 SDVYSFGVVLLELIIGRK 2794
>TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-like
protein, partial (46%)
Length = 941
Score = 147 bits (371), Expect = 6e-36
Identities = 79/194 (40%), Positives = 116/194 (59%)
Frame = +3
Query: 469 LGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVILCVKLQHRNLVKVLGCCIE 528
LG GGFG VY + G +A+KR + SEQG EF+ E+ + KL+HR+LV ++G C E
Sbjct: 15 LGVGGFGKVYYGEVDGGTKVAIKRGNPLSEQGVHEFQTEIEMLSKLRHRHLVSLIGYCEE 194
Query: 529 GDEKLLIYEYMPNKSLDSFLFDPTQSKLLSWSMRLNILNAIARGIQYLHQDSRLRIIHRD 588
E +L+Y++M +L L+ TQ L W RL I ARG+ YLH ++ IIHRD
Sbjct: 195 NTEMILVYDHMAYGTLREHLY-KTQKPPLPWKQRLEICIGAARGLHYLHTGAKYTIIHRD 371
Query: 589 LKASNILLDNEMDPKISDFGMARMCGGDQIEGKTRRIVGTYGYMAPEYVIHGLFSIKSDV 648
+K +NILLD + K+SDFG+++ + + G++GY+ PEY + KSDV
Sbjct: 372 VKTTNILLDEKWVAKVSDFGLSKTGPTLDNTHVSTVVKGSFGYLDPEYFRRQQLTDKSDV 551
Query: 649 FSFGVLLLETISGK 662
+SFGV+L E + +
Sbjct: 552 YSFGVVLFEILCAR 593
>TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase, partial
(30%)
Length = 728
Score = 147 bits (371), Expect = 6e-36
Identities = 76/190 (40%), Positives = 122/190 (64%), Gaps = 5/190 (2%)
Frame = +3
Query: 451 FDLATIIKATDNFSTNNKLGEGGFGPVYKATLQDGHV-IAVKRLSGNSEQGSKEFKNEVI 509
F+ + KAT+NF +KLG+GG+G VY+ L + +AVK S + + + +F +E+I
Sbjct: 132 FNYVELKKATNNFDEKHKLGQGGYGVVYRGMLPKEKLEVAVKMFSRDKMKSTDDFLSELI 311
Query: 510 LCVKLQHRNLVKVLGCCIEGDEKLLIYEYMPNKSLDSFLF---DPTQSKLLSWSMRLNIL 566
+ +L+H++LV++ G C + LL+Y+YMPN SLDS +F T + LSW +R I+
Sbjct: 312 IINRLRHKHLVRLQGWCHKNGVLLLVYDYMPNGSLDSHIFCEEGGTITTPLSWPLRYKII 491
Query: 567 NAIARGIQYLHQDSRLRIIHRDLKASNILLDNEMDPKISDFGMARMCGGDQIE-GKTRRI 625
+ +A + YLH + +++HRDLKASNI+LD+E + K+ DFG+AR ++I + +
Sbjct: 492 SGVASALNYLHNEYDQKVVHRDLKASNIMLDSEFNAKLGDFGLARALENEKISYTELEGV 671
Query: 626 VGTYGYMAPE 635
GT GY+APE
Sbjct: 672 HGTMGYIAPE 701
>TC11757 UP|Q91W27 (Q91W27) Tuberoinfundibular 39 residue protein, partial
(10%)
Length = 715
Score = 140 bits (353), Expect = 7e-34
Identities = 67/136 (49%), Positives = 91/136 (66%), Gaps = 1/136 (0%)
Frame = +2
Query: 459 ATDNFSTNNKLGEGGFGPVYKATLQDGHVIAVKRLSGNSEQGSKEFKNEVILCVKLQHRN 518
ATDNFS NK+GEGGFGPVYK L+DG V A+K LS S+QG KEF E+ + +++H N
Sbjct: 308 ATDNFSPVNKIGEGGFGPVYKGVLKDGKVGAIKVLSAESKQGVKEFMTEINVISEIEHEN 487
Query: 519 LVKVLGCCIEGDEKLLIYEYMPNKSLDSFLFDPTQSKL-LSWSMRLNILNAIARGIQYLH 577
LV++ GCC+EG+ ++L+Y Y N SL L S + W R I +ARG+ +LH
Sbjct: 488 LVQLYGCCVEGNHRILVYNYHENNSLAQTLLGGGHSNIHFDWRTRSRICIGVARGLSFLH 667
Query: 578 QDSRLRIIHRDLKASN 593
++ R I+HRD+KASN
Sbjct: 668 EEVRPHIVHRDIKASN 715
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.137 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,418,532
Number of Sequences: 28460
Number of extensions: 209085
Number of successful extensions: 1427
Number of sequences better than 10.0: 270
Number of HSP's better than 10.0 without gapping: 1251
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1265
length of query: 685
length of database: 4,897,600
effective HSP length: 96
effective length of query: 589
effective length of database: 2,165,440
effective search space: 1275444160
effective search space used: 1275444160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146866.11


