

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.23 + phase: 0 /pseudo
(320 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
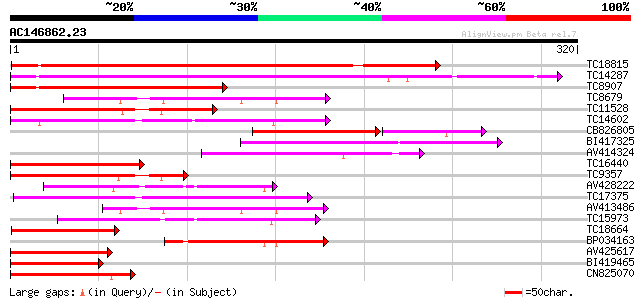
TC18815 similar to UP|Q8W1D5 (Q8W1D5) CBL-interacting protein ki... 229 4e-61
TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein k... 227 2e-60
TC8907 similar to UP|Q8H0X3 (Q8H0X3) Serine/threonine protein ki... 154 1e-38
TC8679 homologue to UP|Q43466 (Q43466) Protein kinase 3 , partia... 121 2e-28
TC11528 homologue to UP|Q39868 (Q39868) Protein kinase , partial... 116 6e-27
TC14602 UP|Q8GS56 (Q8GS56) Phosphoenolpyruvate carboxylase kinas... 116 6e-27
CB826805 113 4e-26
BI417325 108 2e-24
AV414324 102 1e-22
TC16440 similar to UP|Q93Y18 (Q93Y18) SNF1 related protein kinas... 97 3e-21
TC9357 homologue to UP|Q39193 (Q39193) Protein kinase (Protein k... 90 4e-19
AV428222 88 2e-18
TC17375 mitogen-activated kinase kinase kinase alpha [Lotus corn... 81 3e-16
AV413486 80 5e-16
TC15973 homologue to GB|AAL15278.1|16323087|AY057647 AT3g01490/F... 76 7e-15
TC18664 similar to UP|Q944I6 (Q944I6) At2g30360/T9D9.17, partial... 76 9e-15
BP034163 72 1e-13
AV425617 72 1e-13
BI419465 70 5e-13
CN825070 70 6e-13
>TC18815 similar to UP|Q8W1D5 (Q8W1D5) CBL-interacting protein kinase
CIPK25, partial (58%)
Length = 979
Score = 229 bits (585), Expect = 4e-61
Identities = 119/244 (48%), Positives = 159/244 (64%), Gaps = 2/244 (0%)
Frame = +2
Query: 2 VLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAKG 61
++EY+ G ELF K+A KGKL + R+ FQQLI V YCHS+GV HRDLK EN+L+D
Sbjct: 266 IMEYIRGGELFAKVA-KGKLKDDLARRYFQQLISAVDYCHSRGVSHRDLKPENLLLDENE 442
Query: 62 NLKITDFGLSALPQQFRADGLLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYVVL 121
NLK++DFGLS LP+Q R DGLLHT CG+ YVAPE+L +GY+G +D WSCGVILY +L
Sbjct: 443 NLKVSDFGLSGLPEQLRQDGLLHTQCGTPAYVAPEVLRKKGYDGFKTDTWSCGVILYALL 622
Query: 122 TGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDLWF 181
G LPF NL +Y KVL+ +FQ P W S ++ +I +IL +P RIT++ I WF
Sbjct: 623 AGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNRRITISSIMRVSWF 802
Query: 182 KEGYNQAYHEDEEDGVYVDDQAFNIHELSHEEEKSSSQSPIR-INAFQLI-GMSSCLDLS 239
++G++ + + D+ FN S E+ + + + INAF+ I MSS DLS
Sbjct: 803 QKGFSASIPIPDP-----DESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 967
Query: 240 GFFE 243
GFFE
Sbjct: 968 GFFE 979
>TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein kinase-like
protein (CBL-interacting protein kinase 5), partial
(75%)
Length = 1999
Score = 227 bits (578), Expect = 2e-60
Identities = 128/318 (40%), Positives = 190/318 (59%), Gaps = 6/318 (1%)
Frame = +3
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
+V+EYV G ELF K+ +KGKL E + RK FQQLI V +CHS+GV HRDLK EN+L+D
Sbjct: 657 LVMEYVKGGELFTKV-NKGKLNEDDARKYFQQLISAVDFCHSRGVTHRDLKPENLLLDEN 833
Query: 61 GNLKITDFGLSALPQQFRADGLLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYVV 120
+LK++DFGLSALP+Q R DG+L T CG+ YVAPE+L +GY+G+ +D+WSCGVILY +
Sbjct: 834 EDLKVSDFGLSALPEQRRDDGMLVTPCGTPAYVAPEVLKKKGYDGSKADIWSCGVILYAL 1013
Query: 121 LTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDLW 180
L+G+LPF N+ +Y+K K +++ P+W+S A+N+I +L +P+ R ++ EI D W
Sbjct: 1014LSGYLPFQGENVMRIYRKAFKAEYEFPEWISPQAKNLISNLLVADPEKRYSIPEIISDPW 1193
Query: 181 FKEGYNQAYHEDEEDGVYVDDQAFNIHELSHE---EEKSSSQSPIR--INAFQLI-GMSS 234
F+ G+ + + +D E E ++ P R NAF++I +S
Sbjct: 1194FQYGFMRPLAFSINESAVGEDSTI*SFWWGGEXVPELVAAMGKPARPFYNAFEIISSLSH 1373
Query: 235 CLDLSGFFEKEDVSERKIRFTANLSAKVLMEKIEDSVTEMEFRVQKKNGKMKVMQENKDH 294
DL FE S F + SA ++ K+E ++ FRV K MQ K+
Sbjct: 1374GFDLRSLFETRKRSPS--MFISKYSASAVVAKLEGMAKKLNFRVTGKKEFTVRMQGMKEG 1547
Query: 295 KTLGSLSVIIEVMDIQNE 312
+ G L++ +EV ++ E
Sbjct: 1548RK-GKLAMTMEVFEVAPE 1598
>TC8907 similar to UP|Q8H0X3 (Q8H0X3) Serine/threonine protein kinase-like
protein, partial (44%)
Length = 941
Score = 154 bits (390), Expect = 1e-38
Identities = 73/123 (59%), Positives = 96/123 (77%)
Frame = +3
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
MV+EYV G ELF K+ +KGK+ E RK FQQLI V +CHS+GV HRDLK EN+L+D
Sbjct: 567 MVVEYVKGGELFAKL-TKGKMTEVAARKYFQQLISAVDFCHSRGVTHRDLKPENLLLDDN 743
Query: 61 GNLKITDFGLSALPQQFRADGLLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYVV 120
+LK++DFGLS+LP+Q R+DG+L T CG+ YVAPE+L +GY+G+ +D+WSCGVILY +
Sbjct: 744 EDLKVSDFGLSSLPEQRRSDGMLLTPCGTPAYVAPEVLKKKGYDGSKADIWSCGVILYAL 923
Query: 121 LTG 123
L G
Sbjct: 924 LCG 932
>TC8679 homologue to UP|Q43466 (Q43466) Protein kinase 3 , partial (70%)
Length = 972
Score = 121 bits (303), Expect = 2e-28
Identities = 69/163 (42%), Positives = 95/163 (57%), Gaps = 12/163 (7%)
Frame = +2
Query: 31 QQLIDGVSYCHSKGVFHRDLKLENVLVDAKG--NLKITDFGLSALPQQFRADGLLHT--- 85
+QLI GV YCH+ + HRDLKLEN L+D LKI DFG S LLH+
Sbjct: 2 EQLISGVHYCHALQICHRDLKLENTLLDGSPAPRLKICDFGYSK-------SSLLHSRPK 160
Query: 86 -TCGSANYVAPEILANRGYNGASSDVWSCGVILYVVLTGFLPFDD----RNLAVLYQKVL 140
T G+ Y+APE+L+ R Y+G +DVWSCGV LYV+L G PF+D RN Q+++
Sbjct: 161 STVGTPAYIAPEVLSRREYDGKLADVWSCGVTLYVMLVGAYPFEDHDDPRNFRKTIQRIM 340
Query: 141 KGDFQKPKW--LSAGAQNIIKRILDPNPKTRITMAEIKEDLWF 181
++ P + +S ++++ RI NP RI++ EIK WF
Sbjct: 341 AVQYKIPDYVHISQDCRHLLSRIFVANPLRRISLKEIKNHPWF 469
>TC11528 homologue to UP|Q39868 (Q39868) Protein kinase , partial (56%)
Length = 640
Score = 116 bits (290), Expect = 6e-27
Identities = 62/123 (50%), Positives = 79/123 (63%), Gaps = 6/123 (4%)
Frame = +3
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
+VLEY +G ELF++I S G+ E E R FQQLI GVSYCHS + HRDLKLEN L+D
Sbjct: 291 IVLEYASGGELFERICSAGRFSEDEARYFFQQLISGVSYCHSMEICHRDLKLENTLLDGN 470
Query: 61 GN--LKITDFGLSALPQQFRADGLLH----TTCGSANYVAPEILANRGYNGASSDVWSCG 114
+ LKI DFG S +LH +T G+ Y+APE+L+ + Y+G +DVWSCG
Sbjct: 471 PSPRLKICDFGYS-------KSAILHSQPKSTVGTPAYIAPEVLSRKEYDGKVADVWSCG 629
Query: 115 VIL 117
V L
Sbjct: 630 VTL 638
>TC14602 UP|Q8GS56 (Q8GS56) Phosphoenolpyruvate carboxylase kinase, complete
Length = 1256
Score = 116 bits (290), Expect = 6e-27
Identities = 67/187 (35%), Positives = 109/187 (57%), Gaps = 6/187 (3%)
Frame = +2
Query: 1 MVLEYVTGEELFDKI--ASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVD 58
MV+E L D+I A+ +PE E L +QL++ V++CH GV HRD+K +NVL
Sbjct: 296 MVIELCQPLTLLDRIVAANGTSIPEVEAAGLMKQLLEAVAHCHRLGVAHRDVKPDNVLFG 475
Query: 59 AKGNLKITDFGLSALPQQFRADGLLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILY 118
G+LK+ DFG + + F + G+ YVAPE+L R Y G DVWSCGVILY
Sbjct: 476 GGGDLKLADFGSA---EWFGDGRRMSGVVGTPYYVAPEVLMGREY-GEKVDVWSCGVILY 643
Query: 119 VVLTGFLPFDDRNLAVLYQKVLKGDFQKP----KWLSAGAQNIIKRILDPNPKTRITMAE 174
++L+G PF + A +++ V++G+ + P + +S A++++++++ +P RI+ +
Sbjct: 644 IMLSGTPPFYGDSAAEIFEAVIRGNLRFPSRIFRNVSPAAKDLLRKMICRDPSNRISAEQ 823
Query: 175 IKEDLWF 181
WF
Sbjct: 824 ALRHPWF 844
>CB826805
Length = 635
Score = 113 bits (283), Expect = 4e-26
Identities = 53/72 (73%), Positives = 60/72 (82%)
Frame = +2
Query: 138 KVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDLWFKEGYNQAYHEDEEDGV 197
++ KGD Q PKWLSAGAQNIIKRILDPNP+TRITMA IKED WFK+ Y A +DEED V
Sbjct: 2 QIFKGDVQIPKWLSAGAQNIIKRILDPNPETRITMAGIKEDPWFKQDYTPADPDDEEDDV 181
Query: 198 YVDDQAFNIHEL 209
Y+DD AF+IHEL
Sbjct: 182 YIDDDAFSIHEL 217
Score = 72.0 bits (175), Expect = 1e-13
Identities = 46/93 (49%), Positives = 51/93 (54%), Gaps = 34/93 (36%)
Frame = +3
Query: 211 HEEEKSSSQSPIRINAFQLIGMSSCLDLSGFFEKE------------------------- 245
HEEE S SP +INAFQLIGMSS LDLSGFFE+E
Sbjct: 357 HEEEHRSPGSPTQINAFQLIGMSSFLDLSGFFEQEVRMEDN*QDFLEIHHFMRKFHSCSR 536
Query: 246 ---------DVSERKIRFTANLSAKVLMEKIED 269
VSERKIRFT+NLSAK L+E+IED
Sbjct: 537 LMWFVTDLQHVSERKIRFTSNLSAKDLIERIED 635
>BI417325
Length = 449
Score = 108 bits (269), Expect = 2e-24
Identities = 62/149 (41%), Positives = 85/149 (56%), Gaps = 1/149 (0%)
Frame = +2
Query: 131 NLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDLWFKEGYN-QAY 189
NL LY+K+ +F P WLS A+ +I RILDPNP TRIT+AEI ED WFK+ Y +
Sbjct: 5 NLMELYKKISAAEFTCPPWLSFSARKLITRILDPNPMTRITIAEILEDEWFKKDYKPPIF 184
Query: 190 HEDEEDGVYVDDQAFNIHELSHEEEKSSSQSPIRINAFQLIGMSSCLDLSGFFEKEDVSE 249
E+ E + + F E H EK Q P +NAF+LI MS L+L F+ E +
Sbjct: 185 QENGETNLDDVEAVFKDSEEHHVTEKKEEQ-PTAMNAFELISMSKGLNLENLFDVEQGFK 361
Query: 250 RKIRFTANLSAKVLMEKIEDSVTEMEFRV 278
R+ RFT+ A ++ KIE++ + F V
Sbjct: 362 RETRFTSKSPANEIISKIEEAAKPLGFDV 448
>AV414324
Length = 380
Score = 102 bits (253), Expect = 1e-22
Identities = 52/129 (40%), Positives = 75/129 (57%), Gaps = 3/129 (2%)
Frame = +3
Query: 109 DVWSCGVILYVVLTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKT 168
D+WSCGVILYV+L G+LPF D NL +Y+K+ K +F+ P W + + ++ +ILDPNP+T
Sbjct: 3 DIWSCGVILYVLLAGYLPFRDSNLMEMYRKIGKAEFKFPNWFAPDVRRLLSKILDPNPRT 182
Query: 169 RITMAEIKEDLWFKEGYNQ---AYHEDEEDGVYVDDQAFNIHELSHEEEKSSSQSPIRIN 225
RI++A+I E WF++G + D E D F E E S P +N
Sbjct: 183 RISLAKIMESSWFRKGLQKPAITESGDREIAPLDTDGLFGACENGGPVELS---KPCNLN 353
Query: 226 AFQLIGMSS 234
AF +I S+
Sbjct: 354 AFDIISYST 380
>TC16440 similar to UP|Q93Y18 (Q93Y18) SNF1 related protein kinase, partial
(30%)
Length = 549
Score = 97.4 bits (241), Expect = 3e-21
Identities = 42/76 (55%), Positives = 60/76 (78%)
Frame = +1
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
++++Y G ELF KI+ +G+LPE R+ FQQL+ + +CH GV HRDLK +N+L+DA+
Sbjct: 319 LIVDYAGGGELFSKISRRGRLPEPLARRYFQQLVSALCFCHRNGVAHRDLKPQNLLLDAE 498
Query: 61 GNLKITDFGLSALPQQ 76
GNLK++DFGLSALP+Q
Sbjct: 499 GNLKVSDFGLSALPEQ 546
>TC9357 homologue to UP|Q39193 (Q39193) Protein kinase (Protein kinase,
41K) , partial (71%)
Length = 1181
Score = 90.1 bits (222), Expect = 4e-19
Identities = 50/107 (46%), Positives = 66/107 (60%), Gaps = 6/107 (5%)
Frame = +1
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
+V+EY +G ELF++I + G+ E E R FQQLI GVSYCH+ V HRDLKLEN L+D
Sbjct: 574 IVMEYASGGELFERICNAGRFTEDEARFFFQQLISGVSYCHAMQVCHRDLKLENTLLDGS 753
Query: 61 --GNLKITDFGLSALPQQFRADGLLH----TTCGSANYVAPEILANR 101
LKI DFG S +LH +T G+ Y+APE+L+ +
Sbjct: 754 PTPRLKICDFGYS-------KSSVLHSQPKSTVGTPAYIAPEVLSKQ 873
Score = 38.5 bits (88), Expect = 0.002
Identities = 35/128 (27%), Positives = 53/128 (41%), Gaps = 12/128 (9%)
Frame = +3
Query: 77 FRADGLLHTTCG-SANYVAPEILANRGYNGASSDVWSCGVILYVVLTGFLPFDDRN---- 131
F A CG S Y + I+ R SCGV LYV+L G PF+D N
Sbjct: 798 FSASFTTKINCGNSCIYCSRSIIETRVMMARLQMSGSCGVTLYVMLVGAYPFEDPNEPKD 977
Query: 132 LAVLYQKVLKGDFQKPKW--LSAGAQNIIKRILDPNPKTRITMAE-----IKEDLWFKEG 184
Q+VL + P + +S +++I RI +P R + ++++ +G
Sbjct: 978 FRKTIQRVLSVQYSIPDFVQISPECRHLISRIFVFDPAERSPCLKSGT*MVRKNSLLTDG 1157
Query: 185 YNQAYHED 192
YH D
Sbjct: 1158ERYEYH*D 1181
>AV428222
Length = 416
Score = 88.2 bits (217), Expect = 2e-18
Identities = 54/137 (39%), Positives = 80/137 (57%), Gaps = 5/137 (3%)
Frame = +3
Query: 20 KLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLV---DAKGNLKITDFGLSALPQQ 76
+ PE + + + Q+++ V++CH GV HRDLK EN L +A +K+ DFGLS
Sbjct: 3 RYPEDDAKAILLQILNVVAFCHLHGVVHRDLKPENFLFVSKEADSVMKVIDFGLSDF--- 173
Query: 77 FRADGLLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYVVLTGFLPFDDRNLAVLY 136
R D L+ GSA YVAPE+L +R Y+ +D+WS GVI Y++L G PF R + ++
Sbjct: 174 VRPDQRLNDIVGSAYYVAPEVL-HRSYS-VEADLWSIGVISYILLCGSRPFWARTESGIF 347
Query: 137 QKVLKG--DFQKPKWLS 151
+ VL+ +F W S
Sbjct: 348 RSVLRANPNFDDSPWPS 398
>TC17375 mitogen-activated kinase kinase kinase alpha [Lotus corniculatus
var. japonicus]
Length = 1422
Score = 80.9 bits (198), Expect = 3e-16
Identities = 51/171 (29%), Positives = 87/171 (50%), Gaps = 2/171 (1%)
Frame = +3
Query: 3 LEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAKGN 62
LEYV+G + + G E + +Q++ G++Y HS+ HRD+K N+LVD G
Sbjct: 168 LEYVSGGSIHKLLQEYGAFKEPVIQNYTRQIVSGLAYLHSRNTVHRDIKGANILVDPNGE 347
Query: 63 LKITDFGLSALPQQFRADGLLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYVVLT 122
+K+ DFG+S + + + + GS ++APE++ N G D+ S G + + T
Sbjct: 348 IKLADFGMS---KHINSAASMLSFKGSPYWMAPEVVMNTNGYGLPVDISSLGCTILEMAT 518
Query: 123 GFLPFDD-RNLAVLYQKVLKGDFQK-PKWLSAGAQNIIKRILDPNPKTRIT 171
P+ +A +++ D + P+ LS A+N IK+ L +P R T
Sbjct: 519 SKPPWSQFEGVAAIFKIGNSKDMPEIPEHLSDDAKNFIKQCLQRDPLARPT 671
>AV413486
Length = 402
Score = 80.1 bits (196), Expect = 5e-16
Identities = 52/140 (37%), Positives = 77/140 (54%), Gaps = 12/140 (8%)
Frame = +3
Query: 53 ENVLVDAKG--NLKITDFGLSALPQQFRADGLLHT----TCGSANYVAPEILANRGYNGA 106
EN L+D +LKI DFG S +LH+ T G+ Y+APE+L + Y+G
Sbjct: 3 ENTLLDGSPALHLKICDFGYSK-------SSVLHSQPKSTVGTPAYIAPEVLLKQEYDGK 161
Query: 107 SSDVWSCGVILYVVLTGFLPFDD----RNLAVLYQKVLKGDFQKPKW--LSAGAQNIIKR 160
+DVWSCGV LYV+L G PF+D ++ Q+VL + P + +S +++I R
Sbjct: 162 IADVWSCGVTLYVMLVGTYPFEDPDDPKDFRKTIQRVLTVQYSIPDFVQVSPECRHLISR 341
Query: 161 ILDPNPKTRITMAEIKEDLW 180
I +P RIT+ EI ++ W
Sbjct: 342 IFVFDPAERITIPEILKNEW 401
>TC15973 homologue to GB|AAL15278.1|16323087|AY057647 AT3g01490/F4P13_4
{Arabidopsis thaliana;}, partial (49%)
Length = 944
Score = 76.3 bits (186), Expect = 7e-15
Identities = 46/150 (30%), Positives = 79/150 (52%), Gaps = 2/150 (1%)
Frame = +3
Query: 28 KLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAKGNLKITDFGLSALPQQFRADGLLHTTC 87
+L L G+SY HS+ + HRD+K EN+L+D +KI DFG++ + D T
Sbjct: 69 QLALDLARGLSYLHSQKIVHRDVKTENMLLDKTRTVKIADFGVARVEASNPNDMTGET-- 242
Query: 88 GSANYVAPEILANRGYNGASSDVWSCGVILYVVLTGFLPFDDRNLAVLYQKVLKGDFQK- 146
G+ Y+APE+L YN DV+S G+ L+ + +P+ D + + + V++ + +
Sbjct: 243 GTLGYMAPEVLNGNPYN-RKCDVYSFGICLWEIYCCDMPYPDLSFSEITSAVVRQNLRPE 419
Query: 147 -PKWLSAGAQNIIKRILDPNPKTRITMAEI 175
P+ + N++K+ D P R M E+
Sbjct: 420 IPRCCPSSLANVMKKCWDATPDKRPEMDEV 509
>TC18664 similar to UP|Q944I6 (Q944I6) At2g30360/T9D9.17, partial (34%)
Length = 643
Score = 75.9 bits (185), Expect = 9e-15
Identities = 35/61 (57%), Positives = 45/61 (73%)
Frame = +3
Query: 2 VLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAKG 61
V+E+ G ELF +I++KG+ E R+ FQQLI V YCHS+GVFHRDLK EN+L+D K
Sbjct: 321 VMEFAKGGELFARISTKGRFSEDLARRYFQQLISAVGYCHSRGVFHRDLKPENLLLDDKV 500
Query: 62 N 62
N
Sbjct: 501 N 503
>BP034163
Length = 477
Score = 72.0 bits (175), Expect = 1e-13
Identities = 37/97 (38%), Positives = 61/97 (62%), Gaps = 4/97 (4%)
Frame = +1
Query: 88 GSANYVAPEILANRGYNGASSDVWSCGVILYVVLTGFLPFDDRNLAVLYQKVLKG--DFQ 145
GS Y+APE+L R + G D+WS GVILY++L G PF V+ Q +++ DF+
Sbjct: 19 GSPYYMAPEVL--RRHYGPEVDIWSAGVILYILLCGVPPFWAETEQVVAQAIIRSVVDFK 192
Query: 146 KPKW--LSAGAQNIIKRILDPNPKTRITMAEIKEDLW 180
+ W +S A++++K++L+P+PK R+T E+ + W
Sbjct: 193 RDPWPKVSDNAKDLVKKMLNPDPKRRLTAQEVLDHPW 303
>AV425617
Length = 405
Score = 72.0 bits (175), Expect = 1e-13
Identities = 32/58 (55%), Positives = 41/58 (70%)
Frame = +2
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVD 58
+V+EY G ELF++I + G+ E E R FQQLI GV YCH+ + HRDLKLEN L+D
Sbjct: 200 IVMEYAAGGELFERICTAGRFSEDEARYFFQQLISGVCYCHAMQICHRDLKLENTLLD 373
>BI419465
Length = 487
Score = 70.1 bits (170), Expect = 5e-13
Identities = 33/53 (62%), Positives = 39/53 (73%)
Frame = +3
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLE 53
+V+EY +G ELF+KI S G+ E E R FQQLI GVSYCHS + HRDLKLE
Sbjct: 327 IVMEYASGGELFEKIHSAGRFTEDEARFFFQQLISGVSYCHSMQICHRDLKLE 485
>CN825070
Length = 634
Score = 69.7 bits (169), Expect = 6e-13
Identities = 37/74 (50%), Positives = 46/74 (62%), Gaps = 3/74 (4%)
Frame = +3
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVL---V 57
+V+E G ELFD+I KG+ E E KL + +++ V CHS GV HRDLK EN L V
Sbjct: 357 IVMEICEGGELFDRIVQKGQYSEREAAKLIRTIVEVVEACHSLGVMHRDLKPENFLFDTV 536
Query: 58 DAKGNLKITDFGLS 71
+ LK TDFGLS
Sbjct: 537 EEDAKLKTTDFGLS 578
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.135 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,616,899
Number of Sequences: 28460
Number of extensions: 56248
Number of successful extensions: 502
Number of sequences better than 10.0: 174
Number of HSP's better than 10.0 without gapping: 447
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 451
length of query: 320
length of database: 4,897,600
effective HSP length: 90
effective length of query: 230
effective length of database: 2,336,200
effective search space: 537326000
effective search space used: 537326000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146862.23


