

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146864.6 - phase: 0 /pseudo
(180 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
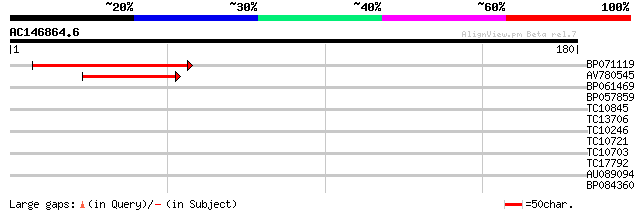
Score E
Sequences producing significant alignments: (bits) Value
BP071119 60 2e-10
AV780545 42 6e-05
BP061469 34 0.013
BP057859 28 1.2
TC10845 weakly similar to UP|WRK6_ARATH (Q9C519) WRKY transcript... 27 1.6
TC13706 similar to UP|AAH60251 (AAH60251) SERTA domain containin... 27 2.1
TC10246 26 3.5
TC10721 similar to UP|MTD_MEDSA (O82515) Probable mannitol dehyd... 26 4.6
TC10703 similar to UP|BAD09184 (BAD09184) Eukaryotic translation... 25 7.8
TC17792 25 7.8
AU089094 25 7.8
BP084360 25 7.8
>BP071119
Length = 412
Score = 60.5 bits (145), Expect = 2e-10
Identities = 28/51 (54%), Positives = 36/51 (69%)
Frame = +2
Query: 8 IRASGLYLLLETNYGQVDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVS 58
+ +GL +++ VD ++ AF ERWH ETSSFHLP GEMTITL+DVS
Sbjct: 257 VHNAGLRWVMKCTSPSVDQSIISAFVERWHPETSSFHLPWGEMTITLEDVS 409
>AV780545
Length = 555
Score = 42.0 bits (97), Expect = 6e-05
Identities = 18/31 (58%), Positives = 24/31 (77%)
Frame = +1
Query: 24 VDHGLLIAFSERWHSETSSFHLPVGEMTITL 54
+D+ L+ A ERW ET++FHL VGEMT+TL
Sbjct: 463 LDNPLISALVERWRRETNTFHLNVGEMTVTL 555
>BP061469
Length = 482
Score = 34.3 bits (77), Expect = 0.013
Identities = 16/40 (40%), Positives = 26/40 (65%)
Frame = -2
Query: 28 LLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGG 67
LL + W SE +F + GE++++L DV+ +L +PVGG
Sbjct: 217 LLTEVMKAWDSERQAFAVGSGEISMSLLDVALILGMPVGG 98
>BP057859
Length = 496
Score = 27.7 bits (60), Expect = 1.2
Identities = 15/39 (38%), Positives = 21/39 (53%)
Frame = -3
Query: 142 RCIRAFLLYLVGCTLFSDKAGNSCCVVYLKYFDDLTTVN 180
+ +R FLL L T+F N CC V + Y+ D T+ N
Sbjct: 263 KSLRPFLLPLK--TIFIADKSNFCCSV*MNYYSDSTSSN 153
>TC10845 weakly similar to UP|WRK6_ARATH (Q9C519) WRKY transcription
factor 6 (WRKY DNA-binding protein 6) (AtWRKY6),
partial (17%)
Length = 635
Score = 27.3 bits (59), Expect = 1.6
Identities = 15/38 (39%), Positives = 23/38 (60%)
Frame = +1
Query: 38 SETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHESL 75
S +S +L V + T D VS L+H+P+ +L FH +L
Sbjct: 34 SLSSCTYLLVYLILWTKDGVSLLIHLPLNPSLSFHPNL 147
>TC13706 similar to UP|AAH60251 (AAH60251) SERTA domain containing 4,
partial (6%)
Length = 523
Score = 26.9 bits (58), Expect = 2.1
Identities = 16/76 (21%), Positives = 33/76 (43%)
Frame = +3
Query: 27 GLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHESLSIHQGTQYLVN 86
G++++ ++ HSET + L + + L +PV G + + G +V
Sbjct: 162 GIVLSRTQPSHSETRALSLSFSSFILRRTQIQIFLRVPVAGGWAVGVLVGVGVG*WLVVM 341
Query: 87 YLGLEFEESAAQTKRL 102
L+ +E + K+L
Sbjct: 342 RKKLKSKEEVMRKKKL 389
>TC10246
Length = 525
Score = 26.2 bits (56), Expect = 3.5
Identities = 10/30 (33%), Positives = 18/30 (59%)
Frame = -2
Query: 143 CIRAFLLYLVGCTLFSDKAGNSCCVVYLKY 172
C+ Y+ G ++KAG++ CV +LK+
Sbjct: 167 CVERVPKYIFGSIHAAEKAGSAACVSHLKH 78
>TC10721 similar to UP|MTD_MEDSA (O82515) Probable mannitol dehydrogenase
(NAD-dependent mannitol dehydrogenase) , complete
Length = 1297
Score = 25.8 bits (55), Expect = 4.6
Identities = 8/22 (36%), Positives = 14/22 (63%)
Frame = -2
Query: 122 EAKSYANQPEEEDSMEWYRTRC 143
EA+S QP+ ++S W+ + C
Sbjct: 522 EARSAGEQPQNQNSHHWFASHC 457
>TC10703 similar to UP|BAD09184 (BAD09184) Eukaryotic translation initiation
factor 3 subunit (EIF-3)-like, partial (16%)
Length = 1066
Score = 25.0 bits (53), Expect = 7.8
Identities = 10/21 (47%), Positives = 15/21 (70%)
Frame = +1
Query: 21 YGQVDHGLLIAFSERWHSETS 41
YG VD+G I ++E +HS+ S
Sbjct: 469 YGSVDNGKKIGWNEDFHSKVS 531
>TC17792
Length = 665
Score = 25.0 bits (53), Expect = 7.8
Identities = 19/65 (29%), Positives = 32/65 (49%)
Frame = +1
Query: 111 TLLSIYTSYLTEAKSYANQPEEEDSMEWYRTRCIRAFLLYLVGCTLFSDKAGNSCCVVYL 170
T+ I+T + E+K+ EE ++ T ++ FLL GCT+ + +SC V +
Sbjct: 193 TVGMIWTGDMPESKAV-----EESMASYFNT--LQGFLLLSHGCTVGAGPTLSSCIHVSV 351
Query: 171 KYFDD 175
K D
Sbjct: 352 KQVVD 366
>AU089094
Length = 639
Score = 25.0 bits (53), Expect = 7.8
Identities = 11/29 (37%), Positives = 17/29 (57%)
Frame = +1
Query: 6 NVIRASGLYLLLETNYGQVDHGLLIAFSE 34
+VI+A + L T YG DHG+ +A +
Sbjct: 166 SVIKAFNMSDRLTTEYGNKDHGIKMALDD 252
>BP084360
Length = 381
Score = 25.0 bits (53), Expect = 7.8
Identities = 9/28 (32%), Positives = 18/28 (64%)
Frame = +1
Query: 80 GTQYLVNYLGLEFEESAAQTKRLRSAHI 107
G + + NY ++F+++ A RLR+ H+
Sbjct: 226 GPESVTNYQEMKFQQTCADFDRLRTFHL 309
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.137 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,540,818
Number of Sequences: 28460
Number of extensions: 47859
Number of successful extensions: 220
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 220
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 220
length of query: 180
length of database: 4,897,600
effective HSP length: 84
effective length of query: 96
effective length of database: 2,506,960
effective search space: 240668160
effective search space used: 240668160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146864.6


