

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
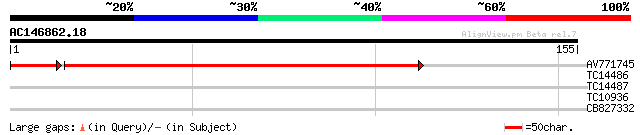
Query= AC146862.18 - phase: 0 /pseudo
(155 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV771745 167 4e-45
TC14486 similar to UP|Q9LID9 (Q9LID9) Gb|AAD15384.1, partial (19%) 27 1.2
TC14487 similar to UP|Q9LID9 (Q9LID9) Gb|AAD15384.1, partial (22%) 27 2.0
TC10936 weakly similar to UP|Q9C887 (Q9C887) Myosin-like protein... 26 3.5
CB827332 25 4.6
>AV771745
Length = 399
Score = 167 bits (422), Expect(2) = 4e-45
Identities = 85/98 (86%), Positives = 90/98 (91%)
Frame = +3
Query: 16 VGDHIYTDVSQSKVHLRWRTALICRELEDEYSALIRCRGDRESLVELINQKEVVGDLFNQ 75
VGDHIYTDVSQSKV+ RWRTA +CRELE+EYSALI R RES VELINQKEV+GDLFNQ
Sbjct: 105 VGDHIYTDVSQSKVNWRWRTAWVCRELEEEYSALIHSRSQRES*VELINQKEVLGDLFNQ 284
Query: 76 LRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVM 113
LRLALQRRSKDRPAQTLAATNM D+DLTESMQKLLIVM
Sbjct: 285 LRLALQRRSKDRPAQTLAATNMKDDDLTESMQKLLIVM 398
Score = 29.3 bits (64), Expect(2) = 4e-45
Identities = 13/14 (92%), Positives = 13/14 (92%)
Frame = +2
Query: 1 MVENSLGIHGDEIL 14
MVENSL IHGDEIL
Sbjct: 59 MVENSLDIHGDEIL 100
>TC14486 similar to UP|Q9LID9 (Q9LID9) Gb|AAD15384.1, partial (19%)
Length = 557
Score = 27.3 bits (59), Expect = 1.2
Identities = 18/90 (20%), Positives = 43/90 (47%)
Frame = -3
Query: 24 VSQSKVHLRWRTALICRELEDEYSALIRCRGDRESLVELINQKEVVGDLFNQLRLALQRR 83
+ + K ++W T C++ +D YS + EL + +V + FNQ +++ R
Sbjct: 411 IGKRKTSIKWTTNTFCQKDKDHYST--------TNYKELTHAPQVYAERFNQKQIS--RA 262
Query: 84 SKDRPAQTLAATNMDDEDLTESMQKLLIVM 113
+ + A L + ++ + ++ L+I++
Sbjct: 261 QQQQIAAQLQEKSQNN*SVFVKIKYLIILL 172
>TC14487 similar to UP|Q9LID9 (Q9LID9) Gb|AAD15384.1, partial (22%)
Length = 493
Score = 26.6 bits (57), Expect = 2.0
Identities = 13/56 (23%), Positives = 27/56 (48%)
Frame = -3
Query: 24 VSQSKVHLRWRTALICRELEDEYSALIRCRGDRESLVELINQKEVVGDLFNQLRLA 79
+ + K ++W T C++ +D YS + EL + +V + FNQ +++
Sbjct: 416 IGKRKTSIKWTTNTFCQKDKDHYST--------TNYKELTHAPQVYAERFNQKQIS 273
>TC10936 weakly similar to UP|Q9C887 (Q9C887) Myosin-like protein, partial
(14%)
Length = 586
Score = 25.8 bits (55), Expect = 3.5
Identities = 17/42 (40%), Positives = 23/42 (54%), Gaps = 1/42 (2%)
Frame = +3
Query: 96 NMDDEDLTESMQKLLIVMQRLDEKI-APMLEADGELFNSRWG 136
NMD + ++SMQ L+ M R EK+ A + ADG WG
Sbjct: 27 NMDIMEQSQSMQTHLVSMAREVEKLRAELANADGR----HWG 140
>CB827332
Length = 511
Score = 25.4 bits (54), Expect = 4.6
Identities = 14/44 (31%), Positives = 20/44 (44%)
Frame = -1
Query: 65 QKEVVGDLFNQLRLALQRRSKDRPAQTLAATNMDDEDLTESMQK 108
Q+ +G L L L LQ K +PA + A + DLT +
Sbjct: 508 QRSSIGSLMLPLXLTLQH*GKSKPASSRPAFVICGHDLTSKYSR 377
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.136 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,283,819
Number of Sequences: 28460
Number of extensions: 23675
Number of successful extensions: 110
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 110
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 110
length of query: 155
length of database: 4,897,600
effective HSP length: 83
effective length of query: 72
effective length of database: 2,535,420
effective search space: 182550240
effective search space used: 182550240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 51 (24.3 bits)
Medicago: description of AC146862.18


