

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146863.4 + phase: 0
(309 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
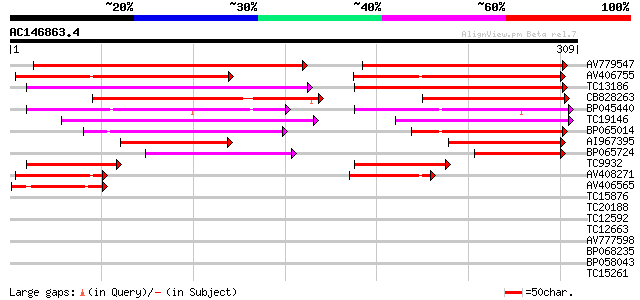
Score E
Sequences producing significant alignments: (bits) Value
AV779547 173 4e-44
AV406755 121 1e-28
TC13186 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidat... 108 1e-24
CB828263 107 3e-24
BP045440 96 6e-21
TC19146 similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-contain... 93 5e-20
BP065014 81 3e-16
AI967395 57 5e-09
BP065724 54 4e-08
TC9932 weakly similar to UP|Q93YA7 (Q93YA7) Resistance gene-like... 54 4e-08
AV408271 50 6e-07
AV406565 42 1e-04
TC15876 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-... 37 0.003
TC20188 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-... 37 0.006
TC12592 35 0.012
TC12663 weakly similar to UP|Q84ZV8 (Q84ZV8) R 3 protein, partia... 34 0.027
AV777598 31 0.23
BP068235 31 0.30
BP058043 28 1.5
TC15261 weakly similar to UP|Q9ZQ55 (Q9ZQ55) At2g22570 protein, ... 28 2.0
>AV779547
Length = 556
Score = 173 bits (438), Expect = 4e-44
Identities = 90/149 (60%), Positives = 110/149 (73%)
Frame = +1
Query: 14 VSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIVVFSK 73
VSFRG DTR NFTDHL GAL K TF+DDT L+KG +I+++LIQAIEGSQ+ IVVFSK
Sbjct: 109 VSFRGEDTRNNFTDHLLGALYGKGFVTFKDDTMLRKGKNISTELIQAIEGSQILIVVFSK 288
Query: 74 NYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRFQDHS 133
YASSTWCL+ELA I + V + VLPV DV SEVRKQSG YGE+F H +RF++
Sbjct: 289 YYASSTWCLQELAKIADGIVGKRQTVLPVVCDVTPSEVRKQSGNYGEAFLKHEERFKEDL 468
Query: 134 NMVQRRRETLQLVGNISGWDLRDKPHHAE 162
MVQR R+ L V ++SGWD+ +KP H +
Sbjct: 469 EMVQRWRKALAEVADLSGWDVTNKPEHEQ 555
Score = 124 bits (312), Expect = 2e-29
Identities = 66/112 (58%), Positives = 77/112 (67%)
Frame = +1
Query: 193 VSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIVVFSK 252
VSFRG DTR NFTDH AL +G F+DDT L+KG+ I+ L +AIE SQ+ IVVFSK
Sbjct: 109 VSFRGEDTRNNFTDHLLGALYGKGFVTFKDDTMLRKGKNISTELIQAIEGSQILIVVFSK 288
Query: 253 NYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
YASSTWCL+EL I + VLPV DV PSEV+KQSG YG+A KH
Sbjct: 289 YYASSTWCLQELAKIADGIVGKRQTVLPVVCDVTPSEVRKQSGNYGEAFLKH 444
>AV406755
Length = 425
Score = 121 bits (304), Expect = 1e-28
Identities = 57/116 (49%), Positives = 82/116 (70%)
Frame = +3
Query: 188 NYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYI 247
+YDVF+SFRG ++R +FT H AL+ GI F D+ +L++GE I+ L +AIE S++ I
Sbjct: 81 SYDVFLSFRGEESRRSFTSHLYTALKNAGIKVFMDN-ELQRGEDISSSLLKAIEDSRIAI 257
Query: 248 VVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSK 303
++FS NY S WCL ELE I+ C + G+ V+PVFY+VDPS+++KQ G G+A K
Sbjct: 258 IIFSTNYTGSKWCLDELEKIIECQRTIGQEVMPVFYNVDPSDIRKQRGTVGEAFRK 425
Score = 117 bits (292), Expect = 3e-27
Identities = 57/119 (47%), Positives = 81/119 (67%)
Frame = +3
Query: 4 PRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEG 63
P+ K + VF+SFRG ++R +FT HL+ AL+ I F D+ LQ+G I+S L++AIE
Sbjct: 66 PKSKWSYDVFLSFRGEESRRSFTSHLYTALKNAGIKVFMDN-ELQRGEDISSSLLKAIED 242
Query: 64 SQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESF 122
S++ I++FS NY S WCL EL I+ C G+ V+PVFY+VD S++RKQ G GE+F
Sbjct: 243 SRIAIIIFSTNYTGSKWCLDELEKIIECQRTIGQEVMPVFYNVDPSDIRKQRGTVGEAF 419
>TC13186 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidate
resistance protein KR1, partial (6%)
Length = 586
Score = 108 bits (269), Expect = 1e-24
Identities = 62/156 (39%), Positives = 92/156 (58%)
Frame = +1
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIV 69
+ VF+S +DTRF F+ L+ AL K I TF DD + + +++ +AIE S++ IV
Sbjct: 103 YDVFISLITSDTRFGFSGDLYTALSDKGILTFIDDGLPRPDDIMSTVNSKAIESSRIAIV 282
Query: 70 VFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRF 129
V S+NYASS++CL EL YIL G + PVFY V+ S+VR Q GG+G++F H +RF
Sbjct: 283 VISENYASSSFCLDELMYILKRREEKGLLIWPVFYQVEPSDVRYQKGGFGKAFASHEERF 462
Query: 130 QDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELEN 165
S V++ R L+ V ++ + +H L N
Sbjct: 463 GSDSAKVRKWRNALKQVADLKAFHFFKHGYHTSLIN 570
Score = 100 bits (248), Expect = 4e-22
Identities = 52/116 (44%), Positives = 74/116 (62%)
Frame = +1
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIV 248
YDVF+S DTRF F+ AL +GI F DD + + ++ +AIE+S++ IV
Sbjct: 103 YDVFISLITSDTRFGFSGDLYTALSDKGILTFIDDGLPRPDDIMSTVNSKAIESSRIAIV 282
Query: 249 VFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
V S+NYASS++CL EL YIL ++ G + PVFY V+PS+V+ Q GG+G A + H
Sbjct: 283 VISENYASSSFCLDELMYILKRREEKGLLIWPVFYQVEPSDVRYQKGGFGKAFASH 450
>CB828263
Length = 570
Score = 107 bits (267), Expect = 3e-24
Identities = 56/131 (42%), Positives = 84/131 (63%), Gaps = 5/131 (3%)
Frame = +2
Query: 46 NLQKGNSIASDLIQAIEGSQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYD 105
+LQ+G+ I+S L++AIE +++ ++VFSKNY +S WCL EL IL C G+ VLPVFYD
Sbjct: 5 DLQRGDEISSSLLRAIEEAKLSVIVFSKNYGNSKWCLDELVKILECKRTRGQIVLPVFYD 184
Query: 106 VDLSEVRKQSGGYGESFNYHGKRFQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAEL-- 163
+D S VR Q+G Y E+F HG+ + VQ+ RE L+ N+SGWD +E+
Sbjct: 185 IDPSHVRNQTGTYAEAFVKHGQ-----VDKVQKWREALREAANLSGWDCSVNRMESEVIE 349
Query: 164 ---ENIIEHIN 171
++++E +N
Sbjct: 350 KIAKDVLEKLN 382
Score = 94.0 bits (232), Expect = 3e-20
Identities = 42/80 (52%), Positives = 58/80 (72%)
Frame = +2
Query: 226 LKKGEFIAPGLFRAIEASQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDV 285
L++G+ I+ L RAIE +++ ++VFSKNY +S WCL EL IL C + G+ VLPVFYD+
Sbjct: 8 LQRGDEISSSLLRAIEEAKLSVIVFSKNYGNSKWCLDELVKILECKRTRGQIVLPVFYDI 187
Query: 286 DPSEVQKQSGGYGDALSKHG 305
DPS V+ Q+G Y +A KHG
Sbjct: 188 DPSHVRNQTGTYAEAFVKHG 247
>BP045440
Length = 535
Score = 96.3 bits (238), Expect = 6e-21
Identities = 54/121 (44%), Positives = 70/121 (57%), Gaps = 2/121 (1%)
Frame = -3
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIV 248
Y +F+SFRG DTR +FT L R G F+DD L+ G I L IE S+ I+
Sbjct: 527 YQIFISFRGMDTRDDFTKPLYQGLCREGFVTFKDDESLEGGSPIEK-LLDDIEESRFAII 351
Query: 249 VFSKNYASSTWCLRELEYILHCSKKHGKH--VLPVFYDVDPSEVQKQSGGYGDALSKHGL 306
+ SKNYA S WCL+EL IL C + GK+ VLP +V P+EV+ Y DA++KH
Sbjct: 350 ILSKNYAESEWCLKELSKILECKDQEGKNQLVLPSSTEVTPTEVRHVINRYADAMAKHEK 171
Query: 307 N 307
N
Sbjct: 170 N 168
Score = 92.4 bits (228), Expect = 8e-20
Identities = 57/146 (39%), Positives = 84/146 (57%), Gaps = 2/146 (1%)
Frame = -3
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIV 69
+ +F+SFRG DTR +FT L+ L R+ TF+DD +L+ G+ I L+ IE S+ I+
Sbjct: 527 YQIFISFRGMDTRDDFTKPLYQGLCREGFVTFKDDESLEGGSPIEK-LLDDIEESRFAII 351
Query: 70 VFSKNYASSTWCLRELAYILNCSVLYGKR--VLPVFYDVDLSEVRKQSGGYGESFNYHGK 127
+ SKNYA S WCL+EL+ IL C GK VLP +V +EVR Y ++ H K
Sbjct: 350 ILSKNYAESEWCLKELSKILECKDQEGKNQLVLPSSTEVTPTEVRHVINRYADAMAKHEK 171
Query: 128 RFQDHSNMVQRRRETLQLVGNISGWD 153
D S +Q ++ L V ++G++
Sbjct: 170 NGID-SKTIQAWKKALFDVCTLTGFE 96
>TC19146 similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-containing protein
TSDC, partial (12%)
Length = 701
Score = 93.2 bits (230), Expect = 5e-20
Identities = 50/141 (35%), Positives = 79/141 (55%), Gaps = 1/141 (0%)
Frame = +1
Query: 29 LFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIVVFSKNYASSTWCLRELAYI 88
L+ AL+R+ TF DD L GN IA L+ A E S++ I+VFS+NY S+WCL EL I
Sbjct: 1 LYHALRREGFKTFMDDEGLVGGNEIAKSLLDASEKSRLSIIVFSENYGYSSWCLDELVKI 180
Query: 89 LNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRFQDHSNMVQRRRETLQLVGN 148
+ C + V P+FY V+ S+V Q Y + H K + S+ V++ R L +
Sbjct: 181 VECMETKKQLVWPIFYKVEKSDVTSQENSYEHAMIGHEKNYGKDSDEVKKWRSALSAMAK 360
Query: 149 ISGWDLRDKPHHAE-LENIIE 168
+ G ++++ + E ++ I+E
Sbjct: 361 LEGHNVKENKYEYEFIKKIVE 423
Score = 77.0 bits (188), Expect = 4e-15
Identities = 41/97 (42%), Positives = 54/97 (55%)
Frame = +1
Query: 211 ALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIVVFSKNYASSTWCLRELEYILHC 270
AL+R G F DD L G IA L A E S++ I+VFS+NY S+WCL EL I+ C
Sbjct: 10 ALRREGFKTFMDDEGLVGGNEIAKSLLDASEKSRLSIIVFSENYGYSSWCLDELVKIVEC 189
Query: 271 SKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKHGLN 307
+ + V P+FY V+ S+V Q Y A+ H N
Sbjct: 190 METKKQLVWPIFYKVEKSDVTSQENSYEHAMIGHEKN 300
>BP065014
Length = 498
Score = 80.9 bits (198), Expect = 3e-16
Identities = 44/111 (39%), Positives = 64/111 (57%)
Frame = -2
Query: 41 FRDDTNLQKGNSIASDLIQAIEGSQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVL 100
F+DD +L+ G I +L+ AIE S+V IVV S+N+A S WCL+EL IL+C VL
Sbjct: 497 FKDDLSLEDGAPI-EELLGAIEESRVAIVVLSENFADSKWCLKELVKILDCRKKKKPLVL 321
Query: 101 PVFYDVDLSEVRKQSGGYGESFNYHGKRFQDHSNMVQRRRETLQLVGNISG 151
P+FY V+ + +R G YGE+ H K+ S V+ + L + + G
Sbjct: 320 PIFYKVEPTNIRHLKGRYGEAMAEHEKKLGKDSQTVREWTQALSDISYLKG 168
Score = 80.5 bits (197), Expect = 3e-16
Identities = 41/85 (48%), Positives = 56/85 (65%)
Frame = -2
Query: 220 FRDDTKLKKGEFIAPGLFRAIEASQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVL 279
F+DD L+ G I L AIE S+V IVV S+N+A S WCL+EL IL C KK VL
Sbjct: 497 FKDDLSLEDGAPIEE-LLGAIEESRVAIVVLSENFADSKWCLKELVKILDCRKKKKPLVL 321
Query: 280 PVFYDVDPSEVQKQSGGYGDALSKH 304
P+FY V+P+ ++ G YG+A+++H
Sbjct: 320 PIFYKVEPTNIRHLKGRYGEAMAEH 246
>AI967395
Length = 205
Score = 56.6 bits (135), Expect = 5e-09
Identities = 27/61 (44%), Positives = 41/61 (66%)
Frame = +3
Query: 61 IEGSQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGE 120
I+ S++ IVVFS++YA S CL EL I++ ++ + V PV+Y V+ E+R Q G YGE
Sbjct: 3 IQESRLIIVVFSEHYAVSASCLNELVNIVDVMIMNNQLVWPVYYGVEAPEIRHQRGRYGE 182
Query: 121 S 121
+
Sbjct: 183 A 185
Score = 53.9 bits (128), Expect = 3e-08
Identities = 27/64 (42%), Positives = 42/64 (65%)
Frame = +3
Query: 240 IEASQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGD 299
I+ S++ IVVFS++YA S CL EL I+ + + V PV+Y V+ E++ Q G YG+
Sbjct: 3 IQESRLIIVVFSEHYAVSASCLNELVNIVDVMIMNNQLVWPVYYGVEAPEIRHQRGRYGE 182
Query: 300 ALSK 303
A++K
Sbjct: 183 AMAK 194
>BP065724
Length = 495
Score = 53.5 bits (127), Expect = 4e-08
Identities = 23/50 (46%), Positives = 33/50 (66%)
Frame = -1
Query: 254 YASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSK 303
YA S+WCL E+ IL C +K+ + V P+FY V+PS+V+ Q Y A+ K
Sbjct: 495 YADSSWCLEEVVKILECKQKNDQLVWPIFYKVEPSDVRHQRNSYEKAMIK 346
Score = 52.0 bits (123), Expect = 1e-07
Identities = 26/82 (31%), Positives = 43/82 (51%)
Frame = -1
Query: 75 YASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRFQDHSN 134
YA S+WCL E+ IL C + V P+FY V+ S+VR Q Y ++ R+ S
Sbjct: 495 YADSSWCLEEVVKILECKQKNDQLVWPIFYKVEPSDVRHQRNSYEKAMIKQETRYGKDSE 316
Query: 135 MVQRRRETLQLVGNISGWDLRD 156
V++ R +L V ++ + ++
Sbjct: 315 KVKKWRSSLSQVCDLKAFHYKE 250
>TC9932 weakly similar to UP|Q93YA7 (Q93YA7) Resistance gene-like, partial
(5%)
Length = 689
Score = 53.5 bits (127), Expect = 4e-08
Identities = 24/52 (46%), Positives = 32/52 (61%)
Frame = -2
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAI 240
YDVF++FRG DTR +F H AL G+N F DD L++G + P L + I
Sbjct: 202 YDVFINFRGKDTRHSFVSHLYTALSNAGVNVFLDDENLERGLELEPELQKLI 47
Score = 47.0 bits (110), Expect = 4e-06
Identities = 21/52 (40%), Positives = 32/52 (61%)
Frame = -2
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAI 61
+ VF++FRG DTR +F HL+ AL + F DD NL++G + +L + I
Sbjct: 202 YDVFINFRGKDTRHSFVSHLYTALSNAGVNVFLDDENLERGLELEPELQKLI 47
>AV408271
Length = 409
Score = 49.7 bits (117), Expect = 6e-07
Identities = 26/47 (55%), Positives = 31/47 (65%)
Frame = +1
Query: 186 EINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFI 232
E YDVFVSFRG D R F H A +++ INAF DD KLK+G+ I
Sbjct: 271 EEKYDVFVSFRGEDIRDGFLSHLADAFRQKKINAFMDD-KLKRGQEI 408
Score = 42.4 bits (98), Expect = 1e-04
Identities = 22/50 (44%), Positives = 30/50 (60%)
Frame = +1
Query: 4 PRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSI 53
P ++ + VFVSFRG D R F HL A ++K+I F DD L++G I
Sbjct: 262 PPQEEKYDVFVSFRGEDIRDGFLSHLADAFRQKKINAFMDD-KLKRGQEI 408
>AV406565
Length = 207
Score = 42.4 bits (98), Expect = 1e-04
Identities = 23/52 (44%), Positives = 36/52 (69%)
Frame = +2
Query: 2 TSPRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSI 53
++P++K+D VF+SFRG DTR+ T HL L R ++ T+ D +LQ+G+ I
Sbjct: 59 SAPQQKHD--VFLSFRGEDTRYTCTGHLQATLTRLQVKTY-IDYDLQRGDEI 205
Score = 36.6 bits (83), Expect = 0.006
Identities = 19/44 (43%), Positives = 27/44 (61%)
Frame = +2
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFI 232
+DVF+SFRG DTR+ T H A L R + + D L++G+ I
Sbjct: 77 HDVFLSFRGEDTRYTCTGHLQATLTRLQVKTY-IDYDLQRGDEI 205
>TC15876 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-containing
protein TSDC, partial (13%)
Length = 558
Score = 37.4 bits (85), Expect = 0.003
Identities = 16/37 (43%), Positives = 24/37 (64%)
Frame = +1
Query: 267 ILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSK 303
IL C +K+ + V P+FY V+PS+V+ Q Y A+ K
Sbjct: 16 ILECXQKNDQLVWPIFYKVEPSDVRHQRNSYEKAMIK 126
Score = 34.3 bits (77), Expect = 0.027
Identities = 19/69 (27%), Positives = 34/69 (48%)
Frame = +1
Query: 88 ILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRFQDHSNMVQRRRETLQLVG 147
IL C + V P+FY V+ S+VR Q Y ++ R+ S V++ R +L V
Sbjct: 16 ILECXQKNDQLVWPIFYKVEPSDVRHQRNSYEKAMIKPETRYGKDSEKVKKWRSSLSQVC 195
Query: 148 NISGWDLRD 156
++ + ++
Sbjct: 196 DLKAFHYKE 222
>TC20188 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-containing
protein TSDC, partial (11%)
Length = 494
Score = 36.6 bits (83), Expect = 0.006
Identities = 19/71 (26%), Positives = 35/71 (48%), Gaps = 1/71 (1%)
Frame = +1
Query: 99 VLPVFYDVDLSEVRKQSGGYGESFNYHGKRFQDHSNMVQRRRETLQLVGNISGWDLRDKP 158
V P+FY++ S+V Q+ YG + H F S VQ+ + L + + G + +
Sbjct: 10 VWPIFYEISXSDVSNQTKSYGHAMLAHENSFGKDSEKVQKWKSALSEMAFLQGEHITENE 189
Query: 159 HHAEL-ENIIE 168
+ +L + I+E
Sbjct: 190 YEYKLIQKIVE 222
Score = 29.6 bits (65), Expect = 0.68
Identities = 11/27 (40%), Positives = 17/27 (62%)
Frame = +1
Query: 278 VLPVFYDVDPSEVQKQSGGYGDALSKH 304
V P+FY++ S+V Q+ YG A+ H
Sbjct: 10 VWPIFYEISXSDVSNQTKSYGHAMLAH 90
>TC12592
Length = 534
Score = 35.4 bits (80), Expect = 0.012
Identities = 16/53 (30%), Positives = 29/53 (54%)
Frame = +3
Query: 99 VLPVFYDVDLSEVRKQSGGYGESFNYHGKRFQDHSNMVQRRRETLQLVGNISG 151
VLPVFY V+ +++R G +GE+ +F SN + ++ L+ + + G
Sbjct: 9 VLPVFYGVEPTDIRNLKGKFGEAMAEIENKFGKDSNTIHEWKQALRDISYLKG 167
Score = 32.3 bits (72), Expect = 0.10
Identities = 11/26 (42%), Positives = 21/26 (80%)
Frame = +3
Query: 278 VLPVFYDVDPSEVQKQSGGYGDALSK 303
VLPVFY V+P++++ G +G+A+++
Sbjct: 9 VLPVFYGVEPTDIRNLKGKFGEAMAE 86
>TC12663 weakly similar to UP|Q84ZV8 (Q84ZV8) R 3 protein, partial (4%)
Length = 494
Score = 34.3 bits (77), Expect = 0.027
Identities = 17/38 (44%), Positives = 24/38 (62%)
Frame = +3
Query: 12 VFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQK 49
VF++FRG+DT F L +L K I+TF +D LQ+
Sbjct: 33 VFLNFRGSDTCHGFISFLCKSLLDKGIYTFINDEELQR 146
>AV777598
Length = 371
Score = 31.2 bits (69), Expect = 0.23
Identities = 13/17 (76%), Positives = 15/17 (87%)
Frame = +3
Query: 189 YDVFVSFRGPDTRFNFT 205
YDVF+SF G DTRF+FT
Sbjct: 309 YDVFLSFCGVDTRFSFT 359
Score = 30.0 bits (66), Expect = 0.52
Identities = 12/21 (57%), Positives = 17/21 (80%)
Frame = +3
Query: 10 HTVFVSFRGTDTRFNFTDHLF 30
+ VF+SF G DTRF+FT +L+
Sbjct: 309 YDVFLSFCGVDTRFSFTGNLY 371
>BP068235
Length = 511
Score = 30.8 bits (68), Expect = 0.30
Identities = 14/42 (33%), Positives = 22/42 (52%)
Frame = -1
Query: 261 LRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALS 302
L L L C K + + P Y V+PS+++ + YG AL+
Sbjct: 475 LMNLSRFLECKKMKNQLLWPCPYKVEPSDIRYRPSSYGTALA 350
>BP058043
Length = 375
Score = 28.5 bits (62), Expect = 1.5
Identities = 13/29 (44%), Positives = 17/29 (57%)
Frame = -3
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGI 217
YDVF+S RG R F + F L ++GI
Sbjct: 154 YDVFISLRGEVLRDTFIECFLKELMQKGI 68
>TC15261 weakly similar to UP|Q9ZQ55 (Q9ZQ55) At2g22570 protein, partial
(75%)
Length = 1031
Score = 28.1 bits (61), Expect = 2.0
Identities = 19/76 (25%), Positives = 31/76 (40%), Gaps = 2/76 (2%)
Frame = -1
Query: 178 PSRTKDLVEINYDVFVSFRGPDTRFNFTDHF--CAALQRRGINAFRDDTKLKKGEFIAPG 235
PS + + + F + R T F +D C GI+ + K K G+F +
Sbjct: 419 PSMHSFRLSVAFGSFSNHRSSGTNFPSSDPIRQCEG*GGSGISG*CESRKAKIGQFFSAN 240
Query: 236 LFRAIEASQVYIVVFS 251
F + S +V+FS
Sbjct: 239 FFERLTDSSTILVIFS 192
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.139 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,349,568
Number of Sequences: 28460
Number of extensions: 70880
Number of successful extensions: 403
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 359
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 398
length of query: 309
length of database: 4,897,600
effective HSP length: 90
effective length of query: 219
effective length of database: 2,336,200
effective search space: 511627800
effective search space used: 511627800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146863.4


