

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.24 - phase: 0
(580 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
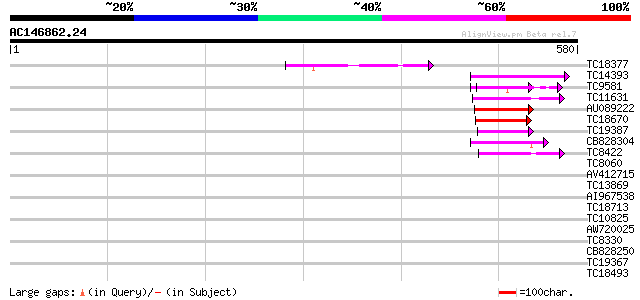
Sequences producing significant alignments: (bits) Value
TC18377 similar to UP|BAC97801 (BAC97801) RelA-SpoT like protein... 93 1e-19
TC14393 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, p... 52 3e-07
TC9581 UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, complete 52 3e-07
TC11631 weakly similar to UP|TCH2_ARATH (P25070) Calmodulin-rela... 50 8e-07
AU089222 49 2e-06
TC18670 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, p... 49 2e-06
TC19387 weakly similar to UP|Q9FYK2 (Q9FYK2) F21J9.28, partial (... 47 7e-06
CB828304 42 4e-04
TC8422 UP|AAQ63461 (AAQ63461) Calmodulin 4 (Fragment), complete 41 5e-04
TC8060 UP|O49184 (O49184) Calmodulin (ESTS AU081349) (Auxin-regu... 40 0.001
AV412715 36 0.020
TC13869 weakly similar to UP|Q9FR00 (Q9FR00) Avr9/Cf-9 rapidly e... 35 0.026
AI967538 35 0.044
TC18713 similar to GB|AAL90989.1|19699246|AY090328 AT3g03000/F13... 35 0.044
TC10825 similar to GB|AAP68339.1|31711966|BT008900 At1g74740 {Ar... 34 0.057
AW720025 34 0.057
TC8330 weakly similar to GB|AAM61257.1|21536925|AY084696 calmodu... 33 0.097
CB828250 33 0.17
TC19367 homologue to UP|Q8QFN0 (Q8QFN0) Guanylate cyclase activa... 32 0.28
TC18493 similar to UP|Q948T9 (Q948T9) Respiratory burst oxidase ... 29 1.8
>TC18377 similar to UP|BAC97801 (BAC97801) RelA-SpoT like protein PsRSH1,
partial (20%)
Length = 449
Score = 92.8 bits (229), Expect = 1e-19
Identities = 57/154 (37%), Positives = 84/154 (54%), Gaps = 3/154 (1%)
Frame = +1
Query: 283 LIDVYKDELLESLKSDPILAELVDDIS---VKGRYKSRYSTMKKLLKDGRRPEDVNDVLG 339
L+D Y D ++ S A + IS + GR+KS YS K+LK +D++D+ G
Sbjct: 19 LVDSYDDVMIASAIERLEQALKDEGISYHVISGRHKSLYSVYCKMLKKKLSIDDIHDIYG 198
Query: 340 LRVVLNPKSRENALEAGERACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDV 399
LR++++ E CY+A ++ +W EIP + KDYIS PK NGY+SLH V
Sbjct: 199 LRLIVDK----------EEDCYKALTVVHKLWYEIPGKLKDYISCPKFNGYQSLHTVV-- 342
Query: 400 SEIGRTRPLMEIQIRTTEMDRLAVGGMASHSLYK 433
+G + +E+QIRT +M A G A+H YK
Sbjct: 343 --MGEGKVPLEVQIRTKDMHLQAEFGFAAHWRYK 438
>TC14393 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, partial
(85%)
Length = 786
Score = 52.0 bits (123), Expect = 3e-07
Identities = 31/103 (30%), Positives = 52/103 (50%), Gaps = 2/103 (1%)
Frame = +3
Query: 472 EFDQRDRVFRLLDKNGDGKISIEELTEVIEELGA--PGEDAHDMMLLLDSNSDGSLSSDE 529
+ ++ RVF D NGDGKIS+ EL V+ LG+ P E+ +M+ LD + DG ++ E
Sbjct: 141 DMEELKRVFTRFDTNGDGKISVNELDNVLRSLGSGVPPEELKRVMVDLDGDHDGFINLSE 320
Query: 530 FQMFQKQVEMVRNLEDRDDEYKKILDEKLHMADDSGLIQVYNK 572
F F + + D ++ +K + + L +V N+
Sbjct: 321 FAAFCRSDTADGGASELHDAFELYDQDKNGLISAAELCKVLNR 449
Score = 39.3 bits (90), Expect = 0.002
Identities = 20/54 (37%), Positives = 34/54 (62%), Gaps = 2/54 (3%)
Frame = +3
Query: 480 FRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEFQ 531
F L D++ +G IS EL +V+ LG E+ H M+ +DS+ DG+++ +EF+
Sbjct: 381 FELYDQDKNGLISAAELCKVLNRLGMNCSEEECHKMIKSVDSDGDGNVNFEEFK 542
>TC9581 UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, complete
Length = 1049
Score = 51.6 bits (122), Expect = 3e-07
Identities = 29/90 (32%), Positives = 53/90 (58%), Gaps = 2/90 (2%)
Frame = +3
Query: 478 RVFRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEFQMFQK 535
RVF++ D+NGDG+I+ +EL + +E LG P ++ M+ +D N DG + DEF +
Sbjct: 357 RVFQMFDRNGDGRITKKELNDSLENLGIFIPDKELTQMIERIDVNGDGCVDIDEFGELYQ 536
Query: 536 QVEMVRNLEDRDDEYKKILDEKLHMADDSG 565
+ +++RD+E + + E ++ D +G
Sbjct: 537 SI-----MDERDEE--EDMREAFNVFDQNG 605
Score = 43.5 bits (101), Expect = 9e-05
Identities = 25/68 (36%), Positives = 37/68 (53%), Gaps = 4/68 (5%)
Frame = +3
Query: 472 EFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPG----EDAHDMMLLLDSNSDGSLSS 527
E D R+ F + D+NGDG I++EEL V+ LG ED M++ +D + DG +
Sbjct: 561 EEDMRE-AFNVFDQNGDGFITVEELRTVLASLGIKQGRTVEDCKKMIMKVDVDGDGMVDY 737
Query: 528 DEFQMFQK 535
EF+ K
Sbjct: 738 KEFKQMMK 761
>TC11631 weakly similar to UP|TCH2_ARATH (P25070) Calmodulin-related protein
2, touch-induced, partial (68%)
Length = 530
Score = 50.4 bits (119), Expect = 8e-07
Identities = 36/98 (36%), Positives = 51/98 (51%), Gaps = 4/98 (4%)
Frame = +3
Query: 474 DQRDRVFRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMMLLLDSNSDGSLSSDEFQ 531
D+ ++F DKNGDGKIS EL E++ LG E+ MM +D N DG + EF
Sbjct: 96 DEVRKIFNKFDKNGDGKISCTELKEMLCALGTKTSSEEVKRMMAEIDQNGDGYIDLKEFA 275
Query: 532 MFQKQVEMVRNLEDR-DDEYKKILDE-KLHMADDSGLI 567
F +L D +D+ K++ D L+ D +GLI
Sbjct: 276 EF--------HLHDAGEDDSKELRDACDLYDLDKNGLI 365
Score = 31.2 bits (69), Expect = 0.48
Identities = 30/121 (24%), Positives = 56/121 (45%), Gaps = 20/121 (16%)
Frame = +3
Query: 454 ELAALRLKDFPSANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGED---- 509
+++ LK+ A ++ R+ +D+NGDG I ++E E L GED
Sbjct: 144 KISCTELKEMLCALGTKTSSEEVKRMMAEIDQNGDGYIDLKEFAEF--HLHDAGEDDSKE 317
Query: 510 AHDMMLLLDSNSDGSLSSDEFQMFQKQV----------EMVRNLE-DRD-----DEYKKI 553
D L D + +G +S++E K++ +M+ N++ D D +E+KK+
Sbjct: 318 LRDACDLYDLDKNGLISANELHSVLKKLGEKCSLGDCRKMISNVDADGDGNVNFEEFKKM 497
Query: 554 L 554
+
Sbjct: 498 M 500
>AU089222
Length = 345
Score = 49.3 bits (116), Expect = 2e-06
Identities = 25/61 (40%), Positives = 38/61 (61%), Gaps = 1/61 (1%)
Frame = +3
Query: 476 RDRVFRLLDKNGDGKISIEELTEVIEELGA-PGEDAHDMMLLLDSNSDGSLSSDEFQMFQ 534
R+R+F+ D NGDG+IS EL E ++ LG+ E+ MM +D + DG +S +EF F
Sbjct: 78 RERIFKRFDANGDGQISSSELGEALKTLGSVTAEEVQRMMDEIDIDGDGFISYEEFTSFA 257
Query: 535 K 535
+
Sbjct: 258 R 260
>TC18670 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, partial
(41%)
Length = 329
Score = 49.3 bits (116), Expect = 2e-06
Identities = 26/59 (44%), Positives = 36/59 (60%), Gaps = 2/59 (3%)
Frame = +3
Query: 477 DRVFRLLDKNGDGKISIEELTEVIEELGA--PGEDAHDMMLLLDSNSDGSLSSDEFQMF 533
+RVF D N DGKIS++EL V+ LG+ P E+ +M LD++ DG +S EF F
Sbjct: 120 ERVFNRFDANSDGKISVDELDSVLRTLGSGVPPEELQQVMEDLDTDRDGFISLPEFAAF 296
>TC19387 weakly similar to UP|Q9FYK2 (Q9FYK2) F21J9.28, partial (45%)
Length = 631
Score = 47.4 bits (111), Expect = 7e-06
Identities = 24/59 (40%), Positives = 34/59 (56%), Gaps = 2/59 (3%)
Frame = +1
Query: 479 VFRLLDKNGDGKISIEELTEVIEELGA--PGEDAHDMMLLLDSNSDGSLSSDEFQMFQK 535
+F+ D+NGDGKIS EL +++ LG+ E+ MM LD N DG + EF F +
Sbjct: 58 IFKKFDRNGDGKISCSELKDLMAALGSKTTAEEVRRMMAELDRNGDGHIDLKEFGEFHR 234
Score = 31.2 bits (69), Expect = 0.48
Identities = 20/85 (23%), Positives = 40/85 (46%), Gaps = 1/85 (1%)
Frame = +1
Query: 454 ELAALRLKDFPSANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIE-ELGAPGEDAHD 512
+++ LKD +A ++ R+ LD+NGDG I ++E E G ++ +
Sbjct: 91 KISCSELKDLMAALGSKTTAEEVRRMMAELDRNGDGHIDLKEFGEFHRGGGGGNSKEIRE 270
Query: 513 MMLLLDSNSDGSLSSDEFQMFQKQV 537
+ D + +G +S+ E K++
Sbjct: 271 AFEMYDLDKNGLISAKELHSVMKKL 345
>CB828304
Length = 489
Score = 41.6 bits (96), Expect = 4e-04
Identities = 24/87 (27%), Positives = 43/87 (48%), Gaps = 7/87 (8%)
Frame = +1
Query: 472 EFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAP--GEDAHDMMLLLDSNSDGSLSSDE 529
+ +Q +F D + DG +++ EL ++ LG G+ HD++ +DSN +GS+ DE
Sbjct: 109 QLNQLREIFARFDMDSDGSLTMLELAALLRSLGLKPSGDQLHDLLSNMDSNGNGSVEFDE 288
Query: 530 FQM-----FQKQVEMVRNLEDRDDEYK 551
+ E++ N E D +K
Sbjct: 289 LVRTILPDLKNNAEVLLNQEQLLDVFK 369
>TC8422 UP|AAQ63461 (AAQ63461) Calmodulin 4 (Fragment), complete
Length = 832
Score = 41.2 bits (95), Expect = 5e-04
Identities = 28/90 (31%), Positives = 46/90 (51%), Gaps = 2/90 (2%)
Frame = +1
Query: 480 FRLLDKNGDGKISIEELTEVIEELGAPGEDA--HDMMLLLDSNSDGSLSSDEFQMFQKQV 537
F L DK+GDG I+ +EL V+ LG +A DM+ +D++ +G++ EF
Sbjct: 184 FSLFDKDGDGSITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNL---- 351
Query: 538 EMVRNLEDRDDEYKKILDEKLHMADDSGLI 567
M R ++D D E + ++ D +G I
Sbjct: 352 -MARKMKDTDSEEELKEAFRVFDKDQNGFI 438
>TC8060 UP|O49184 (O49184) Calmodulin (ESTS AU081349) (Auxin-regulated
calmodulin) (Calmodulin TACAM1-1) (Calmodulin TACAM1-2)
(Calmodulin TACAM1-3) (Calmodulin TACAM3-1) (Calmodulin
TACAM3-2) (Calmodulin TACAM3-3) (Calmodulin TACAM4-1)
(CaM protein), complete
Length = 615
Score = 40.0 bits (92), Expect = 0.001
Identities = 28/90 (31%), Positives = 46/90 (51%), Gaps = 2/90 (2%)
Frame = +3
Query: 480 FRLLDKNGDGKISIEELTEVIEELGAPGEDA--HDMMLLLDSNSDGSLSSDEFQMFQKQV 537
F L DK+GDG I+ +EL V+ LG +A DM+ +D++ +G++ EF
Sbjct: 117 FSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNL---- 284
Query: 538 EMVRNLEDRDDEYKKILDEKLHMADDSGLI 567
M R ++D D E + ++ D +G I
Sbjct: 285 -MARKMKDTDSEEELKEAFRVFDKDQNGFI 371
>AV412715
Length = 300
Score = 35.8 bits (81), Expect = 0.020
Identities = 19/47 (40%), Positives = 30/47 (63%), Gaps = 1/47 (2%)
Frame = +1
Query: 460 LKDFPSANHKGIEFD-QRDRVFRLLDKNGDGKISIEELTEVIEELGA 505
+ DF ++ +E Q +VF+ +D NGDGKIS EL+E++ LG+
Sbjct: 157 ISDFSVSSFLHMEVSSQFQQVFKFIDSNGDGKISATELSELLLCLGS 297
>TC13869 weakly similar to UP|Q9FR00 (Q9FR00) Avr9/Cf-9 rapidly elicited
protein 31, partial (45%)
Length = 547
Score = 35.4 bits (80), Expect = 0.026
Identities = 31/116 (26%), Positives = 54/116 (45%), Gaps = 3/116 (2%)
Frame = +3
Query: 425 GMASHSLYKAGLTNPEEAKRLKTIMLAAAELAALRLKDFPSANHKGIEFDQRDRVFRLLD 484
GM + LT+P + RL++ L +LRL+ R+F + D
Sbjct: 198 GMVHMGQEEITLTSPSRSFRLRS-----PSLNSLRLR----------------RIFDMFD 314
Query: 485 KNGDGKISIEELTEVIEELGAPGEDAH-DMML--LLDSNSDGSLSSDEFQMFQKQV 537
KNGD I++EE+++ + LG E A D M+ + ++G L+ D+F + +
Sbjct: 315 KNGDCMITVEEISQALNLLGLEAEVAEIDSMIRSYIRPGNEG-LTYDDFMALHESI 479
>AI967538
Length = 432
Score = 34.7 bits (78), Expect = 0.044
Identities = 18/41 (43%), Positives = 24/41 (57%), Gaps = 2/41 (4%)
Frame = +3
Query: 458 LRLKDFPSANHKG--IEFDQRDRVFRLLDKNGDGKISIEEL 496
+ ++F A KG I + FR+ DKNGDGKIS EE+
Sbjct: 306 INFEEFMEAQKKGGGIRTMEIQSAFRMFDKNGDGKISAEEV 428
>TC18713 similar to GB|AAL90989.1|19699246|AY090328 AT3g03000/F13E7_5
{Arabidopsis thaliana;}, partial (88%)
Length = 868
Score = 34.7 bits (78), Expect = 0.044
Identities = 22/81 (27%), Positives = 35/81 (43%), Gaps = 14/81 (17%)
Frame = +1
Query: 464 PSANHKGI------------EFDQRDRVFRLLDKNGDGKISIEELTEVIEELG--APGED 509
P NHK I + + +FR D+N DG ++ EL+ ++ LG E
Sbjct: 145 PKKNHKEIMSKKQAVNLDEEQIAELREIFRSFDRNNDGSLTQLELSSLLRSLGLKPSAEQ 324
Query: 510 AHDMMLLLDSNSDGSLSSDEF 530
+ D+NS+G + EF
Sbjct: 325 VEGFIQKADTNSNGLIEFSEF 387
>TC10825 similar to GB|AAP68339.1|31711966|BT008900 At1g74740 {Arabidopsis
thaliana;}, partial (32%)
Length = 1091
Score = 34.3 bits (77), Expect = 0.057
Identities = 26/92 (28%), Positives = 44/92 (47%), Gaps = 2/92 (2%)
Frame = +1
Query: 479 VFRLLDKNGDGKISIEELTEVIEELGAPGEDAHDMMLL--LDSNSDGSLSSDEFQMFQKQ 536
+F L+D + DG++S EEL + ++G+ D ML+ D + +G L EF
Sbjct: 28 MFTLMDTDKDGRVSYEELKAGLRKVGSQLADQEIKMLMEVADVDGNGVLDYGEFVAVTIH 207
Query: 537 VEMVRNLEDRDDEYKKILDEKLHMADDSGLIQ 568
++ + N D+ + K K D SG I+
Sbjct: 208 LQKMEN----DEHFHKAF--KFFDKDGSGYIE 285
>AW720025
Length = 563
Score = 34.3 bits (77), Expect = 0.057
Identities = 27/81 (33%), Positives = 40/81 (49%), Gaps = 3/81 (3%)
Frame = +3
Query: 480 FRLLDKNGDGKISIEELTEVIEELGAPG---EDAHDMMLLLDSNSDGSLSSDEFQMFQKQ 536
F+LL G IS E L LG G EDA +M+ D + DGSL+ EF + +
Sbjct: 210 FKLLSDPETGLISSESLMRNSALLGMDGMTKEDAEEMVKQGDLDGDGSLNETEFCILMVR 389
Query: 537 VEMVRNLEDRDDEYKKILDEK 557
+ +ED + +K L+E+
Sbjct: 390 LS-PGMMEDAESWLEKALEEE 449
>TC8330 weakly similar to GB|AAM61257.1|21536925|AY084696 calmodulin-like
protein {Arabidopsis thaliana;} , partial (72%)
Length = 1174
Score = 33.5 bits (75), Expect = 0.097
Identities = 18/50 (36%), Positives = 29/50 (58%), Gaps = 3/50 (6%)
Frame = +3
Query: 480 FRLLDKNGDGKISIEELTEVIEELGA---PGEDAHDMMLLLDSNSDGSLS 526
FRL+D++ DG +S +L V+ LGA E+ M+ +D + GS+S
Sbjct: 306 FRLIDRDNDGVVSRADLVAVLTRLGALQLSDEEVEVMLREVDGDGRGSIS 455
Score = 31.2 bits (69), Expect = 0.48
Identities = 20/65 (30%), Positives = 33/65 (50%), Gaps = 3/65 (4%)
Frame = +3
Query: 480 FRLLDKNGDGKISIEELTEVIEELG---APGEDAHDMMLLLDSNSDGSLSSDEFQMFQKQ 536
F + D + DG+IS EEL V +G ++ M+ +D N DG + F+ F +
Sbjct: 519 FEVFDTDRDGRISAEELLRVFTAIGDERCTLDECRRMIEGVDRNGDGFVC---FEDFSRM 689
Query: 537 VEMVR 541
+E+ R
Sbjct: 690 MELQR 704
>CB828250
Length = 485
Score = 32.7 bits (73), Expect = 0.17
Identities = 17/50 (34%), Positives = 29/50 (58%), Gaps = 2/50 (4%)
Frame = +1
Query: 467 NHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELG--APGEDAHDMM 514
N+K + + VF+++DK+GDGK+S +L + G A ED + M+
Sbjct: 313 NNKPLGCGAMEEVFKVMDKDGDGKLSHHDLKSYMASAGFSATDEDINAMI 462
>TC19367 homologue to UP|Q8QFN0 (Q8QFN0) Guanylate cyclase activator protein
2 (Guanylate cyclase activating protein 2), partial (7%)
Length = 526
Score = 32.0 bits (71), Expect = 0.28
Identities = 20/68 (29%), Positives = 38/68 (55%), Gaps = 10/68 (14%)
Frame = +1
Query: 480 FRLLDKNGDGKISIEELT---EVIEELGA---PGEDAHDMMLLL----DSNSDGSLSSDE 529
F ++D+NGDG++S +E+ + LG+ P ++ DM+ L+ D + +GSL E
Sbjct: 106 FAMVDENGDGELSRDEVRGGFGLFMPLGSESQPKQEVDDMLDLIFKRFDEDQNGSLDLKE 285
Query: 530 FQMFQKQV 537
F+ ++
Sbjct: 286 FKSLMTEI 309
>TC18493 similar to UP|Q948T9 (Q948T9) Respiratory burst oxidase homolog,
partial (15%)
Length = 553
Score = 29.3 bits (64), Expect = 1.8
Identities = 14/29 (48%), Positives = 21/29 (72%), Gaps = 1/29 (3%)
Frame = +1
Query: 473 FDQRDRVF-RLLDKNGDGKISIEELTEVI 500
FD R + F L+DKN DG+I+ E++ E+I
Sbjct: 379 FDSRVQTFFDLVDKNADGRINEEQVKEII 465
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.135 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,432,999
Number of Sequences: 28460
Number of extensions: 108043
Number of successful extensions: 714
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 699
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 709
length of query: 580
length of database: 4,897,600
effective HSP length: 95
effective length of query: 485
effective length of database: 2,193,900
effective search space: 1064041500
effective search space used: 1064041500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146862.24


