

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
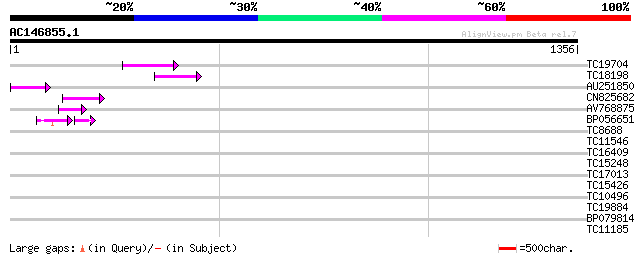
Query= AC146855.1 + phase: 0 /pseudo
(1356 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC19704 weakly similar to UP|Q9SZ87 (Q9SZ87) RNA-directed DNA po... 86 3e-17
TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, p... 74 2e-13
AU251850 66 3e-11
CN825682 65 6e-11
AV768875 60 2e-09
BP056651 42 4e-07
TC8688 homologue to UP|AAS45400 (AAS45400) Endo-1,4-beta-glucana... 30 2.0
TC11546 homologue to UP|Q940S0 (Q940S0) T5E21.14/T5E21.14 (At1g1... 30 3.4
TC16409 weakly similar to UP|O22880 (O22880) At2g40550 protein, ... 29 4.4
TC15248 weakly similar to UP|Q8GT21 (Q8GT21) Benzoyl coenzyme A:... 29 4.4
TC17013 similar to UP|O24625 (O24625) Anther-specific protein, p... 29 5.7
TC15426 weakly similar to UP|HS7E_DROME (P29845) Heat shock 70 k... 28 7.5
TC10496 28 7.5
TC19884 similar to UP|Q84KA9 (Q84KA9) RING/C3HC4/PHD zinc finger... 28 7.5
BP079814 28 9.8
TC11185 similar to UP|Q9SXW6 (Q9SXW6) Ring finger protein (Fragm... 28 9.8
>TC19704 weakly similar to UP|Q9SZ87 (Q9SZ87) RNA-directed DNA
polymerase-like protein, partial (3%)
Length = 530
Score = 86.3 bits (212), Expect = 3e-17
Identities = 43/135 (31%), Positives = 79/135 (57%), Gaps = 1/135 (0%)
Frame = -3
Query: 269 SNIVTKLSYCIEDMKYWSKANHPHFNQRKQQLKNQIDVMRNTSDASVDPRLIELQNS-LA 327
SN+ K + + W N ++ +LKNQ++ ++N S++ D + I+ + L
Sbjct: 465 SNLSGKTTNTKLALTEWHHQNFSRADKEIAELKNQLEHLQNYSNSGEDWQEIKQRKMRLK 286
Query: 328 NLILQEDVYWRQRSKIFWLKDGDKNSKFFHLTASSRRRKNTISKLRHPNGFWLTSQDDMS 387
+L QE+++W QRS+I WLK GDKN+KFFH + RR +N I +++ G W+ SQ ++
Sbjct: 285 DLWRQEELFWSQRSRIKWLKSGDKNTKFFHASTVQRRERNRIERIKDMQGTWVRSQLAIN 106
Query: 388 AHIHDYFSGLFQAIN 402
+ D++ ++++ N
Sbjct: 105 RDVPDFYPDIYKSTN 61
>TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, partial
(14%)
Length = 592
Score = 73.6 bits (179), Expect = 2e-13
Identities = 43/113 (38%), Positives = 61/113 (53%), Gaps = 1/113 (0%)
Frame = -3
Query: 346 LKDGDKNSKFFHLTASSRRRKNTISKLRHPNGFWLTSQDDMSAHIHDYFSGLFQAINGDQ 405
LK GD N++FFH + RR N I K++ +G W+ Q ++ +YFS ++ +
Sbjct: 578 LK*GDHNTRFFHASTIQRRDFNRILKIKDAHGIWVEGQKKVNEAAVNYFSDIYSSSPLQY 399
Query: 406 -QPVITRIQARVTENDNATLTKPFSIDEFKEAVFSMHSDKSPGPDGLNPGFYQ 457
+ + I VT+ N L S DE KEAVFSM K+PG DG+N FYQ
Sbjct: 398 LRECLAPIPKLVTDRLNHKLCAQVSTDEIKEAVFSMGDLKAPGMDGINGLFYQ 240
>AU251850
Length = 343
Score = 66.2 bits (160), Expect = 3e-11
Identities = 34/101 (33%), Positives = 55/101 (53%), Gaps = 5/101 (4%)
Frame = +2
Query: 1 MNVIAWNCRGLGNVKAVPCIKDLVRVYKPDIVILIETLCNNNKISGLKYAIGFDYHFSVD 60
M +++WNCRGLG+ +AV ++ LVR P +V L+ET + + G VD
Sbjct: 38 MKLLSWNCRGLGSPRAVEALQRLVRKEIPQLVFLMETKLKKFETEKISLGFGLSNCLIVD 217
Query: 61 CIG----RSGGIAVLWRNSAHCSILNYS-QHFINMSIQDPV 96
C+G R GG+A+ W S S++++S H + ++D V
Sbjct: 218 CVGDEKERKGGLAMWWNASVDVSLVSFSGNHIVVACVEDDV 340
>CN825682
Length = 626
Score = 65.5 bits (158), Expect = 6e-11
Identities = 33/101 (32%), Positives = 52/101 (50%)
Frame = -3
Query: 126 SQSDDPWCIIGDFNDHLSPSDKRGGPDRPHWLIRGFQEAVSDCNLIDLPLNGYQFTWFKS 185
S ++ W GDFN+ L +K G D+ I+ F++ + NL+DL L G +FTW +
Sbjct: 318 SNTNTFWTCYGDFNEILHHQEKEGARDQDPTRIQVFRDFLDGANLMDLELKGCRFTWLSN 139
Query: 186 IGTSHAKEARIDRALCTAPWLELFPHASLQTLVAPMSDHTP 226
+ + R+DR L W +P+A+ L SDH+P
Sbjct: 138 LRNGLVTKERLDRVLVNWTWRLDYPNATAMALPIISSDHSP 16
>AV768875
Length = 232
Score = 60.5 bits (145), Expect = 2e-09
Identities = 29/67 (43%), Positives = 40/67 (59%)
Frame = +1
Query: 118 WELLRSLHSQSDDPWCIIGDFNDHLSPSDKRGGPDRPHWLIRGFQEAVSDCNLIDLPLNG 177
W+ L ++ + PW +IGD N+ L PS RGG P+ R F + DCNLIDL + G
Sbjct: 16 WQHLSTIRASISIPWLLIGDMNEILFPSQVRGGEFHPNRAQR-FATVLDDCNLIDLGMVG 192
Query: 178 YQFTWFK 184
+FTWF+
Sbjct: 193 GKFTWFR 213
>BP056651
Length = 564
Score = 42.0 bits (97), Expect(2) = 4e-07
Identities = 25/92 (27%), Positives = 43/92 (46%), Gaps = 6/92 (6%)
Frame = +1
Query: 64 RSGGIAVLWRNSAHCSILNYSQHFINMSIQDPVKGPWR-----LTAFYGYPDHGRRRDSW 118
R+GG+ +W+ S + + F+ +KG W+ + Y G +R W
Sbjct: 103 RAGGLLCVWKPS----VFTVEECFLGEGFLG-LKGKWKGVSCVIINVYSPCSRGEKRQLW 267
Query: 119 ELLRSLHSQ-SDDPWCIIGDFNDHLSPSDKRG 149
E L++L + +D WC+ GDFN ++RG
Sbjct: 268 EDLKALRRRLEEDIWCMAGDFNSVRRSEERRG 363
Score = 30.0 bits (66), Expect(2) = 4e-07
Identities = 17/50 (34%), Positives = 25/50 (50%)
Frame = +3
Query: 156 WLIRGFQEAVSDCNLIDLPLNGYQFTWFKSIGTSHAKEARIDRALCTAPW 205
W G Q L D+PL G ++TW +S G + +R+DR L + W
Sbjct: 390 W*NGGIQRLY**MELDDIPLVGRKYTWVRSNGKA---RSRLDRFLVSHQW 530
>TC8688 homologue to UP|AAS45400 (AAS45400) Endo-1,4-beta-glucanase ,
partial (18%)
Length = 750
Score = 30.4 bits (67), Expect = 2.0
Identities = 18/47 (38%), Positives = 24/47 (50%), Gaps = 9/47 (19%)
Frame = -3
Query: 924 IDGRSVMVPASMFGTLVGYVHSLHLNPPLFLLL---------LWLSC 961
+ G + M P S+FG G+ SLH +PPL L L LW +C
Sbjct: 163 LSGPATMAPTSVFG--FGFDVSLHFHPPLQLYLTLFFGMDAPLWWTC 29
>TC11546 homologue to UP|Q940S0 (Q940S0) T5E21.14/T5E21.14
(At1g14670/T5E21.14), partial (20%)
Length = 567
Score = 29.6 bits (65), Expect = 3.4
Identities = 18/55 (32%), Positives = 25/55 (44%)
Frame = +1
Query: 943 VHSLHLNPPLFLLLLWLSCLSTICLIRICPLGIMISSILFLTTKMLQQFFLSLLG 997
+H LHL LL C CL +CP I + + + K + +F L LLG
Sbjct: 256 LHGLHL----LWLLSHARCCRFPCLAALCPPHIQVHHV*VVVVKPVGKFLLFLLG 408
>TC16409 weakly similar to UP|O22880 (O22880) At2g40550 protein, partial
(23%)
Length = 688
Score = 29.3 bits (64), Expect = 4.4
Identities = 11/41 (26%), Positives = 24/41 (57%)
Frame = -3
Query: 292 HFNQRKQQLKNQIDVMRNTSDASVDPRLIELQNSLANLILQ 332
HFN + L + + + +SD V P+L++++ + N +L+
Sbjct: 407 HFNLSLRSLSRSLTICQCSSDRLVSPKLLDMRRPIVNNLLK 285
>TC15248 weakly similar to UP|Q8GT21 (Q8GT21) Benzoyl coenzyme A: benzyl
alcohol benzoyl transferase, partial (52%)
Length = 1085
Score = 29.3 bits (64), Expect = 4.4
Identities = 15/40 (37%), Positives = 20/40 (49%), Gaps = 3/40 (7%)
Frame = -2
Query: 210 PHASLQTLVAPMSDHTPLLLQLDPPPWRAP---HNSFRFN 246
P +S ++L S H P PPPW+ P NS+R N
Sbjct: 604 PRSSHRSLS*SASPHAPAASHKAPPPWQPPQGTRNSYRSN 485
>TC17013 similar to UP|O24625 (O24625) Anther-specific protein, partial
(42%)
Length = 617
Score = 28.9 bits (63), Expect = 5.7
Identities = 12/29 (41%), Positives = 15/29 (51%)
Frame = +1
Query: 211 HASLQTLVAPMSDHTPLLLQLDPPPWRAP 239
H + + P H LLQ +PPPWR P
Sbjct: 418 HQRMGKIHRPHHPHCLRLLQRNPPPWRRP 504
>TC15426 weakly similar to UP|HS7E_DROME (P29845) Heat shock 70 kDa protein
cognate 5, partial (3%)
Length = 1216
Score = 28.5 bits (62), Expect = 7.5
Identities = 19/56 (33%), Positives = 29/56 (50%)
Frame = +3
Query: 1073 KKESLVPIPVCIVMFSQKRIHISSLCVLRLLLVGSCCNLTQSSVTCFLLRMIFPLF 1128
KK+ + P+ C+V KRI I S LR++ S L+ S C LLR + ++
Sbjct: 468 KKQIISPLRSCLVGVLMKRILILSYQNLRVI---SWSMLSSSHTDCALLRKVITIW 626
>TC10496
Length = 562
Score = 28.5 bits (62), Expect = 7.5
Identities = 12/38 (31%), Positives = 21/38 (54%)
Frame = +3
Query: 6 WNCRGLGNVKAVPCIKDLVRVYKPDIVILIETLCNNNK 43
WN RGLG+ K ++ KP++ +L E+ + N+
Sbjct: 444 WNVRGLGSSTKREAAKKILLKIKPEVCLLQESKLDENR 557
>TC19884 similar to UP|Q84KA9 (Q84KA9) RING/C3HC4/PHD zinc finger-like
protein, partial (22%)
Length = 373
Score = 28.5 bits (62), Expect = 7.5
Identities = 17/59 (28%), Positives = 21/59 (34%)
Frame = +1
Query: 181 TWFKSIGTSHAKEARIDRALCTAPWLELFPHASLQTLVAPMSDHTPLLLQLDPPPWRAP 239
+W S+ TS A T+P L P A P S H+ PPW P
Sbjct: 193 SWASSLSTSDTAPTPPQPASATSPALPAVPAAERAASTLPSSKHS--------PPWNTP 345
>BP079814
Length = 517
Score = 28.1 bits (61), Expect = 9.8
Identities = 19/48 (39%), Positives = 21/48 (43%)
Frame = -3
Query: 201 CTAPWLELFPHASLQTLVAPMSDHTPLLLQLDPPPWRAPHNSFRFNNS 248
C APWL F L TL P S H L Q + W A N +N S
Sbjct: 389 CVAPWLRNFVEHLLCTLHYPFSQH---LCQEEKLLWVAYINGSLWNTS 255
>TC11185 similar to UP|Q9SXW6 (Q9SXW6) Ring finger protein (Fragment), partial
(79%)
Length = 807
Score = 28.1 bits (61), Expect = 9.8
Identities = 9/16 (56%), Positives = 12/16 (74%)
Frame = +1
Query: 1144 RQWSYGACGKIVILSC 1159
R WSYGAC + +L+C
Sbjct: 529 RPWSYGACNRDAVLTC 576
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.338 0.147 0.484
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,163,392
Number of Sequences: 28460
Number of extensions: 468345
Number of successful extensions: 3934
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 3860
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3930
length of query: 1356
length of database: 4,897,600
effective HSP length: 101
effective length of query: 1255
effective length of database: 2,023,140
effective search space: 2539040700
effective search space used: 2539040700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146855.1


