

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146855.3 - phase: 0
(186 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
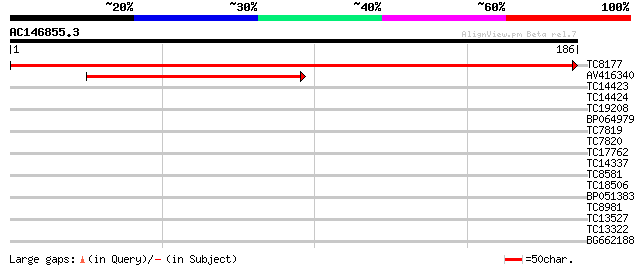
Score E
Sequences producing significant alignments: (bits) Value
TC8177 homologue to UP|RL15_ARATH (O23515) 60S ribosomal protein... 356 2e-99
AV416340 144 9e-36
TC14423 similar to UP|Q8VWX2 (Q8VWX2) Gamma-tocopherol methyltra... 32 0.051
TC14424 similar to UP|Q8VWX2 (Q8VWX2) Gamma-tocopherol methyltra... 32 0.051
TC19208 UP|O23582 (O23582) Ribosomal protein L15, partial (5%) 31 0.11
BP064979 31 0.11
TC7819 similar to UP|Q43817 (Q43817) Lipoxygenase , partial (97%) 30 0.25
TC7820 similar to UP|LOX1_LENCU (P38414) Lipoxygenase , partial... 30 0.25
TC17762 27 2.8
TC14337 similar to UP|Q94JQ8 (Q94JQ8) AT3g57880/T10K17_90, parti... 26 4.8
TC8581 similar to UP|Q94II4 (Q94II4) NAM-like protein, partial (9%) 26 4.8
TC18506 26 4.8
BP051383 25 6.2
TC8981 similar to UP|SERA_ARATH (O04130) D-3-phosphoglycerate de... 25 6.2
TC13527 similar to UP|Q39629 (Q39629) Oleosin-like protein, part... 25 8.2
TC13322 similar to UP|Q9ZS13 (Q9ZS13) RNA helicase (Fragment), p... 25 8.2
BG662188 25 8.2
>TC8177 homologue to UP|RL15_ARATH (O23515) 60S ribosomal protein L15,
complete
Length = 896
Score = 356 bits (913), Expect = 2e-99
Identities = 170/186 (91%), Positives = 178/186 (95%)
Frame = +3
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRF+QRVRCWEYRQQ SIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPV KG
Sbjct: 90 MRFLQRVRCWEYRQQPSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVPKG 269
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDST+KY+E+ILVDVA
Sbjct: 270 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTYKYYEIILVDVA 449
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
H+AIRNDPRINWL NPVHKHREL G TSAGK NRGL GKGHR+HKARPSRRA WK+NN+L
Sbjct: 450 HTAIRNDPRINWLCNPVHKHRELTGKTSAGKHNRGLVGKGHRHHKARPSRRATWKKNNSL 629
Query: 181 SLRRYR 186
SLRRYR
Sbjct: 630 SLRRYR 647
>AV416340
Length = 219
Score = 144 bits (363), Expect = 9e-36
Identities = 71/72 (98%), Positives = 72/72 (99%)
Frame = +3
Query: 26 RPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKGIVYGKPTNQGVTQLKFQRSKRSVAE 85
RPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKGIVYGKPTNQGVTQLKFQRSKRSVAE
Sbjct: 3 RPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKGIVYGKPTNQGVTQLKFQRSKRSVAE 182
Query: 86 ERAGRKLGGLRV 97
ERAGRKLGGL+V
Sbjct: 183ERAGRKLGGLKV 218
>TC14423 similar to UP|Q8VWX2 (Q8VWX2) Gamma-tocopherol methyltransferase,
partial (59%)
Length = 1239
Score = 32.3 bits (72), Expect = 0.051
Identities = 12/13 (92%), Positives = 12/13 (92%)
Frame = +3
Query: 131 NWLTNPVHKHREL 143
NWL NPVHKHREL
Sbjct: 975 NWLCNPVHKHREL 1013
>TC14424 similar to UP|Q8VWX2 (Q8VWX2) Gamma-tocopherol methyltransferase,
partial (13%)
Length = 512
Score = 32.3 bits (72), Expect = 0.051
Identities = 12/13 (92%), Positives = 12/13 (92%)
Frame = +1
Query: 131 NWLTNPVHKHREL 143
NWL NPVHKHREL
Sbjct: 244 NWLCNPVHKHREL 282
>TC19208 UP|O23582 (O23582) Ribosomal protein L15, partial (5%)
Length = 495
Score = 31.2 bits (69), Expect = 0.11
Identities = 18/31 (58%), Positives = 20/31 (64%)
Frame = +1
Query: 131 NWLTNPVHKHRELRGLTSAGKENRGLSGKGH 161
N + NPVHKHREL L + G NRG KGH
Sbjct: 247 N*VCNPVHKHRELL-L*TVGSTNRG--AKGH 330
>BP064979
Length = 506
Score = 31.2 bits (69), Expect = 0.11
Identities = 18/31 (58%), Positives = 20/31 (64%)
Frame = -2
Query: 131 NWLTNPVHKHRELRGLTSAGKENRGLSGKGH 161
N + NPVHKHREL L + G NRG KGH
Sbjct: 289 N*VCNPVHKHRELL-L*TVGSTNRG--AKGH 206
>TC7819 similar to UP|Q43817 (Q43817) Lipoxygenase , partial (97%)
Length = 2842
Score = 30.0 bits (66), Expect = 0.25
Identities = 18/43 (41%), Positives = 21/43 (47%)
Frame = -2
Query: 110 KYFEVILVDVAHSAIRNDPRINWLTNPVHKHRELRGLTSAGKE 152
K EV+L+DV H T V HRELRGL + G E
Sbjct: 2829 KNIEVVLIDVMHQ-----------TYMVTTHRELRGLDTKGNE 2734
>TC7820 similar to UP|LOX1_LENCU (P38414) Lipoxygenase , partial (12%)
Length = 497
Score = 30.0 bits (66), Expect = 0.25
Identities = 18/43 (41%), Positives = 21/43 (47%)
Frame = -1
Query: 110 KYFEVILVDVAHSAIRNDPRINWLTNPVHKHRELRGLTSAGKE 152
K EV+L+DV H T V HRELRGL + G E
Sbjct: 494 KNIEVVLIDVMHQ-----------TYMVTTHRELRGLDTKGNE 399
>TC17762
Length = 463
Score = 26.6 bits (57), Expect = 2.8
Identities = 12/20 (60%), Positives = 14/20 (70%)
Frame = -1
Query: 79 SKRSVAEERAGRKLGGLRVL 98
SK SV R+G+K GG RVL
Sbjct: 355 SKNSVVRRRSGKKDGGFRVL 296
>TC14337 similar to UP|Q94JQ8 (Q94JQ8) AT3g57880/T10K17_90, partial (16%)
Length = 1005
Score = 25.8 bits (55), Expect = 4.8
Identities = 9/18 (50%), Positives = 12/18 (66%)
Frame = +2
Query: 163 YHKARPSRRANWKRNNTL 180
+HK +P + NW RN TL
Sbjct: 23 FHKQKPPTQMNWTRNLTL 76
>TC8581 similar to UP|Q94II4 (Q94II4) NAM-like protein, partial (9%)
Length = 1158
Score = 25.8 bits (55), Expect = 4.8
Identities = 18/60 (30%), Positives = 26/60 (43%)
Frame = -2
Query: 10 WEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKGIVYGKPTNQ 69
WE R IV T T K A ++YR RR +R ++G+V +P +Q
Sbjct: 401 WERRDIKVIVSTTTTTTTTKVHHSLVLALLSLLIYRNEPRRS--QRVQNRGVVNVRPIHQ 228
>TC18506
Length = 554
Score = 25.8 bits (55), Expect = 4.8
Identities = 22/101 (21%), Positives = 44/101 (42%), Gaps = 9/101 (8%)
Frame = +3
Query: 42 VVYRVRVRRGGRKRPVSKGIVYGKPTN-QGVTQLKFQRSKRSVAEERAGR-KLGG----- 94
+VY R ++G R V ++ P+ + + + K+ AEE R KL
Sbjct: 174 IVYTRRRKKGKPVRDVIGSVISSLPSYLSDIPRRPRTKKKKFKAEEALPRPKLDSGVFDN 353
Query: 95 --LRVLNSYWVNEDSTFKYFEVILVDVAHSAIRNDPRINWL 133
+++ NS+ ++ F YF+ + + +A D + W+
Sbjct: 354 NLVKIWNSFSEDKRKPFAYFDSLWFSLYRAASSKDKVLTWI 476
>BP051383
Length = 366
Score = 25.4 bits (54), Expect = 6.2
Identities = 20/57 (35%), Positives = 31/57 (54%), Gaps = 2/57 (3%)
Frame = +3
Query: 43 VYRVRVRRGGRKRPVSKGIVYGKP--TNQGVTQLKFQRSKRSVAEERAGRKLGGLRV 97
+Y+++++R R +P+ KP N+G T LK R RS RAG + GG+ V
Sbjct: 66 LYKLQIKRKERYKPMDH-----KPGGLNRG-TALK--RPSRSTGNPRAGPRTGGMVV 212
>TC8981 similar to UP|SERA_ARATH (O04130) D-3-phosphoglycerate
dehydrogenase, chloroplast precursor (3-PGDH) , partial
(15%)
Length = 720
Score = 25.4 bits (54), Expect = 6.2
Identities = 11/16 (68%), Positives = 12/16 (74%)
Frame = +3
Query: 92 LGGLRVLNSYWVNEDS 107
LG L VL +YWVNE S
Sbjct: 96 LG*LAVLETYWVNETS 143
>TC13527 similar to UP|Q39629 (Q39629) Oleosin-like protein, partial (80%)
Length = 603
Score = 25.0 bits (53), Expect = 8.2
Identities = 14/36 (38%), Positives = 17/36 (46%)
Frame = -3
Query: 135 NPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSR 170
NP R S GKE + S +GHRYH+ R
Sbjct: 358 NPRQHRDRRRAEPSRGKEPQ--SKQGHRYHRGHQHR 257
>TC13322 similar to UP|Q9ZS13 (Q9ZS13) RNA helicase (Fragment), partial (7%)
Length = 528
Score = 25.0 bits (53), Expect = 8.2
Identities = 15/47 (31%), Positives = 24/47 (50%)
Frame = +2
Query: 13 RQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSK 59
R+++ + R TR K R+G + K+ R +VRR R+ SK
Sbjct: 173 RRENQSQKEARGTRQSKRHRIGEEEKEEEGEARQQVRRERRRNQRSK 313
>BG662188
Length = 358
Score = 25.0 bits (53), Expect = 8.2
Identities = 19/58 (32%), Positives = 28/58 (47%)
Frame = +2
Query: 20 RLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKGIVYGKPTNQGVTQLKFQ 77
RLTR RRLG ++++R R RRG KR +S ++ + V LK +
Sbjct: 140 RLTRAVPSR*RRRLGRLTSTTFILFRKR-RRGVGKRKLSGKLLKKVQNPRKVLPLKIK 310
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.136 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,089,065
Number of Sequences: 28460
Number of extensions: 38168
Number of successful extensions: 209
Number of sequences better than 10.0: 35
Number of HSP's better than 10.0 without gapping: 209
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 209
length of query: 186
length of database: 4,897,600
effective HSP length: 85
effective length of query: 101
effective length of database: 2,478,500
effective search space: 250328500
effective search space used: 250328500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146855.3


